Abstract
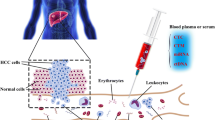
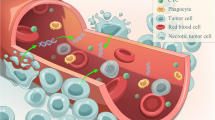
Hepatocellular carcinoma ranks fourth in cancer-related causes of death worldwide and second in China. Patients with hepatocellular carcinoma (HCC) at the early stage have a better prognosis compared to HCC patients at the late stage. Therefore, early screening for HCC is critical for clinical treatment decisions and improving the prognosis of patients. Ultrasound (US), computed tomography (CT), and serum alpha fetoprotein (AFP) have been used to screen HCC, but HCC is still difficult to be diagnosed in the early stage due to the low sensitivity of the above methods. It is urgent to find a method with high sensitivity and specificity for the early diagnosis of HCC. Liquid biopsy is a noninvasive detection method using blood or other bodily fluids. Cell-free DNA (cfDNA) and circulating tumor DNA (ctDNA) are important biomarkers for liquid biopsy. Recently, HCC screening methods using the application of cfDNA and ctDNA have become the hot spot of early HCC diagnostics. In this mini review, we summarize the latest research progress of liquid biopsy based on blood cfDNA in early screening of HCC.
Similar content being viewed by others
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Villanueva A, Hepatocellular Carcinoma N, Engl JM (2019) 380 (15),1450–1462. https://doi.org/10.1056/NEJMra1713263
Heimbach JK, Kulik LM, Finn RS, Sirlin CB, Abecassis MM, Roberts LR, Zhu AX, Murad MH, Marrero J (2018) A. AASLD Guidelines for the treatment of Hepatocellular Carcinoma: Heimbach et Al. Hepatology 67(1):358–380. https://doi.org/10.1002/hep.29086
El-Serag HB (2012) Epidemiology of viral Hepatitis and Hepatocellular Carcinoma. Gastroenterology 142(6):1264–1273. e1
Yang JD, Roberts LR, Hepatocellular Carcinoma (2010) A Global View. Nat Rev Gastroenterol Hepatol 7(8):448–458. https://doi.org/10.1038/nrgastro.2010.100
Yang JD, Hainaut P, Gores GJ, Amadou A, Plymoth A, Roberts LR (2019) A Global View of Hepatocellular Carcinoma: Trends, Risk, Prevention and Management. Nat Rev Gastroenterol Hepatol 16(10):589–604. https://doi.org/10.1038/s41575-019-0186-y
Lampertico P, Agarwal K, Berg T, Buti M, Janssen HLA, Papatheodoridis G, Zoulim F, Tacke F EASL 2017 clinical practice guidelines on the management of Hepatitis B Virus infection. Journal of Hepatology 2017, 67 (2),370–398. https://doi.org/10.1016/j.jhep.2017.03.021
Wu X, Li J, Gassa A, Buchner D, Alakus H, Dong Q, Ren N, Liu M, Odenthal M, Stippel D, Bruns C, Zhao Y, Wahba R (2020) Circulating tumor DNA as an emerging Liquid Biopsy Biomarker for early diagnosis and therapeutic monitoring in Hepatocellular Carcinoma. Int J Biol Sci 16(9):1551–1562. https://doi.org/10.7150/ijbs.44024
Lang H, Sotiropoulos GC, Brokalaki EI, Schmitz KJ, Bertona C, Meyer G, Frilling A, Paul A, Malagó M, Broelsch CE (2007) Survival and recurrence rates after resection for Hepatocellular Carcinoma in noncirrhotic livers. J Am Coll Surg 205(1):27–36. https://doi.org/10.1016/j.jamcollsurg.2007.03.002
NCCN Guidelines for Hepatobiliary Cancers.
