Abstract
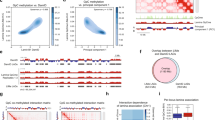
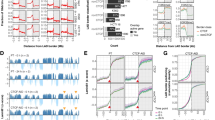
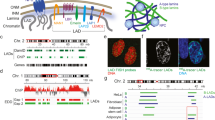
More than one third of the mammalian genome is in a close association with the nuclear lamina, thus these genomic regions were termed lamina-associated domains (LADs). This association is fundamental for many aspects of chromatin biology including transcription, replication, and DNA damage repair. LADs association with the nuclear envelope is thought to be dependent on two major mechanisms: The first mechanism is the interaction between nuclear membrane proteins such as LBR with heterochromatin modifications that are enriched in LADs chromatin. The second mechanism is based on proteins that bind the borders of the LADs and support the association of the LADs with the nuclear envelope. Two factors were suggested to support the second mechanism: CCCTC-binding factor (CTCF) and YY1 based on their enriched binding to LADs borders. However, this mechanism has not been proven yet at a whole genome level. Here, to test if CTCF supports the LADs landscape, we generated melanoma cells with a partial loss of function (pLoF) of CTCF by the CRISPR-Cas9 system and determined the LADs landscape by lamin B ChIP-seq analysis. We found that under regular growth conditions, CTCF pLoF led to modest changes in the LADs landscape that included an increase in the signal of 2% of the LADs and a decrease in the signal of 8% of the LADs. However, CTCF importance for the LADs landscape was much higher upon induction of a chromatin stress. We induced chromatin stress by inhibiting RNA polymerase II, an intervention that is known to alter chromatin compaction and supercoiling. Notably, only in CTCF pLoF cells, the chromatin stress led to the dissociation of 7% of the LADs from the lamina. The CTCF-dependent LADs had almost three times shorter median length than the non-affected LADs, were enriched in CTCF binding at their borders, and were higher in their facultative-status (cell-type specific). Thus, it appears that CTCF is a key factor in facilitating the association of short facultative LADs with the nuclear lamina upon chromatin stress.





Similar content being viewed by others
Abbreviations
- AUGi:
-
Initiation codon
- CRISPR:
-
Clustered regularly interspaced short palindromic repeats
- CTCF:
-
CCCTC-binding factor
- DamID:
-
DNA adenine methyltransferase identification
- DRB:
-
5,6-Dichlorobenzimidazole 1-β-D-ribofuranoside
- INM:
-
Inner nuclear membrane
- LAD:
-
Lamina-associated domain
- LBR:
-
Lamin B receptor
- NPC:
-
Nuclear pore complex
- ONM:
-
Outer nuclear membrane
- pLoF:
-
Partial loss of function
- TAD:
-
Topologically associated domain
References
Andreu MJ, Alvarez-Franco A, Portela M, Gimenez-Llorente D, Cuadrado A, Badia-Careaga C, Tiana M, Losada A, Manzanares M (2021) Establishment of 3D chromatin structure after fertilization and the metabolic switch at the morula-to-blastocyst transition require CTCF. bioRxiv 2021.07.07.451492. https://doi.org/10.1101/2021.07.07.451492
Bian Q, Khanna N, Alvikas J, Belmont AS (2013) β-Globin cis-elements determine differential nuclear targeting through epigenetic modifications. J Cell Biol 203:767–783. https://doi.org/10.1083/jcb.201305027
Bitman-Lotan E, Orian A (2021) Nuclear organization and regulation of the differentiated state. Cell Mol Life Sci 78:3141–3158. https://doi.org/10.1007/s00018-020-03731-4
Braccioli L, de Wit E (2019) CTCF: a Swiss-army knife for genome organization and transcription regulation. Essays Biochem 63:157–165. https://doi.org/10.1042/EBC20180069
Briand N, Collas P (2020) Lamina-associated domains: peripheral matters and internal affairs. Genome Biol 21:85. https://doi.org/10.1186/s13059-020-02003-5
Brunet A, Forsberg F, Fan Q et al (2019) Nuclear lamin b1 interactions with chromatin during the circadian cycle are uncoupled from periodic gene expression. Front Genet 10:917. https://doi.org/10.3389/fgene.2019.00917
