Abstract
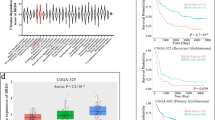
Retinoblastoma-binding protein 8 (RBBP8) affects the prognosis of patients with malignancies through various mechanisms. However, its function in gliomas is unknown. Our study explored the effects of RBBP8 on the prognosis of glioma patients, as well as its regulatory role in the glioma immune microenvironment. We used various bioinformatics methods to analyze the transcriptional profiles and methylation data of RBBP8 in gliomas from multiple databases. Our results showed that the mRNA and protein expression of RBBP8 in gliomas was higher than that in normal tissues and positively correlated with malignant clinical features such as age and WHO grade. A Kaplan–Meier analysis showed that patients with high RBBP8 expression had a poor prognosis. Cox regression demonstrated that RBBP8 was an independent risk indicator and had good diagnostic value for the poor prognosis of glioma. Importantly, RBBP8 was positively correlated with many well-known immune checkpoints (e.g., CTLA4 and PDL-1). Finally, a gene set enrichment analysis revealed that RBBP8 was remarkably enriched in cancer-related pathways such as cell cycle, DNA replication and so on. In conclusion, this study is the first to elaborate on the value of RBBP8 in the pathological process of glioma for anti-tumor immunotherapy. In addition, the expression of RBBP8 and its methylation site, cg05513509, may provide potential targets for glioma therapy.









Similar content being viewed by others
Data Availability
Data will be made available on reasonable request.
Material Availability
Data will be made available on reasonable request.
References
Bothmer A, Rommel PC, Gazumyan A, Polato F, Reczek CR, Muellenbeck MF et al (2013) Mechanism of DNA resection during intrachromosomal recombination and immunoglobulin class switching. J Exp Med 210(1):115–123. https://doi.org/10.1084/jem.20121975
Chen R, Smith-Cohn M, Cohen AL, Colman H (2017) Glioma subclassifications and their clinical significance. Neurotherapeutics 14(2):284–297. https://doi.org/10.1007/s13311-017-0519-x
De Mattos-Arruda L, Blanco-Heredia J, Aguilar-Gurrieri C, Carrillo J, Blanco J (2020) New emerging targets in cancer immunotherapy: the role of neoantigens. ESMO Open 4(Suppl 3):e000684. https://doi.org/10.1136/esmoopen-2020-000684
Desrichard A, Snyder A, Chan TA (2016) Cancer neoantigens and applications for immunotherapy. Clin Cancer Res 22(4):807–812. https://doi.org/10.1158/1078-0432.Ccr-14-3175
Gieryng A, Pszczolkowska D, Walentynowicz KA, Rajan WD, Kaminska B (2017) Immune microenvironment of gliomas. Lab Invest 97(5):498–518. https://doi.org/10.1038/labinvest.2017.19
Gu B, Chen PL (2009) Expression of PCNA-binding domain of CtIP, a motif required for CtIP localization at DNA replication foci, causes DNA damage and activation of DNA damage checkpoint. Cell Cycle 8(9):1409–1420. https://doi.org/10.4161/cc.8.9.8322
Guan X, Zhang C, Zhao J, Sun G, Song Q, Jia W (2018) CMTM6 overexpression is associated with molecular and clinical characteristics of malignancy and predicts poor prognosis in gliomas. EBioMedicine 35:233–243. https://doi.org/10.1016/j.ebiom.2018.08.012
Gusyatiner O, Hegi ME (2018) Glioma epigenetics: from subclassification to novel treatment options. Semin Cancer Biol 51:50–58. https://doi.org/10.1016/j.semcancer.2017.11.010
