Abstract
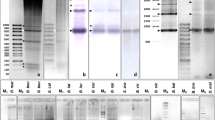
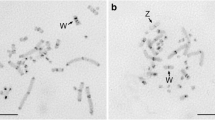
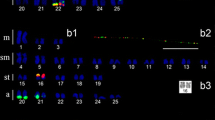
Two highly repeated DNAs, designated NmE1/NmE2 and NmE5, were identified by EcoRV digestion in the chiton Nuttallochiton mirandus (Mollusca: Polyplacophora). The comparison of the sequences obtained showed high similarity in 5′ and 3′ regions and the NmE5 sequence displayed an inserted sequence that might arise from a transposable element. Southern blotting analyses suggested a tandem organization of both satellite DNA families identified. Moreover, dot blot analyses, performed on several molluscan species, revealed a different degree of conservation of the repeated DNAs. Fluorescence in-situ hybridizations (FISH) on metaphase chromosomes showed that both satellite DNAs are located at centromeric regions.
Similar content being viewed by others
References
Altschul SF, Madden TL, Schäffer AA, et al. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25: 3389–3402.
Batistoni R, Pesole G, Marracci S, Nardi I (1995) A tandemly repeated DNA family originated from SINE-related elements in the European plethodontid salamanders (Amphibia, Urodela). J Mol Evol 40(6): 608–615.
Biscotti MA, Olmo E, Canapa A, et al. (2007) Molecular cytogenetics, genome organization and repetitive DNA in Scallop (Pecten maximus). Gene 406: 91–98.
Bradley RD, Wichman HA (1994) Rapidly evolving repetitive DNAs in a conservative genome: a test of factors that affect chromosomal evolution. Chromosome Res 2: 354–360.
Canapa A, Barucca M, Nisi Cerioni P, Olmo E (2000) A satellite DNA containing CENP-B box-like motifs is present in the Antarctic scallop Adamussium colbecki. Gene 247: 175–180.
Canapa A, Cerioni PN, Barucca M, Olmo E, Caputo V (2002) A centomeric satellite DNA may be involved in heterochromatin compactness in gobiid fishes. Chromosome Res 10: 297–304.
Clabby C, Goswami U, Flavin F, Wilkins NP, Houghton JA, Powell R (1996) Cloning, characterization and chromosomal location of a satellite DNA from the Pacific oyster Crassostrea gigas. Gene 168: 205–209.
Flavell RB (1983) Repeated sequences and genome architecture. In: Ciferri O, Dure L, eds. Structure and Function of Plant Genomes. New York: Plenum Press, pp. 1–14.
Galindo LM, Gaitan E, Baccam P, Tohme J (2004) Isolation and characterization of RNase-LTR sequence of Ty1-copia retrotransposon in common bean (Phaseolus vulgaris). Genome 47: 84–95.
Garagna S, Broccoli D, Redi CA, Searle JB, Cooke HJ, Capanna E (1995) Robertsonian metacentrics of the house mouse lose telomeric sequences but retain some minor satellite DNA in the pericentromeric area. Chromosoma 103: 685–692.
Garrido-Ramos MA, Jamelina M, Lozano R, Ruiz Rejòn C, Ruiz Rejòn M (1995) The EcoRI centromeric satellite DNA of the Sparidae family (Pisces, Perciformes) contains a sequence motive common to other vertebrate centromeric satellite DNAs. Cytogenet Cell Genet 71: 345–351.
Junier T, Pagni M (2000) Dotlet: diagonal plots in a web browser. Bioinformatics 16: 178–179.
Kapitonov VV, Holmquist GP, Jurka J (1998) L1 repeat is a basic unit of heterochromatin satellites in cetaceans. Mol Biol Evol 15: 611–612.
Loot C, Santiago N, Sanz A, Casacuberta JM (2006) The proteins encoded by the pogo-like Lemi1 element bind the TIRs and subterminal repeated motifs of the Arabidopsis Emigrant MITE: consequences for the transposition mechanism of MITEs. Nucleic Acids Res 34: 5238–5246.