Busslinger GA, Stocsits RR, van der Lelij P et al (2017) Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl. Nature 544:503–507. https://doi.org/10.1038/nature22063
Chen C-K, Blanco M, Jackson C et al (2016) Xist recruits the X chromosome to the nuclear lamina to enable chromosome-wide silencing. Science 354:468–472. https://doi.org/10.1126/science.aae0047
Chen S, Luperchio TR, Wong X et al (2018) A lamina-associated domain border governs nuclear lamina interactions, transcription, and recombination of the Tcrb locus. Cell Rep 25:1729-1740.e6. https://doi.org/10.1016/j.celrep.2018.10.052
Cong L, Ran FA, Cox D et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823. https://doi.org/10.1126/science.1231143
Czapiewski R, Robson MI, Schirmer EC (2016) Anchoring a Leviathan: how the nuclear membrane tethers the genome. Front Genet 7. https://doi.org/10.3389/fgene.2016.00082
Davidson IF, Goetz D, Zaczek MP, et al (2016) Rapid movement and transcriptional re‐localization of human cohesin on DNA. EMBO J 35:2671–2685. https://doi.org/10.15252/embj.201695402
Donnaloja F, Carnevali F, Jacchetti E, Raimondi MT (2020) Lamin a/c Mechanotransduction in Laminopathies. Cells 9:1306. https://doi.org/10.3390/cells9051306
Dunlevy KL, Medvedeva V, Wilson JE, et al (2020) The PRR14 heterochromatin tether encodes modular domains that mediate and regulate nuclear lamina targeting. J Cell Sci jcs.240416. https://doi.org/10.1242/jcs.240416
Egan B, Yuan C-C, Craske ML et al (2016) An alternative approach to ChIP-Seq normalization enables detection of genome-wide changes in histone H3 lysine 27 trimethylation upon EZH2 inhibition. PLoS ONE 11:e0166438. https://doi.org/10.1371/journal.pone.0166438
Fedoriw AM (2004) Transgenic RNAi reveals essential function for CTCF in H19 gene imprinting. Science 303:238–240. https://doi.org/10.1126/science.1090934
Fracchia A, Asraf T, Salmon-Divon M, Gerlitz G (2020) Increased lamin B1 levels promote cell migration by altering perinuclear actin organization. Cells 9:2161. https://doi.org/10.3390/cells9102161
Gerlitz G, Bustin M (2010) Efficient cell migration requires global chromatin condensation. J Cell Sci 123:2207–2217. https://doi.org/10.1242/jcs.058271
Ghirlando R, Felsenfeld G (2016) CTCF: making the right connections. Genes Dev 30:881–891. https://doi.org/10.1101/gad.277863.116
González-Buendía E, Pérez-Molina R, Ayala-Ortega E et al (2014) Experimental strategies to manipulate the cellular levels of the multifunctional factor CTCF. In: Robles-Flores M (ed) Cancer Cell Signaling. Springer, New York, pp 53–69
Gonzalez-Sandoval A, Towbin BD, Kalck V et al (2015) Perinuclear anchoring of H3K9-methylated chromatin stabilizes induced cell fate in C. elegans embryos. Cell 163:1333–1347. https://doi.org/10.1016/j.cell.2015.10.066
Gruenbaum Y, Foisner R (2015) Lamins: nuclear intermediate filament proteins with fundamental functions in nuclear mechanics and genome regulation. Annu Rev Biochem 84:131–164. https://doi.org/10.1146/annurev-biochem-060614-034115
Guerreiro I, Kind J (2019) Spatial chromatin organization and gene regulation at the nuclear lamina. Curr Opin Genet Dev 55:19–25. https://doi.org/10.1016/j.gde.2019.04.008
Handoko L, Xu H, Li G et al (2011) CTCF-mediated functional chromatin interactome in pluripotent cells. Nat Genet 43:630–638. https://doi.org/10.1038/ng.857
Harr JC, Luperchio TR, Wong X et al (2015) Directed targeting of chromatin to the nuclear lamina is mediated by chromatin state and A-type lamins. J Cell Biol 208:33–52. https://doi.org/10.1083/jcb.201405110
Hetzer MW (2010) The nuclear envelope. Cold Spring Harb Perspect Biol 2:a000539. https://doi.org/10.1101/cshperspect.a000539
Ho CY, Lammerding J (2012) Lamins at a glance. J Cell Sci 125:2087–2093. https://doi.org/10.1242/jcs.087288
Hsieh T-HS, Cattoglio C, Slobodyanyuk E et al (2020) Resolving the 3D landscape of transcription-linked mammalian chromatin folding. Mol Cell 78:539-553.e8. https://doi.org/10.1016/j.molcel.2020.03.002
