Abstract
Background
Proprotein convertase subtilisin/kexin 9 (PCSK9) serves a key regulatory function in the metabolism of low-density lipoprotein (LDL)-cholesterol (LDL-C) through interaction with the LDL receptor (LDLR) followed by its destruction that results in the elevation of the plasma levels of LDL-C. The aims of the present study were to separate and select a number of single-stranded DNA (ssDNA) aptamers against PCSK9 from a library pool (n > 1012) followed by their characterization.
Methods
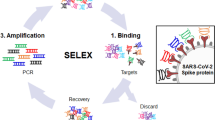
The aptamers obtained from the DNA-PCSK9 complexes which presented the highest affinity against PCSK9 were separated and selected using capillary electrophoresis evolution of ligands by exponential enrichment (CE-SELEX). The selected aptamers were amplified and cloned into a T/A vector. The plasmids from the positive clones were extracted and sequenced. The Mfold web server was used to predict the secondary structure of the aptamers.
Results
Following three rounds of CE-SELEX, the identified anti-PCSK9 ssDNA aptamers, namely aptamer 1 (AP-1) and aptamer 2 (AP-2), presented half maximal inhibitory concentrations of 325 and 327 nM, lowest dissociation constants of 294 and 323 nM, and most negative Gibbs free energy values of − 9.17 and − 8.28 kcal/mol, respectively.
Conclusion
The results indicated that the selected aptamers (AP-1 and AP-2) induced potent inhibitory effects against PCSK9. Further in vivo studies demand to find out AP-1 and AP-2 aptamers as suitable candidates, instead of antibodies, for using in therapeutic purposes in patients with hypercholesterolemia and cardiovascular disease.








Similar content being viewed by others
Data Availability
The datasets used and/or analyzed during the current study are included in this published article.
References
Levine GN, Lange RA, Bairey-Merz CN, Davidson RJ, Jamerson K, Mehta PK, et al. Meditation and cardiovascular risk reduction: a scientific statement from the American Heart Association. J Am Heart Assoc. 2017;6(10):e002218.
Sniderman AD, Islam S, Yusuf S, McQueen MJ. Discordance analysis of apolipoprotein B and non-high density lipoprotein cholesterol as markers of cardiovascular risk in the INTERHEART study. Atherosclerosis. 2012;225(2):444–9.
Defesche JC, Gidding SS, Harada-Shiba M, Hegele RA, Santos RD, Wierzbicki AS. Familial hypercholesterolaemia. Nat Rev Dis Primers. 2017;3:17093.
Wilson PW, D’Agostino RB, Levy D, Belanger AM, Silbershatz H, Kannel WB. Prediction of coronary heart disease using risk factor categories. Circulation. 1998;97(18):1837–47.
Nicholls SJ, Ballantyne CM, Barter PJ, Chapman MJ, Erbel RM, Libby P, et al. Effect of two intensive statin regimens on progression of coronary disease. N Engl J Med. 2011;365(22):2078–87.
Hovingh GK, Davidson MH, Kastelein JJ, O’connor AM. Diagnosis and treatment of familial hypercholesterolaemia. Eur Heart J. 2013;34(13):962–71.
Seidah NG, Benjannet S, Wickham L, Marcinkiewicz J, Jasmin SB, Stifani S, et al. The secretory proprotein convertase neural apoptosis-regulated convertase 1 (NARC-1): liver regeneration and neuronal differentiation. Proc Natl Acad Sci U S A. 2003;100(3):928–33.
Abifadel M, Varret M, Rabès J-P, Allard D, Ouguerram K, Devillers M, et al. Mutations in PCSK9 cause autosomal dominant hypercholesterolemia. Nat Genet. 2003;34(2):154–6.
Burke AC, Dron JS, Hegele RA, Huff MW. PCSK9: regulation and target for drug development for dyslipidemia. Annu Rev Pharmacol Toxicol. 2017;57:223–44.
Seidah NG, Awan Z, Chrétien M, Mbikay M. PCSK9: a key modulator of cardiovascular health. Circ Res. 2014;114(6):1022–36.
Dahagam C, Goud A, Abdelqader A, Hendrani A, Feinstein MJ, Qamar A, et al. PCSK9 inhibitors and their role in high-risk patients in reducing LDL cholesterol levels: evolocumab. Future Cardiol. 2016;12(2):139–48.
Wang X, Raghavan A, Chen T, Qiao L, Zhang Y, Ding Q, et al. CRISPR-Cas9 targeting of PCSK9 in human hepatocytes in vivo. Arterioscler Thromb Vasc Biol. 2016;36(5):783–6.
Gustafsen C, Kjolby M, Nyegaard M, Mattheisen M, Lundhede J, Buttenschøn H, et al. The hypercholesterolemia-risk gene SORT1 facilitates PCSK9 secretion. Cell Metab. 2014;19(2):310–8.
Dadu RT, Ballantyne CM. Lipid lowering with PCSK9 inhibitors. Nat Rev Cardiol. 2014;11(10):563–75.
He N-y, Li Q, Wu C-y, Ren Z, Gao Y, Pan L-h, et al. Lowering serum lipids via PCSK9-targeting drugs: current advances and future perspectives. Acta Pharmacol Sin. 2017;38(3):301.
Ellington AD, Szostak JW. In vitro selection of RNA molecules that bind specific ligands. Nature. 1990;346(6287):818–22.
Zhou J, Rossi J. Aptamers as targeted therapeutics: current potential and challenges. Nat Rev Drug Discov. 2017;16(3):181–202.
Liu J, You M, Pu Y, Liu H, Ye M, Tan W. Recent developments in protein and cell-targeted aptamer selection and applications. Curr Med Chem. 2011;18(27):4117–25.
