Abstract
The tule elk (Cervus canadensis nannodes) is a California endemic subspecies that experienced an extreme bottleneck (potentially two individuals) in the mid-1800s. Through active management, including reintroductions, the subspecies has grown to approximately 6000 individuals spread across 22 recognized populations. The populations tend to be localized and separated by unoccupied intervening habitat, prompting targeted translocations to ensure gene flow. However, little is known about the genetic status or connectivity among adjacent populations in the absence of active translocations. We used 19 microsatellites and a sex marker to obtain baseline data on the genetic effective population sizes and functional genetic connectivity of four of these populations, three of which were established since the 1980s and one of which was established ~ 100 years ago. A Bayesian assignment approach suggested the presence of 5 discrete genetic clusters, which corresponded to the four primary populations and two subpopulations within the oldest of them. Effective population sizes ranged from 15 (95% CI 10–22) to 51 (95% CI 32–88). We detected little or no evidence of gene flow among most populations. Exceptions were a signature of unidirectional gene flow to one population founded by emigrants of the other 30 years earlier, and bidirectional gene flow between subpopulations within the oldest population. We propose that social cohesion more than landscape characteristics explained population structure, which developed over many generations corresponding to population expansion. Whether or which populations can grow and reach sufficient effective population sizes on their own or require translocations to maintain genetic diversity and population growth is unclear. In the future, we recommend pairing genetic with demographic monitoring of these and other reintroduced elk populations, including targeted monitoring following translocations to evaluate their effects and necessity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fragmentation of wildlife populations as a consequence of human activity can result in inbreeding and a loss of genetic variation, leading to lower fitness levels (i.e., inbreeding depression) and reduced adaptive potential (Frankham et al. 2017). Patchily distributed species that have also experienced historical bottlenecks may be especially susceptible to the negative effects of fragmentation (Allendorf and Luikart 2009; Frankham et al. 2017). Landscape connectivity is therefore desirable for maintenance of gene flow, particularly in small populations. In the absence of landscape connectivity, translocations have been used as a management tool to augment genetic diversity and improve fitness of recipient populations (Frankham 2015; Whiteley et al. 2015). Characterizing the genetic structure of populations allows wildlife managers to quantify genetic variation, infer connective corridors and barriers, and inform management activities based on predicted genetic outcomes, such as if and where human-mediated translocations should occur (Buchalski et al. 2015; Frankham et al. 2019).
The tule elk (Cervus canadensis nannodes), a subspecies endemic to California, currently persists in a metapopulation, where demographic and genetic connectivity is thought to be low among populations (McCullough et al. 1996). Although historically numerous, by the late 1800s tule elk rapidly declined to as few as two individuals, culminating in an extreme genetic bottleneck (Kucera 1991; McCullough 1969; Meredith et al. 2007; Sacks et al. 2016). Through intense management efforts, tule elk have recovered demographically from near extinction to nearly 6000 individuals distributed across 22 recognized populations (CDFW 2018; Ciriacy-Wantrup and Phillips 1970; McCullough et al. 1996). Despite this successful demographic recovery, tule elk populations today continue to exhibit low genetic diversity, which could be further reduced if populations do not grow at a sufficient rate or experience gene flow (Meredith et al. 2007; Sacks et al. 2016; Williams et al. 2004). For example, tule elk heterozygosity (0.18) was just over 1/3rd that of Rocky Mountain (C. c. nelsoni) and Manitoban (C. c. Manitobensis) elk (both 0.51) using one set of microsatellite markers (Williams et al. 2004).
Other than cursory investigations of genetic structure of tule elk at the range-wide scale among non-adjacent populations (Kucera 1991; Meredith et al. 2007; Williams et al. 2004), no study has attempted to quantify connectivity among adjacent populations. The conventional wisdom is that reintroduced populations of tule elk tend to remain together and isolated from other populations corresponding to different reintroduction sites, possibly reinforced by heterogeneous habitat (McCullough et al. 1996). However, even occasional dispersal may be sufficient to link populations genetically (Lowe and Allendorf 2010). The most important concern is that gene flow be high enough to prevent inbreeding depression, which can prevent the population growth needed to increase and maintain effective population size.
We used a panel of 19 microsatellites and a sex marker to assess the population structure of tule elk in Mendocino, Lake, and Colusa Counties, CA, in 4 discrete (non-contiguous) populations (CDFW 2018). Two of these populations were established directly through human-mediated reintroduction events, Cache Creek (CC) established with 21 individuals in 1922 and Lake Pillsbury (LPB) established from 94 individuals between 1978 and 1980 (McCullough 1969; McCullough et al. 1996; CDFW 2018). The Potter Valley (PV) population was established in 1978 by an unknown, but presumed large, number of individuals dispersing from the LPB reintroduction (Batter 2020; CDFW 2018). The East Park Reservoir (EPR) population was established in its current location by 1992, presumably from a 1985 reintroduction site several kilometers away that was otherwise unsuccessful (CDFW 2018; McCullough 1969; McCullough et al. 1996). The oldest of these populations, the CC population, gradually expanded its range, and also received multiple augmentation events, all potentially introducing substructure (CDFW 2018).
The connectivity within and among these 4 populations is unknown. Although populations occupy discrete areas on the landscape and use habitat nonrandomly (e.g., Batter 2020), there are no obvious barriers to movements among populations. Therefore, we investigated genetic structure, including contemporary gene flow and genetic effective populations sizes (Ne).