López-Flores I, de la Herrán R, Garrido-Ramos MA, Boudry P, Ruiz-Rejón C, Ruiz-Rejón M (2004) The molecular phylogeny of oysters based on a satellite DNA related to transposons. Gene 339: 181–188.
Martínez-Lage A, Rodríguez F, González-Tizón A, Prats E, Cornudella L, Méndez J (2002) Comparative analysis of different satellite DNAs in four Mytilus species. Genome 45: 922–929.
Martínez-Lage A, Rodríguez-Fariña F, González-Tizón A, Méndez J (2005) Origin and evolution of Mytilus mussel satellite DNAs. Genome 48: 247–256.
Mayr B, Schweizer D, Geber G (1984) NOR activity, heterochromatin differentiation and the Robertsonian polymorphism in Sus scrofa L. J Hered 75: 79–80.
Mravinac B, Plohl M (2007) Satellite junctions identify the potential origin of new repetitive elements in the beetle Tribolium madens. Gene 394: 45–52.
Muchmore ME, Moy GW, Swanson WJ, Vacquier VD (1998) Direct sequencing of genomic DNA for characterization of a satellite DNA in five species of Eastern Pacific abalone. Mol Mar Biol Biotechnol 7: 1–6.
Neumann P, Pozarkova D, Macas J (2003) Highly abundant pea LTR retrotransposon Ogre is constitutively transcribed and partially spliced. Plant Mol Biol 53: 399–410.
Nijman IJ, Lenstra JA (2001) Mutation and recombination in cattle satellite DNA: a feedback model for the evolution of satellite DNA repeats. J Mol Evol 52: 361–371.
Odierna G, Aprea G, Barucca M et al (2008) Karyology of the Antarctic chiton Nuttallochiton mirandus (Thiele, 1906) (Mollusca: Polyplacophora) with some considerations on chromosome evolution in chitons. Chromosome Res 16. doi:10.1007/s10577-008-1247-1.
Petrović V, Plohl M (2005) Sequence divergence and conservation in organizationally distinct subfamilies of Donax trunculus satellite DNA. Gene 362: 37–43.
Plohl M, Cornudella L (1996) Characterization of a complex satellite DNA in the mollusc Donax trunculus: analysis of sequence variations and divergence. Gene 169: 157–164.
Plohl M, Cornudella L (1997) Characterization of interrelated sequence motifs in four satellite DNAs and their distribution in the genome of the mollusc Donax trunculus. J Mol Evol 44: 189–198.
Redi CA, Garagna S, Zuccotti M (1990) Robertsonian chromosome formation and fixation: the genomic scenario. Biol J Linn Soc 41: 235–255.
Ruiz-Lara S, Prats E, Sainz J, Cornudella L (1992) Cloning and characterization of a highly conserved satellite DNA from the mollusc Mytilus edulis. Gene 117: 237–242.
Salser W, Bowen S, Browne D, et al. (1976) Investigation of the organization of mammalian chromosomes at the DNA sequence level. Fed Proc 35: 23–35.
Tu Z, Orphanidis SP (2001) Micruli, a family of miniature subterminal inverted-repeat transposable elements (MSITEs): transposition without terminal inverted repeats. Mol Biol Evol 18: 893–895.
Ugarković D, Plohl M (2002) Variation in satellite DNA profiles- causes and effects. EMBO J 21: 5955–5959.
Ugarković D, Durajlijja S, Plohl M (1996) Evolution of Tribolium madens (Insecta, Coleoptera) satellite DNA through DNA inversion and insertion. J Mol Evol 42: 350–358.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Biscotti, M.A., Barucca, M., Capriglione, T. et al. Molecular and cytogenetic characterization of repetitive DNA in the Antarctic polyplacophoran Nuttallochiton mirandus . Chromosome Res 16, 907–916 (2008). https://doi.org/10.1007/s10577-008-1248-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-008-1248-0