Abstract
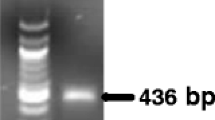
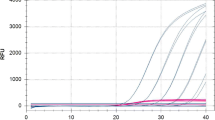
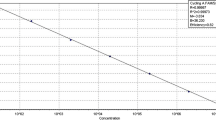
Real-time PCR assays based on SYBR® Green I and TaqMan® technologies were developed for in planta detection and quantification of Phoma tracheiphila, the mitosporic fungus causing ‘mal secco’ disease on citrus. Primers and a hybridization probe were designed on the basis of the internal transcribed spacer (ITS) region of the nuclear rRNA genes. The real-time PCR assays were compared with a classic isolation method in two separate experiments carried out on 6 and 24 month-old sour orange seedlings, artificially inoculated with a conidial suspension of the pathogen. Both technologies made it possible to follow the progression of infection by P. tracheiphila, enabling detection and quantification of the target fungus prior to the development of symptoms. The detection limit was 10 copies of the cloned target sequence and 15 pg of genomic DNA extracted from fungal spores. The values of the cycle threshold (Ct) were linearly correlated with the concentration of the target DNA, indicating that the method is suitable as a qualitative and quantitative assay. The presence of non-target fungal DNA had no effect on the specificity of the assay, but resulted in a 10-fold reduction of sensitivity. Total inhibition of the reaction occurred when conidia of the target pathogen were mixed with an organic soil substrate before extracting DNA using the standard protocol, while an alternative purification kit resulted in a significant decrease in sensitivity. Compared to classic methods, real-time PCR proved faster and easier to perform and showed a higher sensitivity. These results suggest that real-time PCR, based on both chemistries, has a great potential for early diagnosis of ‘mal secco’ disease and for quantitative estimation of fungal growth within host tissue.




Similar content being viewed by others
References
Aljanabi, S. M., & Martinez, I. (1997). Universal and rapid salt-extraction of high quality genomic DNA for PCR based techniques. Nucleic Acids Research, 25, 4692–4693.
Altschul, S. F., Madden, T. L., Schaffer, A. A., Zhang, J., Zhang, Z., & Miller, W., et al. (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research, 25, 3389–3402.
Baldacci, E., & Garofalo, F. (1948). Conoscenze e ricerche sul “mal secco” degli Agrumi. Humus, 4, 21–24.
Balmas, V., Scherm, B., Ghignone, S., Salem, A. O. M., Cacciola, S. O., & Migheli, Q. (2005). Characterisation of Phoma tracheiphila by RAPD-PCR, microsatellite-primed PCR and ITS rDNA sequencing and development of specific primers for in planta PCR detection. European Journal of Plant Pathology, 111, 235–247.
Bridge, P., & Spooner, B. (2001). Soil fungi: Diversity and detection. Plant and Soil, 232, 147–154.
Bruijnesteijn Van Coppenraet, E. S., Lindeboom, J. A., Prins, J. M., Peeters, M. F., Claas, E. C., & Kuijper, E. J., et al. (2004). Real-time PCR assay using fine-needle aspirates and tissue biopsy specimens for rapid diagnosis of mycobacterial lymphadenitis in children. Journal of Clinical Microbiology, 42, 2644–2650.
CABI\EPPO (1997) Deuterophoma tracheiphila. In I. M. Smith, D. G. Mc Namara, P. R. Scott, & M. Holderness (Eds.) Quarantine Pests for Europe (pp. 733-736), 2nd ed. Wallingford (GB): CAB International.
Cacciola, S. O., Pane, A., Magnano di San Lio, G., & Perrotta, G. (1996). Caratterizzazione di mutanti di Phoma tracheiphila (Deuteromycotina, Coelomycetes). Bollettino Accademia Gioenia Scienze Naturali, 29, 147–167 (in Italian, English summary).
Cullen, D. W., & Hirsch, P. R. (1998). Simple and rapid method for direct extraction of microbial DNA from soil for PCR. Soil Biology and Biochemistry, 30, 983–993.
Fogliano, V., Marchese, A., Scaloni, A., Ritieni, A., Visconti, A., & Randazzo, G., et al. (1998). Characterization of a 60 kDa phytotoxic glycoprotein produced by Phoma tracheiphila and its relation to malseccin. Physiological and Molecular Plant Pathology, 53, 149–161.
Gao, H., Beckman, C. H., & Müller, W. C. (1995). The rate of vascular colonization as a measure of the genotypic interaction between various cultivars of tomato and various formae or races of Fusarium oxysporum. Physiological and Molecular Plant Pathology, 46, 29–43.
Harrison, N. A., & Beckman, C. H. (1982). Time/space relationships of colonization and host responses in wilt-resistant and wilt-susceptible cotton (Gossypium) cultivars inoculated with Verticillium dahliae and Fusarium oxysporum f. sp. vasinfectum. Physiological and Molecular Plant Pathology, 21, 193–207.
Hussain, Z., Das, B. C., Husain, S. A., Polipalli, S. K., Ahmed, T., & Begum, N., et al. (2006). Virological course of hepatitis A virus as determined by real time RT-PCR: Correlation with biochemical, immunological and genotypic profiles. World Journal of Gastroenterology, 12, 4683–4688.
Licciardello, G., Grasso, F. M., Bella, P., Cirvilleri, G., Grimaldi, V., & Catara, V. (2006). Identification and detection of Phoma tracheiphila, causal agent of citrus mal secco disease, by realtime polymerase chain reaction. Plant Disease, 90, 1523–1530.