Materials and methods
Study area
The study area encompassed ~ 9500 km2 of the California Coast and Interior Coast mountain ranges. The region has a Mediterranean climate characterized by hot, dry summers and mild, wet winters (Kauffman 2003). Year-round temperatures range from 0 °C in the winter to > 38 °C summer (CDFW 2018). Average annual precipitation is ~ 76 cm, most of which occurs from October through May. This region is characterized by a rugged landscape, with rolling foothills and flats that permeate jagged peaks and valleys with elevations ranging 30–2175 m. Dominant vegetation communities include blue oak (Quercus douglasii) woodland, perennial grasslands, chamise (Adenostoma fasciculatum) and redshank (A. sparsifolium) chaparral, blue oak and foothill pine (Pinus sabiniana) woodlands, mixed conifer and hardwood forests, annual grasslands, lacustrine, and agricultural pastures. In general, female elk were concentrated in valleys dominated by grasslands, with males dispersed more broadly around females (Batter 2020). The LPB site contained ~ 5 km2 lake basin area of grassland and mixed hardwood where elk, especially females, tended to concentrate, and was surrounded by mountain peaks with chaparral and dense-canopy forests that elk tended to avoid. In contrast, valley grasslands generally thought to be good elk habitat spanned the intervening space between EPR and CC, and a mosaic of habitat types variably used by elk spanned CC and intervened between CC and PV, although the latter span included the densely human-populated northern shoreline of Clear Lake.
Genetic sample collection
We collected samples from the four tule elk populations (Fig. 1). We used a combination of fecal DNA and tissue samples, collected as part of a broader study of the Lake-Colusa County tule elk metapopulation (Batter 2020). During fecal pellet field surveys conducted Jun–Aug of 2017–2019 we collected 5–8 fecal pellets from each pellet group. Pellets were stored in 95–100% ethanol at room temperature for 1–4 months until DNA extraction. Tissue samples were obtained from the California Department of Fish and Wildlife (CDFW) from hunter-harvested elk (muscle tissue stored in 10 mm desiccant beads) and elk captured as part of a separate telemetry study, specifically 5–9 mm ear tissue biopsy punch.
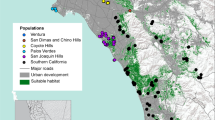
Genetic sample locations of a female tule elk (n = 253), b male tule elk (n = 237), and geographic population units, Potter Valley (PV), Lake Pillsbury (LPB), East Park Reservoir (EPR), Cache Creek-Rocky Quarry (CCRQ), and Cache Creek-Cortina Ridge (CCCR) in Mendocino, Lake, and Colusa counties, CA. Major water bodies are indicated by darker gray polygons
Genetic analyses
We completed all laboratory analyses at the Mammalian Ecology and Conservation Unit of the University of California, Davis Veterinary Genetics Laboratory. For fecal samples we first evaporated ethanol from 1 to 2 pellets at 21 °C overnight; we then agitated the outside of the pellets with ≥ 2 mL of buffer ATL (Qiagen, Valencia, CA, USA) for 1 h to remove epithelial cells from the outer surface of the pellet(s) into the buffer to prepare for extraction. We extracted fecal and tissue DNA using the Qiagen DNeasy Blood and Tissue kits according to the manufacturer’s protocol, except for fecal samples we eluted DNA in 50 μL of buffer AE (Qiagen) to acquire sufficiently concentrated DNA samples. We genotyped the DNA samples using 19 microsatellite markers: TE179, TE85, TE132, TE84, TE185, TE45, TE182, TE68, TE83, T501, TE169, TE105, TE88, T26, T193, T501, T172, T108, and a sex-typing marker from the Y chromosome (SRY) (Jones et al. 2002; Sacks et al. 2016). The PCR reaction conditions were described previously (Sacks et al. 2016). We used an ABI 3730 (Applied Biosystems, Grand Island, NY, USA) and internal size standards (500-LIZ; Applied Biosystems) for electrophoresis, with alleles scored manually using electropherograms visualized in Program STRand (version 2.4.89) (Toonen and Hughes 2001). We amplified each fecal DNA sample in two independent polymerase chain reactions (PCRs).
Individual identification
To ensure that only one genotype per individual was used in analyses, we first conducted a pairwise analysis of genotypes to identify replicate sample genotypes (i.e., genotypes from different samples of the same individual). To maximize accuracy and resolution, we first excluded all sample genotypes with < 18 loci. We then assigned samples to individuals based on their genotypes. To match genotypes, we needed to allow for some number of allele mismatches due to genotyping error, while minimizing the risk of erroneously assigning genotypes from two closely related but distinct individuals (i.e. siblings, parents-offspring, etc.) to the same individual. We used the R package allelematch (v. 2.5) to identify clusters of genotypes identified as unique individuals (Galpern et al. 2012). After assigning individuals, we used only one genotype, the consensus genotype, per individual for analyses.