Tzartzeva K Surveillance imaging and alpha fetoprotein for early detection of Hepatocellular Carcinoma in patients with cirrhosis: a Meta-analysis. 34. https://doi.org/10.1053/j.gastro.2018.01.064
Zhang X, Wang Z, Tang W, Wang X, Liu R, Bao H, Chen X, Wei Y, Wu S, Bao H, Wu X, Shao Y, Fan J, Zhou J (2022) Ultrasensitive and affordable assay for early detection of primary Liver Cancer using plasma cell-free DNA fragmentomics. Hepatology 76(2):317–329. https://doi.org/10.1002/hep.32308
Guo J, Seo Y, Ren S, Hong S, Lee D, Kim S, Jiang Y (2016) Diagnostic performance of contrast-enhanced Multidetector Computed Tomography and Gadoxetic Acid Disodium-Enhanced magnetic resonance imaging in detecting Hepatocellular Carcinoma: direct comparison and a Meta-analysis. Abdom Radiol 41(10):1960–1972. https://doi.org/10.1007/s00261-016-0807-7
The PreCar Team, Chen L, Abou-Alfa GK, Zheng B, Liu J-F, Bai J, Du L-T, Qian Y-S, Fan R, Liu X-L, Wu L, Hou J-L, Wang H-Y (2021) Genome-scale profiling of circulating cell-free DNA signatures for early detection of Hepatocellular Carcinoma in Cirrhotic Patients. Cell Res 31(5):589–592. https://doi.org/10.1038/s41422-020-00457-7
Harris PS, Hansen RM, Gray ME, Massoud OI, McGuire BM, Shoreibah MG (2019) Hepatocellular Carcinoma Surveillance: an evidence-based Approach. WJG 25(13):1550–1559. https://doi.org/10.3748/wjg.v25.i13.1550
Crowley E, Di Nicolantonio F, Loupakis F, Bardelli A (2013) Liquid Biopsy: Monitoring Cancer-Genetics in the blood. Nat Rev Clin Oncol 10(8):472–484. https://doi.org/10.1038/nrclinonc.2013.110
Issa IA, Noureddine M (2017) Colorectal Cancer Screening: An Updated Review of the Available Options. WJG 23 (28), 5086. https://doi.org/10.3748/wjg.v23.i28.5086
Hench IB, Hench J, Tolnay M (2018) Liquid biopsy in clinical management of breast, lung, and Colorectal Cancer. Front Med 5:9. https://doi.org/10.3389/fmed.2018.00009
Morrison GJ, Goldkorn A (2018) Development and application of Liquid Biopsies in metastatic prostate Cancer. Curr Oncol Rep 20(4):35. https://doi.org/10.1007/s11912-018-0683-0
Chung JH, Pavlick D, Hartmaier R, Schrock AB, Young L, Forcier B, Ye P, Levin MK, Goldberg M, Burris H, Gay LM, Hoffman AD, Stephens PJ, Frampton GM, Lipson DM, Nguyen DM, Ganesan S, Park BH, Vahdat LT, Leyland-Jones B, Mughal TI, Pusztai L, O’Shaughnessy J, Miller VA, Ross JS, Ali SM (2017) Hybrid capture-based genomic profiling of circulating Tumor DNA from patients with estrogen receptor-positive metastatic breast Cancer. Ann Oncol 28(11):2866–2873. https://doi.org/10.1093/annonc/mdx490
Pan Y, Ji JS, Jin JG, Kuo WP, Kang H (2017) Cancer Liquid Biopsy: is it ready for clinic? IEEE Pulse 8(1):23–27. https://doi.org/10.1109/MPUL.2016.2630838
Moss J, Magenheim J, Neiman D, Zemmour H, Loyfer N, Korach A, Samet Y, Maoz M, Druid H, Arner P, Fu K-Y, Kiss E, Spalding KL, Landesberg G, Zick A, Grinshpun A, Shapiro AMJ, Grompe M, Wittenberg AD, Glaser B, Shemer R, Kaplan T, Dor Y (2018) Comprehensive Human cell-type methylation Atlas reveals Origins of circulating cell-free DNA in Health and Disease. Nat Commun 9(1):5068. https://doi.org/10.1038/s41467-018-07466-6
Lyu X, Tsui Y-M, Ho DW-H, Ng IO-L (2022) Liquid Biopsy using cell-free or circulating tumor DNA in the management of Hepatocellular Carcinoma. Cell Mol Gastroenterol Hepatol 13(6):1611–1624. https://doi.org/10.1016/j.jcmgh.2022.02.008
