Abstract
Background
Although long non-coding RNA (lncRNA) HCP plays essential roles in human cancers, its function and mechanism in multiple myeloma (MM) have not crystallized.
Methods
HCP5 level in MM was assessed through qRT-PCR. A series of functional investigations were conducted to evaluate the influences of HCP5 on proliferation and apoptosis. Bioinformatics analysis and RIP/RNA pull-down assays were carried out to determine the relationships among HCP5, miR-128-3p, and PLAGL2. Relative protein level was determined through Western blot. A xenograft tumor model was applied for validating the roles of HCP5/miR-128-3p/PLAGL2 axis in vivo.
Results
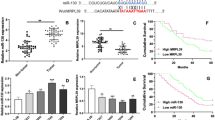
HCP5 was significantly increased in MM. HCP5 knockdown effectively thwarted the proliferative rate and cell cycle of MM cell lines and suppressed tumor growth. HCP5 regulated PLAGL2 expression by sponging miR-128-3p. PLAGL2 overexpression effectively rescued cells from influences by sh-HCP5 on cell proliferative and apoptotic rates. Additionally, HCP5 knockdown significantly inhibited Wnt/β-catenin/cyclin D1 signaling, and these effects were eliminated by PLAGL2 overexpression.
Conclusion
Our study revealed that HCP5/miR-128-3p/PLAGL2 is closely correlated to MM development by modulating Wnt/β-catenin/cyclin D1 signaling.
Graphical abstract
HCP5 promoted cell proliferation and tumor formation of MM cells by activating the Wnt/β‐catenin/CCND1 signaling pathway by sponging miR-128-3p to increase PLAGL2 expression








Similar content being viewed by others
Data availability
Some of the data that support the findings of this study are not publicly available due to their containing information that could compromise the privacy of research participants, but are available on request from the corresponding author: Ruixiang Xia. Department of Hematology, the First Affiliated Hospital of Anhui Medical University, 218 Jixi Road, Shushan District, Hefei 230,022, Anhui, China. Email address: RuixiangXiaHospita@163.com.
References
Adams BD, Parsons C, Walker L, Zhang WC, Slack FJ. Targeting noncoding RNAs in disease. J Clin Investig. 2017;127(3):761–71. https://doi.org/10.1172/jci84424.
Amodio N, D’Aquila P, Passarino G, Tassone P, Bellizzi D. Epigenetic modifications in multiple myeloma: recent advances on the role of DNA and histone methylation. Expert Opin Ther Targets. 2017;21(1):91–101. https://doi.org/10.1080/14728222.2016.1266339.
An QM, Liu YG, Lin L, Liu Y, Lu J, Ge Q. Expression of extracellular matrix protein reelin in patients with multiple myeloma and its relationship with prognosis. Zhongguo Shi Yan Xue Ye Xue Za Zhi. 2018;26(3):789–95. https://doi.org/10.7534/j.issn.1009-2137.2018.03.026.
Baghdadi M, Ishikawa K, Nakanishi S, Murata T, Umeyama Y, Kobayashi T, et al. A role for IL-34 in osteolytic disease of multiple myeloma. Blood Advances. 2019;3(4):541–51. https://doi.org/10.1182/bloodadvances.2018020008.
Baker K, Gordon SL, Melland H, Bumbak F, Scott DJ, Jiang TJ, et al. SYT1-associated neurodevelopmental disorder: a case series. Brain. 2018;141(9):2576–91. https://doi.org/10.1093/brain/awy209.
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global Cancer Statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. https://doi.org/10.3322/caac.21492.
Cao DW, Liu MM, Duan R, Tao YF, Zhou JS, Fang WR, et al. The lncRNA Malat1 functions as a ceRNA to contribute to berberine-mediated inhibition of HMGB1 by sponging miR-181c-5p in poststroke inflammation. Acta Pharmacologica Sinica. 2020;41(1):22–33. https://doi.org/10.1038/s41401-019-0284-y.
Cea M, Cagnetta A, Fulciniti M, Tai YT, Hideshima T, Chauhan D, et al. Targeting NAD+ salvage pathway induces autophagy in multiple myeloma cells via mTORC1 and extracellular signal-regulated kinase (ERK1/2) inhibition. Blood. 2012;120(17):3519–29. https://doi.org/10.1182/blood-2012-03-416776.
Chen M, Li L. SALL4 promotes the tumorigenicity of cervical cancer cells through activation of the Wnt/β-catenin pathway via CTNNB1. Cancer Sci. 2019;110(9):2794–805. https://doi.org/10.1111/cas.14140.
