Abstract
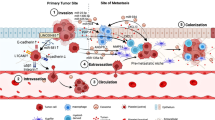
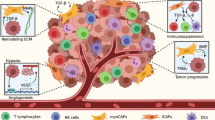
Colorectal cancer (CRC) patients frequently develop liver metastases, which are the major cause of cancer-related mortality. The molecular basis and management of colorectal liver metastases (CRLMs) remain a challenging clinical issue. Recent genomic evidence has demonstrated the liver tropism of CRC and the presence of a stricter evolutionary bottleneck in the liver as a target organ compared to lymph nodes. This bottleneck challenging CRC cells in the liver is organ-specific and requires adaptation not only at the genetic level, but also at the phenotypic level to crosstalk with the hepatic microenvironment. Here, we highlight the emerging evidence on the clonal evolution of CRLM and review recent insights into the molecular mechanisms orchestrating the bidirectional interactions between metastatic CRC cells and the unique liver microenvironment.


Similar content being viewed by others
References
Biller, L. H., & Schrag, D. (2021). Diagnosis and treatment of metastatic colorectal cancer: A review. JAMA, 325(7), 669–685. https://doi.org/10.1001/jama.2021.0106
Dekker, E., Tanis, P. J., Vleugels, J. L. A., Kasi, P. M., & Wallace, M. B. (2019). Colorectal cancer. Lancet, 394(10207), 1467–1480. https://doi.org/10.1016/S0140-6736(19)32319-0
Tsilimigras, D. I., Brodt, P., Clavien, P. A., Muschel, R. J., D'Angelica, M. I., Endo, I., et al. (2021). Liver metastases. Nature Reviews Disease Primers, 7(1), 27. https://doi.org/10.1038/s41572-021-00261-6
Milette, S., Sicklick, J. K., Lowy, A. M., & Brodt, P. (2017). Molecular pathways: Targeting the Microenvironment of liver metastases. Clinical Cancer Research, 23(21), 6390–6399. https://doi.org/10.1158/1078-0432.CCR-15-1636
Hess, K. R., Varadhachary, G. R., Taylor, S. H., Wei, W., Raber, M. N., Lenzi, R., et al. (2006). Metastatic patterns in adenocarcinoma. Cancer, 106(7), 1624–1633. https://doi.org/10.1002/cncr.21778
Paget, S. (1989). The distribution of secondary growths in cancer of the breast. The Lancet, 8(2), 98–101.
Chow, F. C., & Chok, K. S. (2019). Colorectal liver metastases: An update on multidisciplinary approach. World Journal of Hepatology, 11(2), 150–172. https://doi.org/10.4254/wjh.v11.i2.150
Vidal-Vanaclocha, F. (2008). The prometastatic microenvironment of the liver. Cancer Microenviron, 1(1), 113–129. https://doi.org/10.1007/s12307-008-0011-6
Mielgo, A., & Schmid, M. C. (2020). Liver tropism in cancer: The hepatic metastatic niche. Cold Spring Harbor Perspectives in Medicine, 10(3), a037259. https://doi.org/10.1101/cshperspect.a037259
Poisson, J., Lemoinne, S., Boulanger, C., Durand, F., Moreau, R., Valla, D., et al. (2017). Liver sinusoidal endothelial cells: Physiology and role in liver diseases. Journal of Hepatology, 66(1), 212–227. https://doi.org/10.1016/j.jhep.2016.07.009
Miskovic, D., & Mirnezami, R. (2021). Is complete mesocolic excision superior to conventional colectomy for colon cancer? Lancet Oncology, 22(7), 917–918. https://doi.org/10.1016/S1470-2045(21)00256-4
Fujita, S., Mizusawa, J., Kanemitsu, Y., Ito, M., Kinugasa, Y., Komori, K., et al. (2017). Mesorectal excision with or without lateral lymph node dissection for clinical stage II/III lower rectal cancer (JCOG0212): A multicenter, randomized controlled, noninferiority trial. Annals of Surgery, 266(2), 201–207. https://doi.org/10.1097/SLA.0000000000002212
Markowitz, S. D. (2017). Cancer bypasses the lymph nodes. Science, 357(6346), 35–36. https://doi.org/10.1126/science.aan8299
Siravegna, G., Lazzari, L., Crisafulli, G., Sartore-Bianchi, A., Mussolin, B., Cassingena, A., et al. (2018). Radiologic and genomic evolution of individual metastases during HER2 blockade in colorectal cancer. Cancer Cell, 34(1), 148–162. https://doi.org/10.1016/j.ccell.2018.06.004
Russo, M., Siravegna, G., Blaszkowsky, L. S., Corti, G., Crisafulli, G., Ahronian, L. G., et al. (2016). Tumor heterogeneity and lesion-specific response to targeted therapy in colorectal cancer. Cancer Discovery, 6(2), 147–153. https://doi.org/10.1158/2159-8290.CD-15-1283