Genetic structure
We used the program STRUCTURE (v. 2.3.4) (Pritchard et al. 2000), a Bayesian clustering algorithm that uses deviations from Hardy–Weinberg equilibrium (HWE) and linkage equilibrium to assign genetically similar individuals into genetic clusters (Porras-Hurtado et al. 2013). In the absence of a priori spatial data, genetic assignment results from STRUCTURE are independent of geographic sample locations, which allows genetic assignments to be used as an unbiased indicator of spatial patterns of genetic clusters. To assess trends in the likelihood as a function of the number of genetic clusters (K) specified, we ran five independent runs for a range of K = 1–10 at 5,000 burn-in and 15,000 Markov chain Monte Carlo (MCMC) repetitions with an admixture model and assuming correlated allele frequencies (Benestan et al. 2016). We used STRUCTURE Harvester (v. 6.92) to graph likelihood as a function of K (Earl 2019). We then performed one iteration for each level of K at 500,000 burn-in and 1,000,000 MCMC reps (Gilbert et al. 2012). We used STRUCTURE PLOT (v. 2) (Ramasamy et al. 2014) to generate bar plots for selected K values. We next used the R package adegenet (Jombart et al. 2020) to perform a discriminant analysis of principal components (DAPC) to visualize group assignments identified in STRUCTURE in terms of genetic distance (Miller et al. 2020).
Genetic variation
After populations were delineated we estimated genetic summary statistics in the R package diveRsity (Keenan et al. 2013). We tested for departures from HWE using 999 iterations at each locus in each population and estimated the probability of any two unique genotypes matching using GenAlEx 6.5 (Peakall and Smouse 2012). We estimated both the probability of identity (PID; the probability of two randomly selected individuals sharing the same genotype) and the probability of identity of siblings (PSIB; the probability that two siblings share the same genotype) (Waits et al. 2001) using GenAlEx 6.5.
Contemporary genetic effective population size and gene flow
We estimated the contemporary genetic effective population size (Ne) for each genetic cluster (including both sexes) using the NeEstimator (v2.1) (Do et al. 2014) software executed with the RLDNe package in the R environment (https://rdrr.io/github/zakrobinson/RLDNe/). NeEstimator uses a bias correction of the linkage disequilibrium method (Waples 2006). We excluded alleles with a frequency ≤ 0.02 and generated 95% confidence intervals via jackknifing (e.g., Buchalski et al. 2015). We estimated the expected loss of heterozygosity (He) over a single generation of a randomly mating, isolated population of size Ne with the equation He(t1) = 1–He(t0) × 1/(2Ne) (Frankham et al. 2017).
To estimate contemporary gene flow among populations, we used a Bayesian assignment method employed in program BayesAss (v. 3.04; Wilson and Rannala 2003). Based on preliminary test runs, we specified the delta values for migration rates, allele frequencies, and inbreeding coefficients (female: 0.2, 0.3, and 0.3; male: 0.2, 0.3, and 0.4) to reach the recommended acceptance rates. We ran the program with 107 MCMC steps, discarding the first 106 as burn-in, and sampling every 1,000 steps. To assess the consistency of results we performed 3 independent runs with different random number seeds. We used the program Tracer (v1.5) (Rambaut et al. 2018) to assess convergence and to retrieve effective sample size (ESS) values. To express gene flow in terms of migrants per generation (Nm) separately for each sex, we used the equation Nei × Mj→i to infer Nm, where Nei equals the effective population size of population i, estimated above in NeEstimator v2.1, and Mj→i equals the pairwise migration rate from population j into population i, estimated in BayesAss. We considered Mj→i–(1.96 × SD) > 0 to indicate statistically significant gene flow (Wilson and Rannala 2003).
Results
We collected tissue samples from 143 elk along with 1,616 pellet groups across three field seasons. After eliminating genotypes with < 18 of 20 loci, 1145 sample genotypes remained for analysis, from which we identified 490 unique individual elk (257 females, 233 males; Fig. 1a, b; Online resource 1).
Genetic structure
Both males and females exhibited similar patterns of population structure (Fig. 2). Although likelihood increased substantially with increasing numbers of clusters specified up to K = 4 and then gradually from K = 5 to 8, little information was gained for K > 6 (Online resource 2: Table S1, Fig. S1). The structure evident at K = 3–6 clusters was hierarchical (Online resource 2: Fig. S1A, B). At K = 3, three of the original putative populations were differentiated, with PV and LPB clustering together. At K = 4–5, CC split into multiple clusters (Online resource 1: Fig. S1). Division into K = 5 clusters introduced a “ghost cluster” within CC, effectively adding no new information over K = 4. At K = 6, PV emerged as distinct from LPB. Both inter- and intra-population differentiation was geographically evident (Fig. 2a–d). Based on these patterns, we hereafter considered five populations or subpopulations (collectively, “population units”) corresponding to the K = 6 level (i.e., including 1 ghost cluster) for subsequent analyses: PV, LPB, EPR, CC-Rock Quarry (CCRQ), and CC-Cortina Ridge (CCCR). Although bar charts indicated movement of individuals between the two CC subpopulations, none of the 490 unique individuals assigned primarily to another of the primary populations as would be expected for a first-generation migrant (or disperser). One individual (a male) in LPB had approximately half its ancestry assigned to CCRQ, consistent with being progeny of a first-generation migrant. Additionally, CCCR contained several individuals of both sexes with ancestry assignments consistent with second- or third-generation ancestry from the other three primary populations. However, it was unclear whether these admixed individuals in CCCR reflected actual gene flow from the populations under study or artifacts of having been introduced during augmentation events from the same or related source populations.