Howell J, Atkinson SR, Pinato DJ, Knapp S, Ward C, Minisini R, Burlone ME, Leutner M, Pirisi M, Büttner R, Khan SA, Thursz M, Odenthal M, Sharma R (2019) Identification of mutations in circulating cell-free Tumour DNA as a Biomarker in Hepatocellular Carcinoma. Eur J Cancer 116:56–66. https://doi.org/10.1016/j.ejca.2019.04.014
Yan L, Chen Y, Zhou J, Zhao H, Zhang H, Wang G Diagnostic value of circulating cell-free DNA levels for Hepatocellular Carcinoma. International Journal of Infectious Diseases 2018, 67,92–97. https://doi.org/10.1016/j.ijid.2017.12.002
Xu H, Zhu X, Xu Z, Hu Y, Bo S, Xing T, Zhu K (2015) Non-invasive analysis of genomic Copy Number Variation in patients with Hepatocellular Carcinoma by Next Generation DNA sequencing. J Cancer 6(3):247–253. https://doi.org/10.7150/jca.10747
Tao K, Bian Z, Zhang Q, Guo X, Yin C, Wang Y, Zhou K, Wan S, Shi M, Bao D, Yang C, Xing J (2020) Machine learning-based genome-wide interrogation of somatic Copy Number Aberrations in circulating Tumor DNA for early detection of Hepatocellular Carcinoma. EBioMedicine 56:102811. https://doi.org/10.1016/j.ebiom.2020.102811
Guichard C, Amaddeo G, Imbeaud S, Ladeiro Y, Pelletier L, Maad IB, Calderaro J, Bioulac-Sage P, Letexier M, Degos F, Clément B, Balabaud C, Chevet E, Laurent A, Couchy G, Letouzé E, Calvo F, Zucman-Rossi J (2012) Integrated Analysis of somatic mutations and Focal Copy-Number changes identifies key genes and pathways in Hepatocellular Carcinoma. Nat Genet 44(6):694–698. https://doi.org/10.1038/ng.2256
Kaseb AO, Sánchez NS, Sen S, Kelley RK, Tan B, Bocobo AG, Lim KH, Abdel-Wahab R, Uemura M, Pestana RC, Qiao W, Xiao L, Morris J, Amin HM, Hassan MM, Rashid A, Banks KC, Lanman RB, Talasaz A, Mills-Shaw KR, George B, Haque A, Raghav KPS, Wolff RA, Yao JC, Meric-Bernstam F, Ikeda S, Kurzrock R (2019) Molecular profiling of Hepatocellular Carcinoma using circulating cell-free DNA. Clin Cancer Res 25(20):6107–6118. https://doi.org/10.1158/1078-0432.CCR-18-3341
Totoki Y, Tatsuno K, Covington KR, Ueda H, Creighton CJ, Kato M, Tsuji S, Donehower LA, Slagle BL, Nakamura H, Yamamoto S, Shinbrot E, Hama N, Lehmkuhl M, Hosoda F, Arai Y, Walker K, Dahdouli M, Gotoh K, Nagae G, Gingras M-C, Muzny DM, Ojima H, Shimada K, Midorikawa Y, Goss JA, Cotton R, Hayashi A, Shibahara J, Ishikawa S, Guiteau J, Tanaka M, Urushidate T, Ohashi S, Okada N, Doddapaneni H, Wang M, Zhu Y, Dinh H, Okusaka T, Kokudo N, Kosuge T, Takayama T, Fukayama M, Gibbs RA, Wheeler DA, Aburatani H (2014) Shibata, T. Trans-Ancestry Mutational Landscape of Hepatocellular Carcinoma Genomes. Nat Genet 46(12):1267–1273. https://doi.org/10.1038/ng.3126
Qu C, Wang Y, Wang P, Chen K, Wang M, Zeng H, Lu J, Song Q, Diplas BH, Tan D, Fan C, Guo Q, Zhu Z, Yin H, Jiang L, Chen X, Zhao H, He H, Wang Y, Li G, Bi X, Zhao X, Chen T, Tang H, Lv C, Wang D, Chen W, Zhou J, Zhao H, Cai J, Wang X, Wang S, Yan H, Zeng Y-X, Cavenee WK, Jiao Y (2019) Detection of Early-Stage Hepatocellular Carcinoma in Asymptomatic HBsAg-Seropositive Individuals by Liquid Biopsy. Proc. Natl. Acad. Sci. U.S.A. 116 (13), 6308–6312. https://doi.org/10.1073/pnas.1819799116
Huang A, Zhang X, Zhou S-L, Cao Y, Huang X-W, Fan J, Yang X-R, Zhou J (2016) Plasma circulating Cell-Free DNA Integrity as a Promising Biomarker for diagnosis and surveillance in patients with Hepatocellular Carcinoma. J Cancer 7(13):1798–1803. https://doi.org/10.7150/jca.15618