Karoutas A, Akhtar A (2021) Functional mechanisms and abnormalities of the nuclear lamina. Nat Cell Biol 23:116–126. https://doi.org/10.1038/s41556-020-00630-5
Kind J, Pagie L, Ortabozkoyun H et al (2013) Single-cell dynamics of genome-nuclear lamina interactions. Cell 153:178–192. https://doi.org/10.1016/j.cell.2013.02.028
Li W, Shang L, Huang K et al (2017) Identification of critical base pairs required for CTCF binding in motif M1 and M2. Protein Cell 8:544–549. https://doi.org/10.1007/s13238-017-0387-5
Lochs SJA, Kefalopoulou S, Kind J (2019) Lamina associated domains and gene regulation in development and cancer. Cells 8:271. https://doi.org/10.3390/cells8030271
Lukášová E, Kovarˇík A, Bacˇíková A et al (2017) Loss of lamin B receptor is necessary to induce cellular senescence. Biochem J 474:281–300. https://doi.org/10.1042/BCJ20160459
Lund E, Oldenburg AR, Collas P (2014) Enriched domain detector: a program for detection of wide genomic enrichment domains robust against local variations. Nucleic Acids Res 42:e92. https://doi.org/10.1093/nar/gku324
Meuleman W, Peric-Hupkes D, Kind J et al (2013) Constitutive nuclear lamina–genome interactions are highly conserved and associated with A/T-rich sequence. Genome Res 23:270–280. https://doi.org/10.1101/gr.141028.112
Moore JM, Rabaia NA, Smith LE et al (2012) Loss of maternal CTCF Is associated with peri-implantation lethality of Ctcf null embryos. PLoS ONE 7:e34915. https://doi.org/10.1371/journal.pone.0034915
Naughton C, Avlonitis N, Corless S et al (2013) Transcription forms and remodels supercoiling domains unfolding large-scale chromatin structures. Nat Struct Mol Biol 20:387–395. https://doi.org/10.1038/nsmb.2509
Neguembor MV, Martin L, Castells-García Á et al (2021) Transcription-mediated supercoiling regulates genome folding and loop formation. Mol Cell 81:3065-3081.e12. https://doi.org/10.1016/j.molcel.2021.06.009
Nora EP, Goloborodko A, Valton A-L et al (2017) Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169:930-944.e22. https://doi.org/10.1016/j.cell.2017.05.004
Patil S, Sengupta K (2021) Role of A- and B-type lamins in nuclear structure–function relationships. Biol Cell 113:295–310. https://doi.org/10.1111/boc.202000160
Peric-Hupkes D, Meuleman W, Pagie L et al (2010) Molecular maps of the reorganization of genome-nuclear lamina interactions during differentiation. Mol Cell 38:603–613. https://doi.org/10.1016/j.molcel.2010.03.016
Poleshko A, Mansfield KM, Burlingame CC et al (2013) The human protein PRR14 tethers heterochromatin to the nuclear lamina during interphase and mitotic exit. Cell Rep 5:292–301. https://doi.org/10.1016/j.celrep.2013.09.024
Poleshko A, Smith CL, Nguyen SC, et al (2019) H3K9me2 orchestrates inheritance of spatial positioning of peripheral heterochromatin through mitosis. eLife 8:e49278. https://doi.org/10.7554/eLife.49278
Pugacheva EM, Kubo N, Loukinov D et al (2020) CTCF mediates chromatin looping via N-terminal domain-dependent cohesin retention. PNAS 117:2020–2031. https://doi.org/10.1073/pnas.1911708117
Racko D, Benedetti F, Dorier J, Stasiak A (2018) Transcription-induced supercoiling as the driving force of chromatin loop extrusion during formation of TADs in interphase chromosomes. Nucleic Acids Res 46:1648–1660. https://doi.org/10.1093/nar/gkx1123
Racko D, Benedetti F, Dorier J, Stasiak A (2019) Are TADs supercoiled? Nucleic Acids Res 47:521–532. https://doi.org/10.1093/nar/gky1091
Ramirez-Martinez A, Zhang Y, Chen K et al (2021) The nuclear envelope protein Net39 is essential for muscle nuclear integrity and chromatin organization. Nat Commun 12:690. https://doi.org/10.1038/s41467-021-20987-x
Ran FA, Hsu PD, Wright J et al (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308. https://doi.org/10.1038/nprot.2013.143
Robinson JT, Thorvaldsdóttir H, Winckler W et al (2011) Integrative genomics viewer. Nat Biotechnol 29:24–26. https://doi.org/10.1038/nbt.1754
