Abstract
Background
Osteosarcomas (OS) are frequent primary sarcomas of the bone in children and adolescents. The long non-coding RNAs (lncRNAs) can affect the progression of many cancers by their sense transcripts. The present study was designed to probe the role of ZMIZ1-AS1 and the downstream pathway in OS progression.
Methods
Cell proliferation, invasion, and migration were detected by colony formation, transwell, and wound healing assays. The binding of SOX2 or MYC protein with ZMIZ1-AS1 promoter was explored by ChIP assay and dual-luciferase reporter assay. Interaction between PTBP1 protein and ZMIZ1-AS1 (or ZMIZ1 mRNA) was detected by RIP assay.
Results
SOX2 and MYC are the downstream effectors of the Hippo pathway and transcriptionally activated ZMIZ1-AS1. Compared to the controls, OS tissues and cells contained higher ZMIZ1-AS1 expression. Silencing of ZMIZ1-AS1 repressed OS cell viability, proliferation, migration, and invasion. Our findings further showed that ZMIZ1-AS1 recruits RNA-binding protein PTBP1 to stabilize ZMIZ1 mRNA. PTBP1 or ZMIZ1 overexpression rescues the suppressive effects of silenced ZMIZ1-AS1 on OS cellular processes. Importantly, ZMIZ1-AS1 promotes OS growth in vivo by stabilization of ZMIZ1.
Conclusions
Long non-coding RNA ZMIZ1-AS1 promotes OS progression by stabilization of ZMIZ1.
Graphical abstract

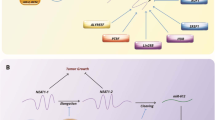
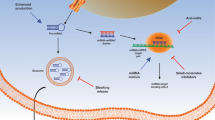
The Hippo pathway is inactivated in osteosarcoma. Transcriptional factors SOX2 and MYC downstream the Hippo pathway induce the upregulation of ZMIZ1-AS1 in osteosarcoma. ZMIZ1-AS1 recruits RNA binding protein PTBP1 that stabilizes ZMIZ1, the sense transcript of ZMIZ1-AS1. ZMIZ1-AS1 promotes osteosarcoma cell viability, proliferation, migration, and invasion by ZMIZ1 in a PTBP1 dependent manner







Similar content being viewed by others
Change history
29 December 2022
A Correction to this paper has been published: https://doi.org/10.1007/s10565-022-09788-0
References
Badouel C, McNeill H. SnapShot: The hippo signaling pathway. Cell. 145:484-484.e481. 2011;145:484–484.e1. https://doi.org/10.1016/j.cell.2011.04.009.
Carrieri C, Cimatti L, Biagioli M, Beugnet A, Zucchelli S, Fedele S, Pesce E, Ferrer I, Collavin L, Santoro C, Forrest ARR, Carninci P, Biffo S, Stupka E, Gustincich S. Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat. Nature. 2012;491:454–7. https://doi.org/10.1038/nature11508.
Cersosimo F, Lonardi S, Bernardini G, Telfer B, Mandelli GE, Santucci A, Vermi W, Giurisato E. Tumor-associated macrophages in osteosarcoma: from mechanisms to therapy. international journal of molecular sciences. 21. 2020;21. https://doi.org/10.3390/ijms21155207.
Cheng C, Ding Q, Zhang Z, Wang S, Zhong B, Huang X, Shao Z. PTBP1 modulates osteosarcoma chemoresistance to cisplatin by regulating the expression of the copper transporter SLC31A1. Journal of cellular and molecular medicine. 2020;24:5274–89. https://doi.org/10.1111/jcmm.15183.
Chu C, Zhang QC, da Rocha ST, Flynn RA, Bharadwaj M, Calabrese JM, Magnuson T, Heard E, Chang HY. Systematic discovery of Xist RNA binding proteins. Cell. 2015;161:404–16. https://doi.org/10.1016/j.cell.2015.03.025.
Corre I, Verrecchia F, Crenn V, Redini F, Trichet V. The osteosarcoma microenvironment: a complex but targetable ecosystem. Cells. 2020;9. https://doi.org/10.3390/cells9040976.
D'Angelo D, Arra C, Fusco A. RPSAP52 lncRNA Inhibits p21Waf1/CIP Expression by interacting with the RNA binding protein HuR. Oncology research. 2020;28:191–201. https://doi.org/10.3727/096504019x15761465603129.
Faghihi MA, Modarresi F, Khalil AM, Wood DE, Sahagan BG, Morgan TE, Finch CE, St. Laurent III G, Kenny PJ, Wahlestedt C. Expression of a noncoding RNA is elevated in Alzheimer's disease and drives rapid feed-forward regulation of beta-secretase. Nature medicine. 2008;14:723–30. https://doi.org/10.1038/nm1784.
Fan TM. Animal models of osteosarcoma. Expert review of anticancer therapy. 2010;10:1327–38. https://doi.org/10.1586/era.10.107.