Sampling locations (n = 490) color coded according to genetic assignment in STRUCTURE in 1 of 4 distinct clusters for a female (n = 257) and b male (n = 233) tule elk in the Potter Valley (PV), Lake Pillsbury (LPB), East Park Reservoir (EPR), Cache Creek-Rock Quarry (CCRQ), and Cache Creek-Cortina Ridge (CCCR) population units. The same sampling locations are shown color coded according to genetic assignment in STRUCTURE in 1 of 6 distinct clusters for c females and d males. Colored circles indicate individuals with ≥ 80% of their ancestry estimated to be from a single genetic cluster; open circles indicate admixed individuals (those with < 80% ancestry across all 4 and/or 6 genetic clusters). Corresponding STRUCTURE bar plots are found under each respective map, where each vertical bar represents an individual elk and the colors within each bar represent proportions of ancestry assigned to different genetic clusters. Vertical dashed lines indicate geographic sample locations along a west-to-east gradient associated with each population unit
The DAPC analysis yielded complementary results to the STRUCTURE analyses for the five population units. Female population units were relatively isolated, except for the two CC units (Fig. 3a). In contrast to females, however, four of five male population units overlapped to some degree, but also with the greatest overlap between the two CC units (Fig. 3b). Only the EPR unit did not overlap with any group in either sex. The proportion of assignments to the population unit in which individuals were sampled ranged 0.71–1 for females and 0.63–1 for males (Table 1).
Discriminant analysis of principal components (DAPC) clusters for tule elk pre-assigned to geographic population units (colored orbs) including a females (n = 257) and b males (n = 233) within the Potter Valley (PV), Lake Pillsbury (LPB), East Park Reservoir (EPR), Cache Creek-Rock Quarry (CCRQ), and Cache Creek-Cortina Ridge (CCCR) population units
Genetic diversity
The five population units followed similar patterns of relatively low levels of genetic variation (Table 2). All loci were polymorphic, with the exception of one locus (TE68) that was monomorphic among PV males (Online resource 2: Table S2). None of the FIS values differed significantly from zero, indicating no evidence of additional substructure within the five population units (Table 2).
Effective population size (N e )
Estimates of Ne were 51.3 (95% CI 33.3–68.5) for LPB, 25.5 (16–45.9) for CCRQ, and 15.2 (10.2–21.5) for CCCR (Table 2). Small sample sizes (< 50) resulted in Ne with upper bounds indistinguishable from infinity for PV [75.5 (13.5–Inf)] and EPR [332.7 (46.0–Inf)]. Because Ne for the EPR population unit exceeded the most recent abundance estimate (Bush et al. 2020), we presumed it biased and excluded it from further analyses. The expected loss of heterozygosity over a single generation for randomly mating populations of the sizes of the other populations were 0.66% (PV), 0.97% (LPB), 1.9% (CCRQ), and 3.3% (CCCR).
Contemporary gene flow
Analyses in BayesAss indicated that both sexes tended to exhibit relatively low migration rates among the four populations, although the two CC subpopulations exchanged higher rates of migration (Table 3). Females exhibited bidirectional gene flow between CCRQ and CCCR (M = 7.2%, 7.8%). In contrast, male gene flow was primarily unidirectional from CCCR to CCRQ (M = 7.2%). We also detected significant unidirectional gene flow from LPB to PV (M = 12.3%), EPR to CCRQ (M = 3.0%), and EPR to CCCR (M = 2.6%) in males. Lastly, we detected no significant gene flow into the LPB and EPR population units for either sex.
Statistically significant estimated numbers of migrants per generation (Nm) ranged up to 1.9 for females and 9.3 for males (Table 4). Female gene flow between CCRQ and CCCR was bidirectional, Nm = 1.9 and Nm = 1.2 (Fig. 4a). Male gene flow was primarily unidirectional from LPB to PV (Nm = 9.3) and from CCCR to CCRQ (Nm = 1.9; Fig. 4b). Low levels of male gene flow (Nm < 1) also potentially connected EPR to both CCRQ and CCCR. Except for male gene flow from LPB, other potentially meaningful levels (Nm > 1) of male and female gene flow into PV were not statistically differentiable from zero.
Schematic diagram showing meaningful (> 1) contemporary migrants per generation (Nm) among five population units, Potter Valley (PV), Lake Pillsbury (LPB), East Park Reservoir (EPR), Cache Creek-Rock Quarry (CCRQ), and Cache Creek-Cortina Ridge (CCCR), of a female and b male tule elk where significant gene flow was detected in BayesAss (Table 3). Nm across pairwise populations are indicated next to the arrow indicating movement. We used the equation Nei × Mj→i to infer Nm, where Nei equals the effective population size of population i, estimated in NeEstimator v2.1, and Mj→i equals the effective immigration rate from population j into population i estimated in BayesAss. Dashed arrows indicate statistically significant gene flow that is Nm < 1 (b: EPR to CCRQ, EPR to CCCR) or that are not statistically significant but for which the point estimates of Nm > 1 (a, b: all other dashed arrows)
Discussion
At the outset of our study, spatially disparate populations with distinct reintroduction histories were thought to be completely isolated from one another with two exceptions: LPB and PV, and the two subpopulations of CC, in both cases known to have shared common origins. As anticipated, each population was associated with its historical release site or a founding population known to have dispersed from one of those release sites (CDFW 2018). The genetic diversity, including Ne, also was generally low, presumably due at least in part to the historical population bottleneck of the 1800s and their polygynous mating system. Below, we expand on these findings in the context of population histories, experiences with other elk populations, and implications for conservation.