Robson MI, de las Heras JI, Czapiewski R et al (2016) Tissue-specific gene repositioning by muscle nuclear membrane proteins enhances repression of critical developmental genes during myogenesis. Molecular Cell 62:834–847. https://doi.org/10.1016/j.molcel.2016.04.035
Segal T, Salmon-Divon M, Gerlitz G (2018) The heterochromatin landscape in migrating cells and the importance of H3K27me3 for associated transcriptome alterations. Cells 7:205. https://doi.org/10.3390/cells7110205
Solovei I, Wang AS, Thanisch K et al (2013) LBR and Lamin A/C sequentially tether peripheral heterochromatin and inversely regulate differentiation. Cell 152:584–598. https://doi.org/10.1016/j.cell.2013.01.009
Soshnikova N, Montavon T, Leleu M et al (2010) Functional analysis of CTCF during mammalian limb development. Dev Cell 19:819–830. https://doi.org/10.1016/j.devcel.2010.11.009
Towbin BD, González-Aguilera C, Sack R et al (2012) Step-wise methylation of histone H3K9 positions heterochromatin at the nuclear periphery. Cell 150:934–947. https://doi.org/10.1016/j.cell.2012.06.051
Tran JR, Paulson DI, Moresco JJ et al (2021) An APEX2 proximity ligation method for mapping interactions with the nuclear lamina. J Cell Biol 220:e202002129. https://doi.org/10.1083/jcb.202002129
Tucker KL, Beard C, Dausmann J et al (1996) Germ-line passage is required for establishment of methylation and expression patterns of imprinted but not of nonimprinted genes. Genes Dev 10:1008–1020. https://doi.org/10.1101/gad.10.8.1008
van Schaik T, Vos M, Peric-Hupkes D, et al (2020) Cell cycle dynamics of lamina-associated DNA. EMBO Rep 21:e50636. https://doi.org/10.15252/embr.202050636
van Steensel B, Belmont AS (2017) Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 169:780–791. https://doi.org/10.1016/j.cell.2017.04.022
Vazquez BN, Thackray JK, Simonet NG et al (2019) SIRT7 mediates L1 elements transcriptional repression and their association with the nuclear lamina. Nucleic Acids Res 47:7870–7885. https://doi.org/10.1093/nar/gkz519
Venables WN, Ripley BD (2002) Modern applied statistics with S. Springer, New York
Wan L-B, Pan H, Hannenhalli S et al (2008) Maternal depletion of CTCF reveals multiple functions during oocyte and preimplantation embryo development. Development 135:2729–2738. https://doi.org/10.1242/dev.024539
Watson LA, Wang X, Elbert A et al (2014) Dual effect of CTCF loss on neuroprogenitor differentiation and survival. J Neurosci 34:2860–2870. https://doi.org/10.1523/JNEUROSCI.3769-13.2014
Wilson KL, Berk JM (2010) The nuclear envelope at a glance. J Cell Sci 123:1973–1978. https://doi.org/10.1242/jcs.019042
Wong X, Cutler JA, Hoskins VE, et al (2021) Mapping the micro-proteome of the nuclear lamina and lamina-associated domains. Life Sci Alliance 4:e202000774. https://doi.org/10.26508/lsa.202000774
Wong X, Stewart CL (2020) The laminopathies and the insights they provide into the structural and functional organization of the nucleus. Annu Rev Genomics Hum Genet 21:263–288. https://doi.org/10.1146/annurev-genom-121219-083616
Zullo JM, Demarco IA, Piqué-Regi R et al (2012) DNA sequence-dependent compartmentalization and silencing of chromatin at the nuclear lamina. Cell 149:1474–1487. https://doi.org/10.1016/j.cell.2012.04.035
Acknowledgements
We thank Orly Reiner, Weizmann Institute of Science for help with CRISPR and Eli Arama for S2 cells.
Funding
The research was supported by the Israel Cancer Association (grant no. 20201181) and Ariel University.
Author information
Authors and Affiliations
Contributions
LSK, NL, MSD, and GG conceived and designed research. LSK, NL, and TS conducted experiments. LSK, MSD, and GG analyzed data. LSK, MSD, and GG wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Irina Solovei
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 460 KB)
Rights and permissions
About this article
Cite this article
Kaczmarczyk, L.S., Levi, N., Segal, T. et al. CTCF supports preferentially short lamina-associated domains. Chromosome Res 30, 123–136 (2022). https://doi.org/10.1007/s10577-022-09686-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-022-09686-5