Hu L, Han Z, Cheng X, Wang S, Feng Y, Lin Z (2021) Expression profile analysis identifies a novel seven immune-related gene signature to improve prognosis prediction of glioblastoma. Front Genet 12:638458. https://doi.org/10.3389/fgene.2021.638458
Joly JH, Lowry WE, Graham NA (2020) Differential gene set enrichment analysis: a statistical approach to quantify the relative enrichment of two gene sets. Bioinformatics. Available at https://www.ncbi.nlm.nih.gov/pubmed/32692836. https://doi.org/10.1093/bioinformatics/btaa658
Kremer PH, Koeleman BP, Pawlikowska L, Weinsheimer S, Bendjilali N, Sidney S et al (2015) Evaluation of genetic risk loci for intracranial aneurysms in sporadic arteriovenous malformations of the brain. J Neurol Neurosurg Psychiatry 86(5):524–529. https://doi.org/10.1136/jnnp-2013-307276
Li B, Severson E, Pignon JC, Zhao H, Li T, Novak J et al (2016) Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol 17(1):174. https://doi.org/10.1186/s13059-016-1028-7
Li T, Fan J, Wang B, Traugh N, Chen Q, Liu JS et al (2017) TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res 77(21): e108–e110. Available at https://www.ncbi.nlm.nih.gov/pubmed/29092952. https://doi.org/10.1158/0008-5472.CAN-17-0307
Li J, Liu L, Liu X, Xu P, Hu Q, Yu Y (2019) The role of upregulated DDX11 as a potential prognostic and diagnostic biomarker in lung adenocarcinoma. J Cancer 10(18):4208–4216. Available at https://www.ncbi.nlm.nih.gov/pubmed/31413739. https://doi.org/10.7150/jca.33457
Li J, Rao B, Yang J, Liu L, Huang M, Liu X et al (2020) Dysregulated m6A-related regulators are associated with tumor metastasis and poor prognosis in osteosarcoma. Front Oncol 10:769. Available at https://www.ncbi.nlm.nih.gov/pubmed/32582536. https://doi.org/10.3389/fonc.2020.00769
Liu F, Lee WH (2006) CtIP activates its own and cyclin D1 promoters via the E2F/RB pathway during G1/S progression. Mol Cell Biol 26(8):3124–3134. https://doi.org/10.1128/mcb.26.8.3124-3134.2006
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK et al (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131(6):803–820. https://doi.org/10.1007/s00401-016-1545-1
Lu YC, Robbins PF (2016) Cancer immunotherapy targeting neoantigens. Semin Immunol 28(1):22–27. https://doi.org/10.1016/j.smim.2015.11.002
Odunsi K (2017) Immunotherapy in ovarian cancer. Ann Oncol, 28(suppl_8):viii1–viii7. https://doi.org/10.1093/annonc/mdx444
Reni M, Mazza E, Zanon S, Gatta G, Vecht CJ (2017) Central nervous system gliomas. Crit Rev Oncol Hematol 113:213–234. https://doi.org/10.1016/j.critrevonc.2017.03.021
Rodríguez-Cerdeira C, Carnero Gregorio M, López-Barcenas A, Sánchez-Blanco E, Sánchez-Blanco B, Fabbrocini G et al (2017) Advances in immunotherapy for melanoma: a comprehensive review. Mediators Inflamm 2017:3264217. https://doi.org/10.1155/2017/3264217
Rolfo C, Caglevic C, Santarpia M, Araujo A, Giovannetti E, Gallardo CD et al (2017) Immunotherapy in NSCLC: a promising and revolutionary weapon. Adv Exp Med Biol 995:97–125. https://doi.org/10.1007/978-3-319-53156-4_5
Schumacher TN, Schreiber RD (2015) Neoantigens in cancer immunotherapy. Science 348(6230):69–74. https://doi.org/10.1126/science.aaa4971
Sjöstedt E, Zhong W, Fagerberg L, Karlsson M, Mitsios N, Adori C et al (2020) An atlas of the protein-coding genes in the human, pig, and mouse brain. Science (New York, NY). https://doi.org/10.1126/science.aay5947