Jiang P, Chan CWM, Chan KCA, Cheng SH, Wong J, Wong VW-S, Wong GLH, Chan SL, Mok TSK, Chan HLY, Lai PBS, Chiu RWK, Lo YMD (2015) Lengthening and Shortening of Plasma DNA in Hepatocellular Carcinoma Patients. Proc. Natl. Acad. Sci. U.S.A. 112 (11). https://doi.org/10.1073/pnas.1500076112
Meng Z, Ren Q, Zhong G, Li S, Chen Y, Wu W, Feng Y, Mao M, Zhang F, Long G (2021) Noninvasive detection of Hepatocellular Carcinoma with circulating Tumor DNA features and α-Fetoprotein. J Mol Diagn 23(9):1174–1184. https://doi.org/10.1016/j.jmoldx.2021.06.003
Jiang P, Sun K, Tong YK, Cheng SH, Cheng THT, Heung MMS, Wong J, Wong VWS, Chan HLY, Chan KCA, Lo YMD, Chiu RWK (2018) Preferred End Coordinates and Somatic Variants as Signatures of Circulating Tumor DNA Associated with Hepatocellular Carcinoma. Proc. Natl. Acad. Sci. U.S.A. 115 (46). https://doi.org/10.1073/pnas.1814616115
Jiang P, Sun K, Peng W, Cheng SH, Ni M, Yeung PC, Heung MMS, Xie T, Shang H, Zhou Z, Chan RWY, Wong J, Wong VWS, Poon LC, Leung TY, Lam WKJ, Chan JYK, Chan HLY, Chan KCA, Chiu RWK, Lo YM (2020) D. plasma DNA end-motif profiling as a fragmentomic marker in Cancer, pregnancy, and transplantation. Cancer Discov 10(5):664–673. https://doi.org/10.1158/2159-8290.CD-19-0622
Manel E Epigenetics in Cancer. N engl j med 2008, 12. https://doi.org/10.1056/NEJMra072067
Chatterjee R, Vinson C, CpG Methylation Recruits Sequence Specific Transcription Factors Essential for Tissue Specific Gene Expression (2012) Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms. 1819(7):763–770. https://doi.org/10.1016/j.bbagrm.2012.02.014
Kisiel JB, Dukek BA, Kanipakam VSR, Ghoz R, Yab HM, Berger TC, Taylor CK, Foote WR, Giama PH, Onyirioha NH, Abdallah K, Burger MA, Slettedahl KN, Mahoney SW, Smyrk DW, Lewis TC, Giakoumopoulos JT, Allawi M, Lidgard HT, Roberts GP, Ahlquist LR (2019) D. A. Hepatocellular Carcinoma Detection by Plasma Methylated DNA: Discovery, Phase I Pilot, and Phase II Clinical Validation. Hepatology hep.30244. https://doi.org/10.1002/hep.30244
Xu R, Wei W, Krawczyk M, Wang W, Luo H, Flagg K, Yi S, Shi W, Quan Q, Li K, Zheng L, Zhang H, Caughey BA, Zhao Q, Hou J, Zhang R, Xu Y, Cai H, Li G, Hou R, Zhong Z, Lin D, Fu X, Zhu J, Duan Y, Yu M, Ying B, Zhang W, Wang J, Zhang E, Zhang C, Li O, Guo R, Carter H, Zhu J, Hao X, Zhang K (2017) Circulating Tumour DNA methylation markers for diagnosis and prognosis of Hepatocellular Carcinoma. Nat Mater 16(11):1155–1161. https://doi.org/10.1038/nmat4997
Luo B, Ma F, Liu H, Hu J, Rao L, Liu C, Jiang Y, Kuangzeng S, Lin X, Wang C, Lei Y, Si Z, Chen G, Zhou N, Liang C, Jiang F, Liu F, Dai W, Liu W, Gao Y, Li Z, Li X, Zhou G, Li B, Zhang Z, Nian W, Luo L, Liu X (2022) Cell-free DNA methylation markers for Differential diagnosis of Hepatocellular Carcinoma. BMC Med 20(1):8. https://doi.org/10.1186/s12916-021-02201-3
Chalasani NP, Ramasubramanian TS, Bhattacharya A, Olson MC, Edwards V, Roberts DK, Kisiel LR, Reddy JB, Lidgard KR, Johnson GP, Bruinsma SC (2021) A novel blood-based panel of methylated DNA and protein markers for detection of early-stage Hepatocellular Carcinoma. Clin Gastroenterol Hepatol 19(12):2597–2605e4. https://doi.org/10.1016/j.cgh.2020.08.065
Chalasani NP, Porter K, Bhattacharya A, Book AJ, Neis BM, Xiong KM, Ramasubramanian TS, Edwards DK, Chen I, Johnson S, Roberts LR, Kisiel JB, Reddy KR, Singal AG, Olson MC, Bruinsma JJ (2022) Validation of a Novel Multitarget Blood Test shows high sensitivity to Detect Early Stage Hepatocellular Carcinoma. Clin Gastroenterol Hepatol 20(1):173–182e7. https://doi.org/10.1016/j.cgh.2021.08.010