Chen X, Qi M, Yang Q, Li JY. MiR-299-3p functions as a tumor suppressor in thyroid cancer by regulating SHOC2. European Review for Medical and Pharmacological Sciences. 2019;23(1):232–40. https://doi.org/10.26355/eurrev_201901_16769.
Clark KC, Hewett DR. Targeted disruption of bone marrow stromal cell-derived Gremlin1 limits multiple myeloma disease progression in vivo. 2020;12(8). doi:https://doi.org/10.3390/cancers12082149.
Dang S, Zhou J, Wang Z, Wang K, Dai S, He S. MiR-299–3p functions as a tumor suppressor via targeting Sirtuin 5 in hepatocellular carcinoma. Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie. 2018;106:966–75. doi:https://doi.org/10.1016/j.biopha.2018.06.042.
Detappe A, Bustoros M, Mouhieddine TH, Ghoroghchian PP. Advancements in nanomedicine for multiple myeloma. Trends Mol Med. 2018;24(6):560–74. https://doi.org/10.1016/j.molmed.2018.04.005.
Fu C, Li D, Zhang X, Liu N, Chi G, Jin X. LncRNA PVT1 Facilitates tumorigenesis and progression of glioma via regulation of MiR-128–3p/GREM1 axis and BMP signaling pathway. Neurotherapeutics. 2018;15(4):1139–57. https://doi.org/10.1007/s13311-018-0649-9.
Garzon R, Marcucci G, Croce CM. Targeting microRNAs in cancer: rationale, strategies and challenges. Nature Reviews Drug Discovery. 2010;9(10):775–89. https://doi.org/10.1038/nrd3179.
Gu Y, Xiao X, Yang S. LncRNA MALAT1 acts as an oncogene in multiple myeloma through sponging miR-509-5p to modulate FOXP1 expression. Oncotarget. 2017;8(60):101984–93. https://doi.org/10.18632/oncotarget.21957.
Gupta N, Kumar R, Seth T, Garg B, Sharma A. Targeting of stromal versican by miR-144/199 inhibits multiple myeloma by downregulating FAK/STAT3 signalling. RNA Biol. 2020;17(1):98–111. https://doi.org/10.1080/15476286.2019.1669405.
Hrašovec S, Hauptman N, Glavač D, Jelenc F, Ravnik-Glavač M. TMEM25 is a candidate biomarker methylated and down-regulated in colorectal cancer. Disease Markers. 2013;34(2):93–104. https://doi.org/10.3233/dma-120948.
Hu R, Lu Z. Long non-coding RNA HCP5 promotes prostate cancer cell proliferation by acting as the sponge of miR-4656 to modulate CEMIP expression. Oncology Reports. 2020;43(1):328–36. https://doi.org/10.3892/or.2019.7404.
Ibrahim AM, Sabet S, El-Ghor AA, Kamel N. Fibulin-2 is required for basement membrane integrity of mammary epithelium. Sci rep. 2018;8(1):14139. https://doi.org/10.1038/s41598-018-32507-x.
Juli G, Oliverio M. Anti-tumor activity and epigenetic impact of the polyphenol oleacein in multiple myeloma. Cancer (Basel). 2019;11(7):990. https://doi.org/10.3390/cancers11070990.
Kong X, Duan Y, Sang Y, Li Y. LncRNA-CDC6 promotes breast cancer progression and function as ceRNA to target CDC6 by sponging microRNA-215. J Cell Physiol. 2019;234(6):9105–17. https://doi.org/10.1002/jcp.27587.
Lai CQ, Parnell LD, Arnett DK, García-Bailo B, Tsai MY, Kabagambe EK, et al. WDTC1, the ortholog of Drosophila adipose gene, associates with human obesity, modulated by MUFA intake. Obesity (silver Spring, Md). 2009;17(3):593–600. https://doi.org/10.1038/oby.2008.561.
Laubach J, Garderet L, Mahindra A, Gahrton G, Caers J, Sezer O, et al. Management of relapsed multiple myeloma: recommendations of the International Myeloma Working Group. Leukemia. 2016;30(5):1005–17. https://doi.org/10.1038/leu.2015.356.
Leone E, Morelli E, Di Martino MT, Amodio N, Foresta U, Gullà A, et al. Targeting miR-21 inhibits in vitro and in vivo multiple myeloma cell growth. Clin Cancer Res. 2013;19(8):2096–106. https://doi.org/10.1158/1078-0432.ccr-12-3325.
Li N, Li D, Du Y, Su C, Yang C, Lin C, et al. Overexpressed PLAGL2 transcriptionally activates Wnt6 and promotes cancer development in colorectal cancer. Oncology Reports. 2019;41(2):875–84. https://doi.org/10.3892/or.2018.6914.