Lillie, S. H., Hanlon, E. Jr., Kelly, J. M. & Rayburn, B. B (2005). Potential military chemical/biological agents and compounds. Army Knowledge Online (www.us.army.mil).
Livak, K. J., Flood, S. P. A., Marmejo, J., Giusti, W., & Deetz, K. (1995). Oligonucleotides with fluorescent dyes at opposite ends provide a quenched probe system useful for detecting PCR products and nucleic acid hybridization. PCR Methods and Applications, 4, 357–362.
Lopez, M. M., Bertolini, E., Olmos, A., Caruso, P., Gorris, M. T., & Llop, P., et al. (2003). Innovative tools for plant pathogenic viruses and bacteria. International Microbiology, 6, 233–243.
Magnano di San Lio, G., & Graniti, A. (1987). Osservazioni sulla condizione nucleare di Phoma tracheiphila (Petri) Kanc. et Ghick. Phytopathologia Mediterranea, 26, 100–107 (in Italian, English summary).
Magnano di San Lio, G., Cacciola, S. O., Pane, A., & Grasso, S. (1992). Relationship between xylem colonization and symptom expression in mal secco infected sour orange seedlings. Proceedings International Society of Citriculture, 2, 873–876.
Maicas, S., Adam, A. C., & Polaina, J. (2000). The ribosomal DNA of the zygomycete Mucor miehei. Current Genetics, 37, 412–419.
Mercado-Blanco, J., Collado-Romero, M., Parrilla-Arraujo, S., Rodriguez-Jurado, D., & Jimenez-Diaz, R. M. (2003). Quantitative monitoring of colonization of olive genotypes by Verticillium dahliae pathotypes with real-time PCR. Physiological and Molecular Plant Pathology, 63, 91–105.
Nachmias, A., Barash, I., Solel, Z., & Strobel, G. A. (1979). Purification and characterization of a phytotoxin produced by Phoma tracheiphila, the causal agent of mal secco disease of citrus. Physiological Plant Pathology, 10, 147–157.
O’Sullivan, C. E., Kasai, M., Francesconi, A., Petraitis, V., Petraitiene, R., & Kelaher, A. M., et al. (2003). Development and validation of a quantitative real-time PCR assay using fluorescence resonance energy transfer technology for detection of Aspergillus fumigatus in experimental invasive pulmonary aspergillosis. Journal of Clinical Microbiology, 12, 5676–5682.
OEPP/EPPO (2005) Phoma tracheiphila. Bulletin OEPP/EPPO Bulletin 35: 307–311. http://www.eppo.org/STANDARDS/standards.htm.
Perrotta, G., & Graniti, A. (1988). Phoma tracheiphila (Petri) Kanchaveli & Gikashvili. In I. M. Smith, J. Dunez, R. A. Lelliott, D. H. Phillips, & S. A. Archer (Eds.) European handbook of plant diseases (pp. 396–398). Oxford, UK: Blackwell Scientific Publications.
Punithalingam, E., & Holliday, P. (1973). Deuterophoma tracheiphila. CMI Descriptions of pathogenic fungi and bacteria No. 399. Kew, UK: Commonweath Mycological Institute.
Rollo, F., Amici, A., Foresi, F., & Di Silvestro, I. (1987). Construction and characterization of a cloned probe for the detection of Phoma tracheiphila in plant tissues. Applied Microbiology and Biotechnology, 26, 352–357.
Rollo, F., Salvi, R., & Torchia, P. (1990). Highly sensitive and fast detection of Phoma tracheiphila by polymerase chain reaction. Applied Microbiology and Biotechnology, 32, 572–576.
Rozen, S., & Skaletsky, H. J. (2000). Primer3 on the WWW for general users and for biologist programmers. Methods in Molecular Biology, 132, 365–386.
Salerno, M., & Perrotta, G. (1966). Ricerche sul mal secco degli agrumi (Deuterophoma tracheiphila, Petri). Virulenza e caratteri colturali del fungo in Sicilia. Rivista di Patologia Vegetale Ser. 4° 2: 203–312 (in Italian).
Schaad, N. W., & Frederick, R. D. (2002). Real-time PCR and its application for rapid plant disease diagnostics. Canadian Journal of Plant Pathology, 24, 250–258.
Solel, Z. (1976). Epidemiology of “mal secco” disease of lemons. Phytopathologische Zeitschrift, 85, 90–92.
Solel, Z., & Salerno, M. (2000). Mal Secco. In L. W. Timmer, S. M. Garnsey, & J. H. Graham (Eds.) Compendium of citrus diseases (pp. 33–35), 2nd ed. St. Paul, MN: American Phytopathological Society Press.
Ward, E., Foster, S. J., Fraaije, B. A., & McCartney, H. A. (2004). Plant pathogen diagnostics: Immunological and nucleic acids based approaches. Annals of Applied Biology, 145, 1–16.
Acknowledgements
This work was funded by the European Union within the framework of INTERREG III A Italy–France–‘Isole’ (Project acronym: CITRUS). The authors wish to thank Franco Masia, Umberto Pirisi and Giuseppina Emonti for excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Demontis, M.A., Cacciola, S.O., Orrù, M. et al. Development of real-time PCR systems based on SYBR® Green I and TaqMan® technologies for specific quantitative detection of Phoma tracheiphila in infected Citrus . Eur J Plant Pathol 120, 339–351 (2008). https://doi.org/10.1007/s10658-007-9222-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-007-9222-9