Mosing RK, Mendonsa SD, Bowser MT. Capillary electrophoresis-SELEX selection of aptamers with affinity for HIV-1 reverse transcriptase. Anal Chem. 2005;77(19):6107–12.
Dong L, Tan Q, Ye W, Liu D, Chen H, Hu H, et al. Screening and identifying a novel ssDNA aptamer against alpha-fetoprotein using CE-SELEX. Sci Rep. 2015;5:15552.
Wang P, Yang Y, Hong H, Zhang Y, Cai W, Fang D. Aptamers as therapeutics in cardiovascular diseases. Curr Med Chem. 2011;18(27):4169–74.
Kaur H, Bruno JG, Kumar A, Sharma TK. Aptamers in the therapeutics and diagnostics pipelines. Theranostics. 2018;8:4016–32.
Povsic TJ, Vavalle JP, Alexander JH, Aberle LH, Zelenkofske SL, Becker RC, et al. Use of the REG1 anticoagulation system in patients with acute coronary syndromes undergoing percutaneous coronary intervention: results from the phase II RADAR-PCI study. EuroIntervention. 2014;10(4):431–8.
Jilma-Stohlawetz P, Gorczyca ME, Jilma B, Siller-Matula J, Gilbert JC, Knöbl P. Inhibition of von Willebrand factor by ARC1779 in patients with acute thrombotic thrombocytopenic purpura. Thromb Haemost. 2011;105(03):545–52.
Hamedani NS, Müller J. Capillary electrophoresis for the selection of DNA aptamers recognizing activated protein C. Methods Mol Biol. 2016;1380:61–75.
Kouhpayeh S, Hejazi Z, Khanahmad H, Rezaei A. Real-time PCR: an appropriate approach to confirm ssDNA generation from PCR product in SELEX process. Iran J Biotechnol. 2017;15(2):143–8.
Zuker M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003;31(13):3406–15.
Chan JC, Piper DE, Cao Q, Liu D, King C, Wang W, et al. A proprotein convertase subtilisin/kexin type 9 neutralizing antibody reduces serum cholesterol in mice and nonhuman primates. Proc Natl Acad Sci U SA. 2009;106(24):9820–5.
Liang H, Chaparro-Riggers J, Strop P, Geng T, Sutton JE, Tsai D, et al. Proprotein convertase substilisin/kexin type 9 antagonism reduces low-density lipoprotein cholesterol in statin-treated hypercholesterolemic nonhuman primates. J Pharmacol Exp Ther. 2012;340(2):228–36.
Heiger DN. High performance capillary electrophoresis: an introduction: a primer. Agilent Technologies; 2000.
Agarwal SK, Avery CL, Ballantyne CM, Catellier D, Nambi V, Saunders J, et al. Sources of variability in measurements of cardiac troponin T in a community-based sample: the atherosclerosis risk in communities study. Clin Chem. 2011;57(6):891–7.
Benjamin EJ, Blaha MJ, Chiuve SE, Cushman M, Das SR, et al. Heart disease and stroke statistics—2017 update: a report from the American Heart Association. Circulation. 2017;135(10):E146.
Cohen JC, Boerwinkle E, Mosley TH Jr, Hobbs HH. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N Engl J Med. 2006;354(12):1264–72.
Abifadel M, Elbitar S, El Khoury P, Ghaleb Y, Chémaly M, Moussalli M-L, et al. Living the PCSK9 adventure: from the identification of a new gene in familial hypercholesterolemia towards a potential new class of anticholesterol drugs. Curr Atheroscler Rep. 2014;16(9):439.
Seidah NG. The PCSK9 revolution and the potential of PCSK9-based therapies to reduce LDL-cholesterol. Glob Cardiol Sci Pract. 2017;1:e201702.
Chaudhary R, Garg J, Shah N, Sumner A. PCSK9 inhibitors: a new era of lipid lowering therapy. World J Cardiol. 2017;9(2):76–91.
Dunn MR, Jimenez RM, Chaput JC. Analysis of aptamer discovery and technology. Nat Rev Chem. 2017;1(10):0076.
Haghighi M, Khanahmad H, Palizban A. Selection and characterization of single-stranded DNA aptamers binding human B-cell surface protein CD20 by cell-SELEX. Molecules. 2018;23(4):e715.
Palizban AA, Salehi R, Nori N, Galehdari H. In vivo transfection rat small intestine K-cell with pGIP/Ins plasmid by DOTAP liposome. J Drug Target. 2007;15(5):351–7.
Stoekenbroek RM, Lambert G, Cariou B, Hovingh GK. Inhibiting PCSK9 - biology beyond LDL control. Nat Rev Endocrinol. 2018;15(1):52–62.
Acknowledgments
The authors acknowledge the Isfahan University of Medical Sciences for providing academic and laboratory supports.
Funding
The present study was supported by the Iran National Science Foundation (INSF grant no. 93016087) and the Isfahan University of Medical Sciences (project no. 395902).
Author information
Authors and Affiliations
Contributions
AAP conceived the study, designed the experiments, performed the capillary electrophoresis evolution of ligands by exponential enrichment, and analyzed the data. RS prepared the samples for capillary electrophoresis and performed the experiments. HK facilitated the molecular biology experiments.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict interests.
Ethics Approval and Consent to Participate
This study was approved by the Ethics Committee (no. 395902) of the Isfahan University of Medical Sciences, Iran.
Patient consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sattari, R., Palizban, A. & Khanahmad, H. Single-Strand DNA-Like Oligonucleotide Aptamer Against Proprotein Convertase Subtilisin/Kexin 9 Using CE-SELEX: PCSK9 Targeting Selection. Cardiovasc Drugs Ther 34, 475–485 (2020). https://doi.org/10.1007/s10557-020-06986-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10557-020-06986-y