Liang L, Xu J, Wang M, Xu G, Zhang N, Wang G, et al. LncRNA HCP5 promotes follicular thyroid carcinoma progression via miRNAs sponge. Cell Death & Disease. 2018;9(3):372. https://doi.org/10.1038/s41419-018-0382-7.
Liu H, Devraj K, Möller K, Liebner S, Hecker M, Korff T. EphrinB-mediated reverse signalling controls junctional integrity and pro-inflammatory differentiation of endothelial cells. Thrombosis and Haemostasis. 2014;112(1):151–63. https://doi.org/10.1160/th13-12-1034.
Liu T, Zhang X, Du L, Wang Y, Liu X, Tian H, et al. Exosome-transmitted miR-128-3p increase chemosensitivity of oxaliplatin-resistant colorectal cancer. Molecular Cancer. 2019a;18(1):43. https://doi.org/10.1186/s12943-019-0981-7.
Liu Y, Wang J, Dong L, Xia L, Zhu H, Li Z, et al. Long noncoding RNA HCP5 regulates pancreatic cancer gemcitabine (GEM) resistance by sponging Hsa-miR-214-3p to target HDGF. OncoTargets and Therapy. 2019b;12:8207–16. https://doi.org/10.2147/ott.s222703.
Lu TX, Rothenberg ME. MicroRNA. J Allergy Clin Immunol. 2018;141(4):1202–7. https://doi.org/10.1016/j.jaci.2017.08.034.
Parks RN, Kim CJ, Al-Safi ZA, Armstrong AA, Zore T, Moatamed NA. Multiple myeloma 1 transcription factor is superior to CD138 as a marker of plasma cells in endometrium. Int J Surg Pathol. 2019;27(4):372–9. https://doi.org/10.1177/1066896918814307.
Pawlyn C, Bright MD, Buros AF, Stein CK, Walters Z, Aronson LI, et al. Overexpression of EZH2 in multiple myeloma is associated with poor prognosis and dysregulation of cell cycle control. Blood Cancer. 2017;7(3):e549. https://doi.org/10.1038/bcj.2017.27.
She X, Yu Z, Cui Y, Lei Q, Wang Z, Xu G, et al. miR-128 and miR-149 enhance the chemosensitivity of temozolomide by Rap1B-mediated cytoskeletal remodeling in glioblastoma. Oncology Reports. 2014;32(3):957–64. https://doi.org/10.3892/or.2014.3318.
Spaan I, Raymakers RA, van de Stolpe A, Peperzak V. Wnt Signaling in Multiple Myeloma: a Central Player in Disease with Therapeutic Potential. 2018;11(1):67. https://doi.org/10.1186/s13045-018-0615-3.
Wang JH, Zheng WW, Cheng ST, Liu BX, Liu FR, Song JQ. Correlation between microRNA-21 and sprouty homolog 2 gene expression in multiple myeloma. Molecular Medicine Reports. 2015;11(6):4220–4. https://doi.org/10.3892/mmr.2015.3280.
Wang L, Luan T, Zhou S, Lin J, Yang Y, Liu W, et al. LncRNA HCP5 promotes triple negative breast cancer progression as a ceRNA to regulate BIRC3 by sponging miR-219a-5p. Cancer Medicine. 2019;8(9):4389–403. https://doi.org/10.1002/cam4.2335.
Weiss GJ, Bemis LT, Nakajima E, Sugita M, Birks DK, Robinson WA, et al. EGFR regulation by microRNA in lung cancer: correlation with clinical response and survival to gefitinib and EGFR expression in cell lines. Ann Oncol. 2008;19(6):1053–9. https://doi.org/10.1093/annonc/mdn006.
Wu H, Liu C, Yang Q, Xin C, Du J, Sun F, et al. MIR145-3p promotes autophagy and enhances bortezomib sensitivity in multiple myeloma by targeting HDAC4. Autophagy. 2020;16(4):683–97. https://doi.org/10.1080/15548627.2019.1635380.
Xu B, Zhang X, Wang S, Shi B. MiR-449a suppresses cell migration and invasion by targeting PLAGL2 in breast cancer. Pathology, Research and Practice. 2018;214(5):790–5. https://doi.org/10.1016/j.prp.2017.12.012.
Yang C, Sun J, Liu W, Yang Y, Chu Z, Yang T, et al. Long noncoding RNA HCP5 contributes to epithelial-mesenchymal transition in colorectal cancer through ZEB1 activation and interacting with miR-139-5p. Am J Transl Res. 2019;11(2):953–63.
Yang J, Li C, Mudd A, Gu X. LncRNA PVT1 predicts prognosis and regulates tumor growth in prostate cancer. Bioscience, Biotechnology, and Biochemistry. 2017;81(12):2301–6. https://doi.org/10.1080/09168451.2017.1387048.