Steven A, Fisher SA, Robinson BW (2016) Immunotherapy for lung cancer. Respirology 21(5):821–833. https://doi.org/10.1111/resp.12789
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102(43):15545–15550. https://doi.org/10.1073/pnas.0506580102
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res, 45(W1):W98–W102. Available at https://www.ncbi.nlm.nih.gov/pubmed/28407145. https://doi.org/10.1093/nar/gkx247
Williams GH, Stoeber K (2012) The cell cycle and cancer. J Pathol 226(2):352–364. https://doi.org/10.1002/path.3022
Yu X, Chen J (2004) DNA damage-induced cell cycle checkpoint control requires CtIP, a phosphorylation-dependent binding partner of BRCA1 C-terminal domains. Mol Cell Biol 24(21):9478–9486. https://doi.org/10.1128/mcb.24.21.9478-9486.2004
Yun MH, Hiom K (2009) CtIP-BRCA1 modulates the choice of DNA double-strand-break repair pathway throughout the cell cycle. Nature 459(7245):460–463. https://doi.org/10.1038/nature07955
Zhang W, Song Y, He X, Liu X, Zhang Y, Yang Z et al (2020) Prognosis value of RBBP8 expression in plasma cell myeloma. Cancer Gene Ther 27(1–2):22–29. https://doi.org/10.1038/s41417-018-0069-3
Zhang Z, Li H, Jiang S, Li R, Li W, Chen H, Bo X (2019) A survey and evaluation of Web-based tools/databases for variant analysis of TCGA data. Brief Bioinform, 20(4):1524–1541. Available at https://www.ncbi.nlm.nih.gov/pubmed/29617727. https://doi.org/10.1093/bib/bby023
Zhao Z, Zhang KN, Wang Q, Li G, Zeng F, Zhang Y et al (2021) Chinese Glioma Genome Atlas (CGGA): a comprehensive resource with functional genomic data from Chinese gliomas. Genomics Proteomics Bioinformatics. Available at https://www.ncbi.nlm.nih.gov/pubmed/33662628. https://doi.org/10.1016/j.gpb.2020.10.005
Funding
This work was supported by the Thousand Talents Plan of Central Plains (ZYQR201912122).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10571_2022_1198_MOESM4_ESM.tif
Supplementary file4 Fig.S1. The gene expression profile across all tumor samples and paired normal tissues. (TIF 5738 kb)
10571_2022_1198_MOESM5_ESM.tif
Supplementary file5 Fig.S2. Relationship between the expression of RBBP8 and immune checkpoints in GBM, including PDCD1 (a), CD274 (b), PDCD1LG2 (c), CTLA4 (d) and HAVCR2 (e); LGG: PDCD1 (f), CD274 (g), PDCD1LG2 (h), CTLA4 (i) and HAVCR2 (j).(TIF 4830 kb)
10571_2022_1198_MOESM6_ESM.tif
Supplementary file6 Fig.S3. GSEA analysis of RBBP8 in glioma using CGGA RNA-Seq dataset. (a) cell cycle signaling pathway, (b) P53 signaling pathway, (c) DNA replication, (d) homologous recombination, (e) mismatch repair, (f) antigen processing and presentation.(TIF 44836 kb)
10571_2022_1198_MOESM7_ESM.tif
Supplementary file7 Fig.S4. GSEA analysis of RBBP8 in glioma using CGGA microarray dataset. (a) cell cycle signaling pathway, (b) P53 signaling pathway, (c) DNA replication, (d) homologous recombination, (e) mismatch repair, (f) antigen processing and presentation. (TIF 5779 kb)
Rights and permissions
About this article
Cite this article
Liu, Z., Cheng, X., Lin, S. et al. Prognostic Significance of mRNA Expression RBBP8 or Its Methylation in Gliomas. Cell Mol Neurobiol 43, 409–422 (2023). https://doi.org/10.1007/s10571-022-01198-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-022-01198-4