Demographic history and population structure
Microsatellite genotypes clustered tule elk into four major genetic populations corresponding to the locations of their initial establishment. Among these, only the CC population exhibited substructure. Possible explanations for the maintenance of genetic distance observed among populations include geographic or anthropogenic barriers and social cohesion of individuals that were introduced together. Although the landscape was highly heterogeneous with respect to settlement habitat—valleys, riparian, and lacustrine areas of grassland and low canopy cover (Batter 2020)—we doubt that habitats poor for elk settlement (e.g., dense-canopy forest, chaparral-covered peaks) necessarily posed significant dispersal barriers or alone could explain the observed distinctiveness of populations (Hilty et al. 2012; Zecherle et al. 2020). Post-release movements of elk suggest that poor settlement habitat does not necessarily present a barrier to movement. For example, the PV population was established from “dispersers” (soon after introduction) from LPB, between which the landscape is primarily characterized by poor settlement habitat. Additionally, founders of the EPR population ostensibly originated from an introduction to a site (Bartlett Springs/Potato Hill) surrounded by inhospitable habitat that they would have to have traversed (CDFW 2018; Batter 2020). Although it is possible that human or livestock density between some of these populations discouraged regular movement, it seems likely that the genetic isolation among populations stems primarily from non-landscape-related factors.
In particular, social cohesion related to the legacy of past reintroductions could be the principle driver of population structure in our study region, and perhaps among reintroduced elk populations, generally. In a study of reintroduced elk in the eastern United States, both male and female elk introduced from multiple sources to the same site segregated into breeding groups corresponding to their source populations for at least two generations after reintroduction (Muller et al. 2018). This example contrasts with a natural population of elk in Idaho found to experience substantial gene flow among patchily distributed populations (Aycrigg and Garton 2014), suggesting that under natural circumstances or once populations have achieved some critical threshold in size, elk exchange genes more fluidly across the landscape. The CC population in our study, which had been reintroduced a century earlier, demonstrated aspects of both of these extremes. As with more newly reintroduced populations, both in our study area and those of the eastern United States, the CC population was relatively isolated from other populations. More in line with the natural populations in Idaho, however, the CC population exhibited substructure, whereby each subpopulation was genetically diagnosable, yet connected through regular gene flow. These patterns suggest that elk may tend to remain within their social groups, emigrating from an area only when abundance exceeds carrying capacity and necessitates it, and then moving only as far as necessary to settle in suitable habitat. Additionally, social memory may facilitate gene flow between subpopulations established this way, in contrast to those established from independent sources. Such a model could explain the observed lack of connectivity among the four primary populations in our study as a result of both low abundance relative to carrying capacity of the broader region and social independence among some of the reintroduced populations. A prediction of this model is that social inertia and small population size must be overcome before reintroduced elk populations can develop conservation-independent connectivity.
Elk also may respond differently depending on the local conditions of reintroduction sites. Elk introduced to our study area at different periods during the twentieth century apparently responded in one of two ways: either they remained where they were originally introduced, expanding slowly, or they left the reintroduction site, establishing elsewhere (Batter 2020; CDFW 2018). These different responses seem to correspond to differences in local carrying capacities. The two reintroduction sites in our study area that experienced immediate exodus and exploratory dispersal by reintroduced individuals, Bartlett Springs/Potato Hill and LPB, involved relatively small areas of grassland surrounded by large expanses of poor tule elk habitat (primarily closed-canopy coniferous forests). In contrast, the CC site occurred within a large mosaic of interconnected riparian, grassland, and oak woodland habitats, allowing population expansion from CCCR into suitable habitat to the west, establishing the CCRQ, without requiring long-distance dispersal (CDFW 2018; McCullough et al. 1996). The resulting proximity, along with social memory, could help to explain ongoing gene flow between these subpopulations. Likewise, the two populations that were established “naturally” by elk themselves (PV, EPR) occurred in similar expanses of suitable habitat suggesting that landscape-level habitat selection favors a stepping-stone configuration of interconnected habitat patches, which also facilitates genetic connectivity as populations grow and expand.
Genetic diversity and effective population size
As expected, we found low overall levels of genetic diversity and small effective population sizes within the five population units (McCullough et al. 1996; Meredith et al. 2007; Sacks et al. 2016). As a general guideline, the 50/500 rule suggests an effective population size of Ne > 50 is needed to avoid the risk of inbreeding depression, whereas Ne > 500 is needed to keep a population from losing adaptive potential due to genetic drift (Franklin 1980). This rule was later revised upward (the 100/1000 rule) (Frankham et al. 2014). More specifically, simulations aimed at addressing loss of diversity in tule elk populations indicate that up to 44% of allelic diversity and 95% of heterozygosity could be lost in 25 generations in an isolated population with Ne = 20 (Williams et al. 2004). Our estimates of contemporary Ne for the focal tule elk population units were near or below these lower thresholds and well below the higher thresholds. These results are not surprising given the expectations of a polygynous mating system and variation in reproductive success (Waples et al. 2016), where only about 17% of mature males successfully breed with roughly 90% of mature females (Johnson et al. 2007; McCullough et al. 1996). Ultimately, however, the implications of the effective population sizes of the population units we studied and other tule elk populations depends on their growth rate and connectivity.