Klug M, Schmidhofer S, Gebhard C, Andreesen R, Rehli M (2013) 5-Hydroxymethylcytosine is an essential Intermediate of active DNA demethylation processes in primary human monocytes. 8. https://doi.org/10.1186/gb-2013-14-5-r46
Cai J, Chen L, Zhang Z, Zhang X, Lu X, Liu W, Shi G, Ge Y, Gao P, Yang Y, Ke A, Xiao L, Dong R, Zhu Y, Yang X, Wang J, Zhu T, Yang D, Huang X, Sui C, Qiu S, Shen F, Sun H, Zhou W, Zhou J, Nie J, Zeng C, Stroup EK, Zhang X, Chiu BC-H, Lau WY, He C, Wang H, Zhang W, Fan J (2019) Genome-wide mapping of 5-Hydroxymethylcytosines in circulating cell-free DNA as a Non-Invasive Approach for early detection of Hepatocellular Carcinoma. Gut 68(12):2195–2205. https://doi.org/10.1136/gutjnl-2019-318882
Ye W, Siwko S, Tsai RYL (2021) Sex and race-related DNA methylation changes in Hepatocellular Carcinoma. IJMS 22(8):3820. https://doi.org/10.3390/ijms22083820
Lin N, Lin Y, Xu J, Liu D, Li D, Meng H, Gallant MA, Kubota N, Roy D, Li JS, Gorospe EC, Sherman M, Gish RG, Abou-Alfa GK, Nguyen MH, Taggart DJ, Van Etten RA, Hoshida Y, Li W (2022) A Multi‐analyte cell‐free DNA –Based blood test for early detection of Hepatocellular Carcinoma. Hepatol Commun 6(7):1753–1763. https://doi.org/10.1002/hep4.1918
Yang X, Zhou J, Fan J et al (2022) Early detection of hepatocellular carcinoma with methylation and fragmentation signatures of circulating tumour DNA: a prospective, multicentre, case-control, observational study. Lancet Oncol 23:S15. https://doi.org/10.1016/S1470-2045(22)00414-4
Lewin J, Kottwitz D, Aoyama J et al (2021) Plasma cell free DNA methylation markers for hepatocellular carcinoma surveillance in patients with cirrhosis: a case control study. BMC Gastroenterol 21(1):136. https://doi.org/10.1186/s12876-021-01714-8
Kotoh Y, Suehiro Y, Saeki I et al (2020) Novel Liquid Biopsy Test based on a sensitive methylated SEPT9 assay for diagnosing Hepatocellular Carcinoma. Hepatol Commun 0(0):10. https://doi.org/10.1002/hep4.1469
Oussalah A, Rischer S, Bensenane M et al (2018) Plasma mSEPT9: a novel circulating cell-free DNA-Based epigenetic biomarker to diagnose Hepatocellular Carcinoma. EBioMedicine 30:138–147. https://doi.org/10.1016/j.ebiom.2018.03.029
Qi T, Pan M, Shi H, Wang L, Bai Y, Ge Q, Cell-Free DNA (2023) Fragmentomics: The Novel Promising Biomarker. IJMS. ;24(2):1503. doi:https://doi.org/10.3390/ijms24021503
van der Pol Y, Moldovan N, Verkuijlen S et al (2022) The Effect of Preanalytical and physiological variables on cell-free DNA fragmentation. Clin Chem 68(6):803–813. https://doi.org/10.1093/clinchem/hvac029
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
Shiqi Hu: Investigation, Data Curation, Formal analysis, Writing - Original Draft;Yaqin Liu: Investigation, Data Curation, Writing - Original Draft;Qidong Yang: Investigation, Data Curation, Writing - Original Draft;Lin Chen: Data Curation;Huizi Chai: Data Curation;Mingzhe Xiao: Formal analysis, Writing - Review & Editing;Chuang Qi: Formal analysis, Writing - Review & Editing;Wei Qiu [Corresponding author]: Conceptualization, Methodology, Writing - Review & Editing;.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Competing interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, S., Liu, Y., Yang, Q. et al. Liquid biopsy using cell-free DNA in the early diagnosis of hepatocellular carcinoma. Invest New Drugs 41, 532–538 (2023). https://doi.org/10.1007/s10637-023-01363-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10637-023-01363-6