Yu Y, Shen HM, Fang DM, Meng QJ, Xin YH. LncRNA HCP5 promotes the development of cervical cancer by regulating MACC1 via suppression of microRNA-15a. Eur Rev Med Pharmacol Sci. 2018;22(15):4812–9. https://doi.org/10.26355/eurrev_201808_15616.
Yu Y, Zhao JD, Yang H. MiR-299-3p inhibits proliferation and invasion of cervical cancer cell via targeting TCF4. Eur Rev Med Pharmacol Sci. 2019;23(13):5621–7. https://doi.org/10.26355/eurrev_201907_18296.
Yun WK, Hu YM, Zhao CB, Yu DY, Tang JB. HCP5 promotes colon cancer development by activating AP1G1 via PI3K/AKT pathway. European Review for Medical and Pharmacological Sciences. 2019;23(7):2786–93. https://doi.org/10.26355/eurrev_201904_17553.
Zhang Y, Chao T, Li R, Liu W, Chen Y, Yan X, et al. MicroRNA-128 inhibits glioma cells proliferation by targeting transcription factor E2F3a. J Mol Med (Berl). 2009;87(1):43–51. https://doi.org/10.1007/s00109-008-0403-6.
Zhao J, Li D, Fang L. MiR-128–3p suppresses breast cancer cellular progression via targeting LIMK1. Biomed Pharmacother. 2019;115:108947. doi:https://doi.org/10.1016/j.biopha.2019.108947.
Zhou J, Liu H, Zhang L, Liu X, Zhang C, Wang Y, et al. DJ-1 promotes colorectal cancer progression through activating PLAGL2/Wnt/BMP4 axis. Cell Death & Disease. 2018;9(9):865. https://doi.org/10.1038/s41419-018-0883-4.
Zhou Z, Wu L, Liu Z, Zhang X, Han S, Zhao N, et al. MicroRNA-214-3p targets the PLAGL2-MYH9 axis to suppress tumor proliferation and metastasis in human colorectal cancer. Aging. 2020;12(10):9633–57. https://doi.org/10.18632/aging.103233.
Acknowledgements
We thank the major projects of Natural Science Research Projects of Universities in Anhui Province for the support.
Funding
This work was supported by the Anhui Medical University 2020 School Research Fund (2020xky163).
Author information
Authors and Affiliations
Contributions
Qinhua Liu, Ruixiang Xia: study concepts, literature research, clinical studies, data analysis, experimental studies, manuscript writing and review; Ruonan Ran: study design, literature research, experimental studies, and manuscript editing; Mingyue Song: definition of intellectual content, clinical studies, data acquisition, and statistical analysis; Xiaodan Li1: data acquisition, manuscript preparation, and data analysis; Zhengsheng Wu, Guanrong Dai: data acquisition and statistical analysis. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Informed consent was obtained from all individual participants included in the study. All producers were approved by the Human and Animal Ethics Committee of the First Affiliated Hospital of Anhui Medical University. Procedures operated in this research were completed in keeping with the standards set out in the Announcement of Helsinki and Laboratory Guidelines of Research in China.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
•HCP5 was significantly upregulated in MM tissues and cancer cell lines.
•HCP5 knockdown effectively inhibited proliferation and cell cycle of MM cell lines in vitro and suppressed tumor growth in vivo.
•HCP5 could function as competitive endogenous RNA (ceRNA) via directly sponging of miR-128-3p, which further regulates PLAGL2 expression.
Supplementary Information
Below is the link to the electronic supplementary material.

10565_2021_9628_MOESM1_ESM.jpg
Supplementary file1 PLAGL2 overexpression promoted proliferation of MM cell lines. NCI-H929 cells were transfected with PLAGL2 (pcDNA3.1-PLAGL2 overexpression plasmid) and corresponding negative control (the empty vector), and MM.1S cells were transfected with si-PLAGL2 and negative control si-NC. (A) Transfection efficiency was evaluated by Western blot. (B) Cell viability was evaluated by CCK-8 assay. (C and D) Cell proliferation was evaluated by colony formation assay (C) and EdU staining assay (D). (E) Cell cycle was evaluated by flow cytometry. * P < 0.05, ** P < 0.01, *** P < 0.001. Each experiment was repeated three times. (JPG 1329 KB)
Rights and permissions
About this article
Cite this article
Liu, Q., Ran, R., Song, M. et al. LncRNA HCP5 acts as a miR-128-3p sponge to promote the progression of multiple myeloma through activating Wnt/β‐catenin/cyclin D1 signaling via PLAGL2. Cell Biol Toxicol 38, 979–993 (2022). https://doi.org/10.1007/s10565-021-09628-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10565-021-09628-7