Gene flow
Because elk persist in a matrilineal social structure with a polygynous mating system (Muller et al. 2018; Nussey et al. 2005; Raedeke et al. 2002; Smith and Anderson 2001), we expected to detect more gene flow among male than female elk. Although comparison of corresponding female and male BayesAss estimates of gene flow did not consistently indicate higher gene flow among males than females, this may have partly reflected the biparental inheritance of markers, which are expected to reflect similarly in males and females after the first generation. In contrast, however, the DAPC analysis generally supported this expectation. In particular, the DAPC analysis in females showed clear separation of the four clusters corresponding to the four primary populations, whereas the same analysis of males only resolved 3 clusters, with the PV samples overlapping LPB and CC clusters. On the whole, gene flow was generally low in both sexes. Although our estimate of gene flow in males from LPB to PV was very high (nearly 10 migrants per generation), this estimate may partly reflect the founding of PV by LPB individuals several generations ago. Indeed, the STRUCTURE analysis at K = 6 provided no evidence of recent (e.g., second-generation) gene flow in either sex. We also found no evidence of gene flow into EPR for either sex.
From the perspective of maintaining genetic diversity, gene flow from either sex benefits both sexes. A heuristic rule is that gene flow in excess of one migrant per generation may be sufficient to counter genetic drift and inbreeding within small, isolated population units (Mills and Allendorf 1996; Wright 1931). Our estimates of Nm suggest that all populations could be susceptible to negative genetic consequences without substantial population growth or external gene flow. Although we observed substantial gene flow within CC (i.e., between subpopulations), the combined Ne was nevertheless small (Ne ~ 40). Because we lack baseline data for the historical diversity in this population, it is impossible to assess whether its Ne has declined or increased in the century since its establishment. However, this population has grown demographically in relative isolation, with a larger current census population size (N ~ 400) than any of the other populations (N < 200) (Batter 2020; Bush et al. 2020; Moran et al. 2020). While such resilience in the short-term appears to be a common feature among fragmented tule elk populations, it does not necessarily assure protection against future deleterious genetic effects (CDFW 2018; McCullough et al. 1996; Williams et al. 2004).
Conservation implications
Elk conservation in California has been successful on the whole, but in the future could benefit from regular genetic monitoring and a systematic genetic management plan that incorporates goals and assessments. The relatively large increase in the range-wide abundance of tule elk over the past century indicates that reintroductions have been successful in growing numbers and, therefore, stemming the loss of genetic diversity at the level of the subspecies. Indeed, the human-assisted recovery of the subspecies from as few as 2 to > 6000 individuals marks a major conservation success.
However, presumptive fragmentation among reestablished populations (the motivation for this study) continues to pose concerns over the genetic health at the local level (Williams et al. 2004; CDFW 2018), which has prompted the practice of using human-mediated augmentations to artificially impose gene flow among reintroduced populations with the aim of maintaining or increasing their genetic diversity. While our findings in this study support the broad assumptions that have motivated this basic management strategy (CDFW 2018)—low genetic effective populations sizes and gene flow—the success or necessity of particular augmentations remains unclear.
Because augmentations are typically opportunistic rather than components of a systematic genetic management plan that includes genetic monitoring, few data exist with which to evaluate whether translocated individuals successfully breed and integrate genetically into recipient populations (CDFW 2018). Based on other systems, successful integration of translocated individuals into restored populations can vary substantially and be influenced by demographic, behavioral, and environmental conditions, numbers and sex of individuals released, and even the pre-release circumstances of source populations, such as whether captive or free-ranging and the genetic diversity of source stocks (Flesch et al. 2020; Mertes et al. 2019; Ralls et al. 2018; Renan et al. 2018; Youngmann et al. 2020; Zecherle et al. 2020). Future augmentations should therefore incorporate post-release demographic and genetic monitoring programs, including telemetry of reintroduced individuals and pedigree reconstruction to evaluate social integration and reproductive success of released individuals.
Also unknown is the extent to which successful introductions (i.e., resulting in genetic integration) have influenced Ne or when Ne would have continued to grow without augmentation. In the future, monitoring trends in Ne could provide valuable data to inform management. For example, populations shown to have declining Ne would most warrant augmentations, whereas those for which Ne was growing could continue to be monitored without augmentation. In populations receiving translocations, monitoring of Ne along with reproductive success of introduced individuals would inform on the efficacy of augmentation efforts and illuminate factors that may affect its success (Williams et al. 2004). Importantly, genetic monitoring could help to identify populations for which Ne grows sufficiently or that expand connectivity with other populations to the point of conservation-independence, which, where feasible, should be the ultimate goal of genetic management. One final attribute in need of monitoring is potential admixture among subspecies, particularly in locations where tule elk come into close proximity with Roosevelt (C. c. roosevelti) or Rocky Mountain elk populations. Admixture among subspecies can potentially have positive and negative effects on fitness leading to a variety of conservation implications. Admixture also may have social and economic impacts on stakeholders, such as hunters or communities relying on hunting revenues, as records would be invalidated by subspecific admixture.
Conclusions
Our findings underscore the importance of obtaining baseline genetic data for tule elk populations and point to the need to do so for other fragmented elk populations throughout California. By identifying distinct genetic clusters and quantifying both genetic effective population size and gene flow among them in the present study, we were able to provisionally assess whether natural processes were sufficient to maintain existing genetic diversity and counter inbreeding and loss of genetic diversity (Allendorf and Luikart 2009; Frankham et al. 2017). The answer was a tentative ‘no,’ with the qualifier that given sufficient time and ability to expand naturally, populations could potentially escape the need for conservation-dependence. Thus, monitoring of both demographic and genetic parameters in the context of experimentation is essential toward developing a strategy to achieve self-sustaining populations.
