Abstract
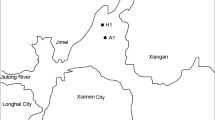
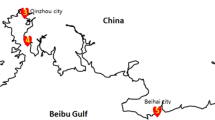
In order to increase our understanding of the microbial diversity associated with seagrass Thalassia hemprichii in Xincun Bay, South China Sea, 16S rRNA gene was identified by highthrough sequencing method. Bacteria associated with seagrass T. hemprichii belonged to 37 phyla, 99 classes. The diversity of bacteria associated with seagrass was similar among the geographically linked coastal locations of Xincun Bay. Proteobacteria was the dominant bacteria and the α-proteobacteria had adapted to the seagrass ecological niche. As well, α-proteobacteria and Pseudomonadales were associated microflora in seagrass meadows, but the interaction between the bacteria and plant is needed to further research. Burkholderiales and Verrucomicrobiae indicated the influence of the bay from anthropogenic activities. Further, Cyanobacteria could imply the difference of the nutrient conditions in the sites. γ-proteobacteria, Desulfobacterales and Pirellulales played a role in the cycle of sulfur, organic mineralization and meadow ecosystem, respectively. In addition, the less abundance bacteria species have key functions in the seagrass meadows, but there is lack knowledge of the interaction of the seagrass and less abundance bacteria species. Microbial communities can response to surroundings and play key functions in the biochemical cycle.






Similar content being viewed by others
References
Andrews JH, Harris RF (2000) The ecology and biogeography of microorganisms on plant surfaces. Annu Rev Phytopathol 38:145–180
Aravindraja C, Viszwapriya D, Pandian SK (2013) Ultradeep 16S rRNA sequencing analysis of geographically similar but diverse unexplored marine samples reveal varied bacterial community composition. PLoS One 8:e76724
Beck MW et al (2001) The identification, conservation, and management of estuarine and marine nurseries for fish and invertebrates. Bioscience 51:633–641
Boetius A et al (2000) A marine microbial consortium apparently mediating anaerobic oxidation of methane. Nature 407:623–626
Caporaso JG et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Caporaso JG et al (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624
Chao A (1987) Estimating the population size for capture-recapture data with unequal catchability. Biometrics 43:783–791
Chen M et al (2014) Dynamic succession of soil bacterial community during continuous cropping of peanut (Arachis hypogaea L.). PLoS One 9:e101355
Crump BC, Koch EW (2008) Attached bacterial populations shared by four species of aquatic angiosperms. Appl Environ Microbiol 74:5948–5957
Delille D, Canon C, Windeshausen F (1996) Comparison of planktonic and benthic bacterial communities associated with a Mediterranean Posidonia seagrass system. Bot Mar 39:239–250
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200
Feng BW, Li XR, Wang JH, Hu ZY, Meng H, Xiang LY, Quan ZX (2009) Bacterial diversity of water and sediment in the Changjiang estuary and coastal area of the East China Sea. FEMS Microbiol Ecol 70:236–248
Frankovich TA, Fourqurean JW (1997) Seagrass epiphyte loads along a nutrient availability gradient, Florida Bay, USA. Mar Ecol Prog Ser 159:37–50
Gallego S, Vila J, Nieto JM, Urdiain M, Rossello-Mora R, Grifoll M (2010) Breoghania corrubedonensis gen. nov. sp. nov., a novel alphaproteobacterium isolated from a Galician beach (NW Spain) after the Prestige fuel oil spill, and emended description of the family Cohaesibacteraceae and the species Cohaesibacter gelatinilyticus. Syst Appl Microbiol 33:316–321
Gonenc IE, Wolflin JP (2004) Coastal lagoons: ecosystem processes and modeling for sustainable use and development. CRC Press, USA
Good IJ (1952) Rational decisions. J R Stat Soc Ser B (Methodol) 14:107–114
Gutierrez JL et al (2011) Physical ecosystem engineers and the functioning of estuaries and coasts. Functioning of Estuaries and Coastal Ecosystems. Academic press, Waltham
Hamisi MI, Lyimo TJ, Muruke M (2007) Cyanobacterial occurrence and diversity in seagrass meadows in coastal Tanzania. West Indian Ocean J Mar Sci 3:113–122
Herbert R (1999) Nitrogen cycling in coastal marine ecosystems. FEMS Microbiol Rev 23:563–590
Hirano SS, Upper CD (2000) Bacteria in the leaf ecosystem with emphasis on pseudomonas syringae—a pathogen, ice nucleus, and epiphyte. Microbiol Mol Biol Rev 64:624–653
Jackson SA, Flemer B, McCann A, Kennedy J, Morrissey JP, O’Gara F, Dobson AD (2013) Archaea appear to dominate the microbiome of inflatella pellicula deep sea sponges. PLoS One 8:e84438
Kalyuzhnaya MG, Hristova KR, Lidstrom ME, Chistoserdova L (2008) Characterization of a novel methanol dehydrogenase in representatives of Burkholderiales: implications for environmental detection of methylotrophy and evidence for convergent evolution. J Bacteriol 190:3817–3823
Kreyszig E (1979) Applied Mathematics. Wiley Press, New York, p 880
Kurtz JC, Yates DF, Macauley JM, Quarles RL, Genthner FJ, Chancy CA, Devereux R (2003) Effects of light reduction on growth of the submerged macrophyte Vallisneria americana and the community of root-associated heterotrophic bacteria. J Exp Mar Biol Ecol 291:199–218