Fernandes I, Melo-Alvim C, Lopes-Brás R, Esperança-Martins M, Costa L. Osteosarcoma pathogenesis leads the way to new target treatments. International journal of molecular sciences. 2021;22. https://doi.org/10.3390/ijms22020813.
Ferrè F, Colantoni A, Helmer-Citterich M. Revealing protein-lncRNA interaction. Briefings in bioinformatics. 2016;17:106–16. https://doi.org/10.1093/bib/bbv031.
Fu D, Lu C, Qu X, Li P, Chen K, Shan L, et al. (2019) LncRNA TTN-AS1 regulates osteosarcoma cell apoptosis and drug resistance via the miR-134-5p/MBTD1 axis. Aging. 11:8374-8385 doi:10.18632/aging.102325
Guil S, Esteller MJNs, biology m. Cis-acting noncoding RNAs: friends and foes. 2012;19:1068–75. https://doi.org/10.1038/nsmb.2428.
Gumbiner BM, Kim NG. The Hippo-YAP signaling pathway and contact inhibition of growth. Journal of cell science. 2014;127:709–17. https://doi.org/10.1242/jcs.140103.
Harvey KF, Zhang X, Thomas DM. The Hippo pathway and human cancer. Nature reviews Cancer. 2013;13:246–57. https://doi.org/10.1038/nrc3458.
Huang Q, Shi SY, Ji HB, Xing SX. LncRNA BE503655 inhibits osteosarcoma cell proliferation, invasion/migration via Wnt/β-catenin pathway. Bioscience reports. 2019;39. https://doi.org/10.1042/bsr20182200.
Ji XY, Zhong G, Zhao B (2017) Molecular mechanisms of the mammalian Hippo signaling pathway. Yi chuan = Hereditas. 39:546-567 doi:10.16288/j.yczz.17-094
Kim J, Abdelmohsen K, Yang X, De S, Grammatikakis I, Noh JH, et al. LncRNA OIP5-AS1/cyrano sponges RNA-binding protein HuR. Nucleic acids research. 2016;44:2378–92. https://doi.org/10.1093/nar/gkw017.
Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic acids research. 2014;42:D92–7. https://doi.org/10.1093/nar/gkt1248.
Liu Z, Dai J, Shen HJBebaMbod. Systematic analysis reveals long noncoding RNAs regulating neighboring transcription factors in human cancers. 2018;1864:2785–92. https://doi.org/10.1016/j.bbadis.2018.05.006.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods (San Diego, Calif). 2001;25:402–8. https://doi.org/10.1006/meth.2001.1262.
Matsui K, Nishizawa M, Ozaki T, Kimura T, Hashimoto I, Yamada M, Kaibori M, Kamiyama Y, Ito S, Okumura T. Natural antisense transcript stabilizes inducible nitric oxide synthase messenger RNA in rat hepatocytes. Hepatology (Baltimore, Md). 2008;47:686–97. https://doi.org/10.1002/hep.22036.
Rakowski LA, Garagiola DD, Li CM, Decker M, Caruso S, Jones M, Kuick R, Cierpicki T, Maillard I, Chiang MY. Convergence of the ZMIZ1 and NOTCH1 pathways at C-MYC in acute T lymphoblastic leukemias. Cancer research. 2013;73:930–41. https://doi.org/10.1158/0008-5472.Can-12-1389.
Rogers LM, Riordan JD, Swick BL, Meyerholz DK, Dupuy AJ. Ectopic expression of Zmiz1 induces cutaneous squamous cell malignancies in a mouse model of cancer. The Journal of investigative dermatology. 2013;133:1863–9. https://doi.org/10.1038/jid.2013.77.
Shen Y, Xu J, Pan X, Zhang Y, Weng Y, Zhou D, He S. LncRNA KCNQ1OT1 sponges miR-34c-5p to promote osteosarcoma growth via ALDOA enhanced aerobic glycolysis. Cell Death Dis. 2020;11:278. https://doi.org/10.1038/s41419-020-2485-1.
Song N, Kim K, Shin A, Park JW, Chang HJ, Shi J, Cai Q, Kim DY, Zheng W, Oh JH. Colorectal cancer susceptibility loci and influence on survival. Genes, chromosomes & cancer. 2018;57:630–7. https://doi.org/10.1002/gcc.22674.
Song N, Shin A, Park JW, Kim J, Oh JH. Common risk variants for colorectal cancer: an evaluation of associations with age at cancer onset. Scientific reports. 2017;7:40644. https://doi.org/10.1038/srep40644.
Su X, Teng J, Jin G, Li J, Zhao Z, Cao X, Guo Y, Guo M, Li X, Wu J, Wang C, Guo Z, Guo Q. ELK1-induced upregulation of long non-coding RNA MIR100HG predicts poor prognosis and promotes the progression of osteosarcoma by epigenetically silencing LATS1 and LATS2. Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie. 2019;109:788–97. https://doi.org/10.1016/j.biopha.2018.10.029.