Data availability
All data generated or analysed during this study are included in this published article (and its supplementary information files).
References
Allendorf FW, Luikart G (2009) Conservation and the genetics of populations. Wiley, Hoboken
Aycrigg JL, Garton EO (2014) Linking metapopulation structure to elk population management in Idaho: a genetic approach. J Mammal 95:597–614
Batter TJ (2020) Development and implementation of DNA-based survey methods for population monitoring of Tule Elk (Cervus canadensis nannodes) in the interior coast ranges of Northern California. Dissertation. University of California, Davis
Benestan LM et al (2016) Conservation genomics of natural and managed populations: building a conceptual and practical framework. Mol Ecol 25:2967–2977
Buchalski MR et al (2015) Genetic population structure of Peninsular bighorn sheep (Ovis canadensis nelsoni) indicates substantial gene flow across US–Mexico border. Biol Conserv 184:218–228
Bush J, Batter T, Landers R, Denryter K (2020) Report on aerial surveys of tule elk (Cervus canadensis nannodes) in 2018-2019 in Bear Valley, Cache Creek, East Park Reservoir, and Lake Pillsbury tule elk hunt zones. Internal report. California Department of Fish and Wildlife, Sacramento, CA. Sacramento, California, USA. https://doi.org/10.13140/RG.2.2.11670.98888
CDFW (2018) Conservation and management plan for elk. California Department of Fish and Wildlife, Sacramento
Ciriacy-Wantrup S, Phillips WE (1970) Conservation of the California Tule Elk: a socioeconomic study of a survival problem. Biol Cons 3:23–32
Do C, Waples RS, Peel D, Macbeth G, Tillett BJ, Ovenden JR (2014) NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour 14:209–214
Earl D (2019) Structure Harvester vO. 56.4
Flesch EP, Graves TA, Thomson JM, Proffitt KM, White P, Stephenson TR, Garrott RA (2020) Evaluating wildlife translocations using genomics: a bighorn sheep case study. Ecol Evol 10:13687–13704
Frankham R (2015) Genetic rescue of small inbred populations: Meta-analysis reveals large and consistent benefits of gene flow. Mol Ecol 24:2610–2618
Frankham R, Bradshaw CJ, Brook BW (2014) Genetics in conservation management: revised recommendations for the 50/500 rules, Red List criteria and population viability analyses. Biol Conserv 170:56–63
Frankham R et al (2017) Genetic management of fragmented animal and plant populations. Oxford University Press, Oxford
Frankham R et al (2019) A practical guide for genetic management of fragmented animal and plant populations. Oxford University Press, Oxford
Franklin IR (1980) Evolutionary change in small populations. In: Soule ME and Wilcox BA (eds) Conservation biology: an evolutionary-ecological perspective. Sinauer Associates, Sunderland, MA, pp 135–140
Galpern P, Manseau M, Hettinga P, Smith K, Wilson P (2012) Allelematch: an R package for identifying unique multilocus genotypes where genotyping error and missing data may be present. Mol Ecol Resour 12:771–778
Gilbert KJ et al (2012) Recommendations for utilizing and reporting population genetic analyses: the reproducibility of genetic clustering using the program STRUCTURE. Mol Ecol 21:4925–4930
Hilty JA, Lidicker WZ Jr, Merenlender AM (2012) Corridor ecology: the science and practice of linking landscapes for biodiversity conservation. Island Press, Washington
Johnson HE, Bleich VC, Krausman PR, Koprowski JL (2007) Effects of antler breakage on mating behavior in male tule elk (Cervus elaphus nannodes). Eur J Wildl Res 53:9–15
Jombart T et al. (2020) Adegenet: exploratory analysis of genetic and genomic data
Jones KC, Levine KF, Banks JD (2002) Characterization of 11 polymorphic tetranucleotide microsatellites for forensic applications in California elk (Cervus elaphus canadensis). Mol Ecol Notes 2:425–427
Kauffman E (2003) Atlas of the biodiversity of California. Climate and topography. California Department of Fish and Game, Sacramento, pp 12–15
Keenan K, McGinnity P, Cross T, Crozier W, Prodöhl P (2013) DiveRsity: an R package for the estimation of population genetics parameters and their associated errors. Methods Ecol Evol. https://doi.org/10.1111/2041-210X.12067
Kucera T (1991) GENETIC-VARIABILITY IN TULE ELK. Calif Fish Game 77:70–78
Lowe WH, Allendorf FW (2010) What can genetics tell us about population connectivity? Mol Ecol 19:3038–3051
McCullough DR (1969) The tule elk: its history, behavior, and ecology. University of California Publications in Zoology 88: 1–209
McCullough DR, Fischer JK, Ballou JD (1996) From bottleneck to metapopulation: recovery of the tule elk in California. In: McCullough DR (ed) Metapopulations and wildlife conservation. Island Press, Washington, pp 375–403
Meredith E, Rodzen J, Banks J, Schaefer R, Ernest HB, Famula T, May B (2007) Microsatellite analysis of three subspecies of elk (Cervus elaphus) in California. J Mammal 88:801–808
Mertes K et al (2019) Management background and release conditions structure post-release movements in reintroduced ungulates. Front Ecol Evol 7:470