Kusel K, Pinkart HC, Drake HL, Devereux R (1999) Acetogenic and sulfate-reducing bacteria inhabiting the rhizoplane and deep cortex cells of the sea grass Halodule wrightii. Appl Environ Microbiol 65:5117–5123
Lovell CR, Decker PV, Bagwell CE, Thompson S, Matsui GY (2008) Analysis of a diverse assemblage of diazotrophic bacteria from Spartina alterniflora using DGGE and clone library screening. J Microbiol Methods 73:160–171
Magoc T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963
Mcroy CP, Goering JJ (1974) Nutrient transfer between the seagrass Zostera marina and its epiphytes. Nature 248:173–174
Mengoni A, Pini F, Huang L-N, Shu W-S, Bazzicalupo M (2009) Plant-by-plant variations of bacterial communities associated with leaves of the nickel hyperaccumulator Alyssum bertolonii Desv. Microb Ecol 58:660–667
Mohamed NM, Saito K, Tal Y, Hill RT (2009) Diversity of aerobic and anaerobic ammonia-oxidizing bacteria in marine sponges. ISME J 4:38–48
Newby B-mZ, Cutright T, Barrios CA, Xu Q (2006) Zosteric acid—an effective antifoulant for reducing fresh water bacterial attachment on coatings. J Coat Technol Res 3:69–76
Nielsen JT, Liesack W, Finster K (1999) Desulfovibrio zosterae sp. nov., a new sulfate reducer isolated from surface-sterilized roots of the seagrass Zostera marina. Int J Syst Bacteriol 49:859–865
Ning XR, Chai F, Xue H, Cai Y, Liu C, Shi J (2004) Physical-biological oceanographic coupling influencing phytoplankton and primary production in the South China Sea. J Geophys Res: Oceans (1978–2012) 109:1–20
Nourisson DH, Scapini F, Massi L, Lazzara L (2013) Optical characterization of coastal lagoons in Tunisia: ecological assessment to underpin conservation. Ecol Inf 14:79–83
Nubel U et al (1996) Sequence heterogeneities of genes encoding 16S rRNAs in Paenibacillus polymyxa detected by temperature gradient gel electrophoresis. J Bacteriol 178:5636–5643
Ondov BD, Bergman NH, Phillippy AM (2011) Interactive metagenomic visualization in a Web browser. BMC Bioinformatics 12:385
Palenik B et al (2003) The genome of a motile marine synechococcus. Nature 424:1037–1042
Partensky F, Hess W, Vaulot D (1999) Prochlorococcus, a marine photosynthetic prokaryote of global significance. Microbiol Mol Biol Rev 63:106–127
Perez Pantoja D, Donoso R, Agullo L, Cordova M, Seeger M, Pieper DH, Gonzalez B (2012) Genomic analysis of the potential for aromatic compounds biodegradation in Burkholderiales. Environ Microbiol 14:1091–1117
Pini F, Frascella A, Santopolo L, Bazzicalupo M, Biondi EG, Scotti C, Mengoni A (2012) Exploring the plant-associated bacterial communities in Medicago sativa L. BMC Microbiol 12:1471–2180
Rabus R, Hansen TA, Widdel F (2006) Dissimilatory sulfate-and sulfur-reducing prokaryotes. The prokaryotes. Springer, New York, pp 659–768
Scanlan DJ, West NJ (2002) Molecular ecology of the marine cyanobacterial genera Prochlorococcus and Synechococcus. FEMS Microbiol Ecol 40:1–12
Soetaert K, Heip C (1990) Sample-size dependence of diversity indices and the determination of sufficient sample size in a high-diversity deep-sea environment. Mar Ecol Prog Ser 59:305–307
Sundqvist MK, Sanders NJ, Wardle DA (2013) Community and ecosystem responses to elevational gradients: processes, mechanisms, and insights for global change. Annu Rev Ecol Evol Syst 44:261–280
Uku J (2007) Characterization and comparison of prokaryotic epiphytes associated with three East African seagrasses. J Phycol 43:768–779
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Weidner S, Arnold W, Stackebrandt E, Pühler A (2000) Phylogenetic analysis of bacterial communities associated with leaves of the seagrass Halophila stipulacea by a culture-independent small-subunit rRNA gene approach. Microb Ecol 39:22–31
Yang D, Huang DJ (2011) Impacts of Typhoons Tianying and Dawei on seagrass distribution in Xincun Bay, Hainan Province, China. Acta Oceanol Sin 30:32–39
Yang D, Yang C (2009) Detection of seagrass distribution changes from 1991 to 2006 in Xincun Bay, Hainan, with satellite remote sensing. Sensors 9:830–844
Yu ZQ, Deng H, Wu KW, Du J, Ma M (2012) Nutrient contents of dominant seagrass species and their affecting factors in Hainan Province. J East China Norm Univ (Nat Sci) 4:017
Zhang Y, Zhao Z, Dai M, Jiao N, Herndl GJ (2014) Drivers shaping the diversity and biogeography of total and active bacterial communities in the South China Sea. Mol Ecol 23:2260–2274
Acknowledgments
This research was supported by the National Natural Science Foundation of China (Nos. 41276113, 41276114, 41406191 and 41430966), the National High Technology Research and Development Program of China (Nos. 2012AA092104, 2013AA092901 and 2013AA092902), the Strategic Priority Research Program of the Chinese Academy of Sciences (No. XDA11020202), the Science and technology cooperation Projects of Sanya (No. 2013YD74), and the Sanya Station Database and the Information System of CERN, the Open Fund of Key Laboratory for Ecological Environment in Coastal Areas, State Oceanic Administration (No. 201304).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jiang, YF., Ling, J., Dong, JD. et al. Illumina-based analysis the microbial diversity associated with Thalassia hemprichii in Xincun Bay, South China Sea. Ecotoxicology 24, 1548–1556 (2015). https://doi.org/10.1007/s10646-015-1511-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10646-015-1511-z