Tan ML, Choong PF, Dass CR. Osteosarcoma: conventional treatment vs. gene therapy. Cancer biology & therapy. 2009;8:106–17. https://doi.org/10.4161/cbt.8.2.7385.
Tsukamoto S, Errani C, Angelini A, Mavrogenis AF. Current treatment considerations for osteosarcoma metastatic at presentation. Orthopedics. 2020;43:e345–58. https://doi.org/10.3928/01477447-20200721-05.
Wang M, Mao C, Ouyang L, Liu Y, Lai W, Liu N, Shi Y, Chen L, Xiao D, Yu F, Wang X, Zhou H, Cao Y, Liu S, Yan Q, Tao Y, Zhang B. Correction to: long noncoding RNA LINC00336 inhibits ferroptosis in lung cancer by functioning as a competing endogenous RNA. Cell death and differentiation. 2020;27:1447. https://doi.org/10.1038/s41418-019-0394-6.
Yao XY, Liu JF, Luo Y, Xu XZ, Bu J (2021) LncRNA HOTTIP facilitates cell proliferation, invasion, and migration in osteosarcoma by interaction with PTBP1 to promote KHSRP level. Cell cycle (Georgetown, Tex). 20:283-297 doi:10.1080/15384101.2020.1870820
Yi W, Li J, Zhu X, Wang X, Fan L, Sun W, Liao L, Zhang J, Li X, Ye J, Chen F, Taipale J, Chan KM, Zhang L, Yan J. CRISPR-assisted detection of RNA-protein interactions in living cells. Nature methods. 2020;17:685–8. https://doi.org/10.1038/s41592-020-0866-0.
Yu FX, Zhao B, Guan KL. Hippo pathway in organ size control, tissue homeostasis, and cancer. Cell. 2015;163:811–28. https://doi.org/10.1016/j.cell.2015.10.044.
Zhang H, Liu Y, Yan L, Zhang M, Yu X, Du W, et al. Increased levels of the long noncoding RNA, HOXA-AS3, promote proliferation of A549 cells. Cell death & disease. 2018;9:707. https://doi.org/10.1038/s41419-018-0725-4.
Zhang J, Ma X, Zhou R, Zhou Y. TRPS1 and YAP1 regulate cell proliferation and drug resistance of osteosarcoma via competitively binding to the target of circTADA2A - miR-129-5p. OncoTargets and therapy. 2020;13:12397–407. https://doi.org/10.2147/ott.S276953.
Zhao D, Wang S, Chu X, Han D. LncRNA HIF2PUT inhibited osteosarcoma stem cells proliferation, migration and invasion by regulating HIF2 expression. Artificial cells, nanomedicine, and biotechnology. 2019;47:1342–8. https://doi.org/10.1080/21691401.2019.1596934.
Zheng S, Jiang F, Ge D, Tang J, Chen H, Yang J, Yao Y, Yan J, Qiu J, Yin Z, Ni Y, Zhao L, Chen X, Li H, Yang L. LncRNA SNHG3/miRNA-151a-3p/RAB22A axis regulates invasion and migration of osteosarcoma. Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie. 2019;112:108695. https://doi.org/10.1016/j.biopha.2019.108695.
Zhou Y, Jin Q, Xiao W, Sun C. Tankyrase1 antisense oligodeoxynucleotides suppress the proliferation, migration and invasion through Hippo/YAP pathway in human osteosarcoma cells. Pathology, research and practice. 2019;215:152381. https://doi.org/10.1016/j.prp.2019.03.010.
Zhu G, Wang Y, Mijiti M, Wang Z, Wu PF, Jiafu D. Upregulation of miR-130b enhances stem cell-like phenotype in glioblastoma by inactivating the Hippo signaling pathway. Biochemical and biophysical research communications. 2015;465:194–9. https://doi.org/10.1016/j.bbrc.2015.07.149.
Availability of data and material
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
Not applicable.
Author information
Authors and Affiliations
Contributions
Yichi Zhou and Chengjun Sun conceived and designed the research; Yichi Zhou, Qi Jin, Jianzhong Chang, and Zufa Zhao performed the research; Yichi Zhou, Qi Jin and Chengjun Sun analyzed the data; Yichi Zhou wrote the paper; Chengjun Sun edited the manuscript. All authors approved final version of manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Written informed consent was obtained from all participants. The study complied with the Declaration of Helsinki and was approved by the Ethics Committee of CR & WISCO General Hospital (Hubei, China).
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
Supplementary Fig. 1 Quantitative data of the rescue assays. A Quantitative data of colony assays in Fig. 6B. B Quantitative data of wound healing assays in Fig. 6C. (C) Quantitative data of transwell invasion assays in Fig. 6D. p**<0.01, p***<0.001 (TIF 436 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, Y., Jin, Q., Chang, J. et al. Long non-coding RNA ZMIZ1-AS1 promotes osteosarcoma progression by stabilization of ZMIZ1. Cell Biol Toxicol 38, 1013–1026 (2022). https://doi.org/10.1007/s10565-021-09641-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10565-021-09641-w