Miller JM, Cullingham CI, Peery RM (2020) The influence of a priori grouping on inference of genetic clusters: simulation study and literature review of the DAPC method. Heredity 125:269–280
Mills LS, Allendorf FW (1996) The one-migrant-per-generation rule in conservation and management. Conserv Biol 10:1509–1518
Moran A, Morefield K, Denryter K (2020) Report on spring aerial surveys of tule elk (Cervus canadensis nannodes) in the Mendocino Elk Management Unit Internal report California Department of Fish and Wildlife, Sacramento, CA Sacramento, California, USA. https://doi.org/10.13140/RG.2.2.15269.22243
Muller LI et al (2018) Genetic structure in elk persists after translocation. J Wildl Manag 82:1124–1134
Nussey D et al (2005) Rapidly declining fine-scale spatial genetic structure in female red deer. Mol Ecol 14:3395–3405
Peakall R, Smouse P (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Porras-Hurtado L, Ruiz Y, Santos C, Phillips C, Carracedo Á, Lareu M (2013) An overview of STRUCTURE: applications, parameter settings, and supporting software. Front Genet 4:98
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raedeke KJ, JJ M, PE C (2002) Population characteristics. In: DE T, JW T (eds) North American elk: ecology and management. Smithsonian Institution Press, Washington, pp 449–491
Ralls K et al (2018) Call for a paradigm shift in the genetic management of fragmented populations. Conserv Lett 11:e12412
Ramasamy RK, Ramasamy S, Bindroo BB, Naik VG (2014) STRUCTURE PLOT: a program for drawing elegant STRUCTURE bar plots in user friendly interface. SpringerPlus 3:1–3
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67:901
Renan S et al (2018) Fission-fusion social structure of a reintroduced ungulate: implications for conservation. Biol Cons 222:261–267
Sacks BN, Lounsberry ZT, Kalani T, Meredith EP, Langner C (2016) Development and characterization of 15 polymorphic dinucleotide microsatellite markers for tule elk using HiSeq3000. J Hered 107:666–669
Smith BL, Anderson SH (2001) Does dispersal help regulate the Jackson elk herd? Wildl Soc Bull 29:331–341
Toonen RJ, Hughes S (2001) Increased throughput for fragment analysis on an ABI Prism® 377 automated sequencer using a membrane comb and STRand software. Biotechniques 31:1320–1325
Waits LP, Luikart G, Taberlet P (2001) Estimating the probability of identity among genotypes in natural populations: cautions and guidelines. Mol Ecol 10:249–256
Waples RS (2006) A bias correction for estimates of effective population size based on linkage disequilibrium at unlinked gene loci. Conserv Genet 7:167
Waples RK, Larson WA, Waples RS (2016) Estimating contemporary effective population size in non-model species using linkage disequilibrium across thousands of loci. Heredity 117:233–240
Whiteley AR, Fitzpatrick SW, Funk WC, Tallmon DA (2015) Genetic rescue to the rescue. Trends Ecol Evolut 30:42–49
Williams CL, Lundrigan B, Rhodes OE Jr (2004) Microsatellite DNA variation in tule elk. J Wildl Manag 68:109–119
Wilson GA, Rannala B (2003) Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163:1177–1191
Wright S (1931) Evolution in Mendelian populations. Genetics 16:97
Youngmann JL, Deyoung RW, Demarais S, Strickland BK, Jenkins G (2020) Genetic characteristics of restored elk populations in Kentucky. J Wildl Manag 84:515–523
Zecherle L, Bar-David S, Nichols H, Templeton A, Hipperson H, Horsburgh G, Brown R (2020) Landscape resistance affects individual habitat selection but not genetic relatedness in a reintroduced desert ungulate. Biol Cons 252:108845
Acknowledgements
Funding was from the California Department of Fish and Wildlife (CDFW), Big Game Management Account through cooperative agreement with the University of California, Davis (P1680034), supplemented through the Mammalian Ecology and Conservation Unit of the Veterinary Genetics Laboratory at UC Davis. C. Sanchez, K. Barnitz, A. Hemphill, C. White, and J. Owen assisted with field work. S. Vanderzwan provided laboratory oversight, training, and assistance with laboratory analyses. J.M. Statham assisted with data analysis. We thank the U.S. Bureau of Land Management, U.S.Forest Service, U.S. Bureau of Reclamation, and private land owners who provided access to property to conduct research activities. We thank K. Denryter, N. Galloway, and an anonymous reviewer for helpful comments on an earlier draft of this manuscript.
Funding
California Department of Fish and Wildlife, Agreement No. P1680034.
Author information
Authors and Affiliations
Contributions
BNS and TJB designed the study, TJB and JPB collected data, TJB and BNS analyzed data, TJB, BNS, and JPB wrote manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts.
Ethical approval
Methods approved by CDFW Wildlife Investigations Laboratory, in accordance with CDFW Animal Welfare Policy.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Batter, T.J., Bush, J.P. & Sacks, B.N. Assessing genetic diversity and connectivity in a tule elk (Cervus canadensis nannodes) metapopulation in Northern California. Conserv Genet 22, 889–901 (2021). https://doi.org/10.1007/s10592-021-01371-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-021-01371-0