Pietrantonio, F., Oddo, D., Gloghini, A., Valtorta, E., Berenato, R., Barault, L., et al. (2016). MET-driven resistance to dual EGFR and BRAF blockade may be overcome by switching from EGFR to MET inhibition in BRAF-mutated colorectal cancer. Cancer Discovery, 6(9), 963–971. https://doi.org/10.1158/2159-8290.CD-16-0297
Ryser, M. D., Mallo, D., Hall, A., Hardman, T., King, L. M., Tatishchev, S., et al. (2020). Minimal barriers to invasion during human colorectal tumor growth. Nature Communications, 11(1), 1280. https://doi.org/10.1038/s41467-020-14908-7
Hu, Z., Li, Z., Ma, Z., & Curtis, C. (2020). Multi-cancer analysis of clonality and the timing of systemic spread in paired primary tumors and metastases. Nature Genetics, 52(7), 701–708. https://doi.org/10.1038/s41588-020-0628-z
Naxerova, K., Reiter, J. G., Brachtel, E., Lennerz, J. K., van de Wetering, M., Rowan, A., et al. (2017). Origins of lymphatic and distant metastases in human colorectal cancer. Science, 357(6346), 55–60. https://doi.org/10.1126/science.aai8515
Househam, J., Heide, T., Cresswell, G. D., Spiteri, I., Kimberley, C., Zapata, L., et al. (2022). Phenotypic plasticity and genetic control in colorectal cancer evolution. Nature, 611(7937), 744–753. https://doi.org/10.1038/s41586-022-05311-x
Wu, Y., Yang, S., Ma, J., Chen, Z., Song, G., Rao, D., et al. (2021). Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discovery, 12(1), 134–153. https://doi.org/10.1158/2159-8290.CD-21-0316
Li, C., Sun, Y. D., Yu, G. Y., Cui, J. R., Lou, Z., Zhang, H., et al. (2020). Integrated omics of metastatic colorectal cancer. Cancer Cell, 38(5), 734–747 e739. https://doi.org/10.1016/j.ccell.2020.08.002
Liu, Y., Zhang, Q., Xing, B., Luo, N., Gao, R., Yu, K., et al. (2022). Immune phenotypic linkage between colorectal cancer and liver metastasis. Cancer Cell, 40(4), 424–437. https://doi.org/10.1016/j.ccell.2022.02.013
Obenauf, A. C., & Massague, J. (2015). Surviving at a distance: Organ-specific metastasis. Trends in Cancer, 1(1), 76–91. https://doi.org/10.1016/j.trecan.2015.07.009
Wei, Q., Ye, Z., Zhong, X., Li, L., Wang, C., Myers, R. E., et al. (2017). Multiregion whole-exome sequencing of matched primary and metastatic tumors revealed genomic heterogeneity and suggested polyclonal seeding in colorectal cancer metastasis. Annals of Oncology, 28(9), 2135–2141. https://doi.org/10.1093/annonc/mdx278
Dang, H. X., Krasnick, B. A., White, B. S., Grossman, J. G., Strand, M. S., Zhang, J., et al. (2020). The clonal evolution of metastatic colorectal cancer. Science Advances, 6(24), eaay9691. https://doi.org/10.1126/sciadv.aay9691
Alves, J. M., Prado-Lopez, S., Cameselle-Teijeiro, J. M., & Posada, D. (2019). Rapid evolution and biogeographic spread in a colorectal cancer. Nature Communications, 10(1), 5139. https://doi.org/10.1038/s41467-019-12926-8
Yachida, S., Jones, S., Bozic, I., Antal, T., Leary, R., Fu, B., et al. (2010). Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature, 467(7319), 1114–1117. https://doi.org/10.1038/nature09515
Campbell, P. J., Yachida, S., Mudie, L. J., Stephens, P. J., Pleasance, E. D., Stebbings, L. A., et al. (2010). The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature, 467(7319), 1109–1113. https://doi.org/10.1038/nature09460
Turajlic, S., & Swanton, C. (2016). Metastasis as an evolutionary process. Science, 352(6282), 169–175. https://doi.org/10.1126/science.aaf2784
Turajlic, S., Furney, S. J., Lambros, M. B., Mitsopoulos, C., Kozarewa, I., Geyer, F. C., et al. (2012). Whole genome sequencing of matched primary and metastatic acral melanomas. Genome Research, 22(2), 196–207. https://doi.org/10.1101/gr.125591.111
Gundem, G., Van Loo, P., Kremeyer, B., Alexandrov, L. B., Tubio, J. M. C., Papaemmanuil, E., et al. (2015). The evolutionary history of lethal metastatic prostate cancer. Nature, 520(7547), 353–357. https://doi.org/10.1038/nature14347
Gkountela, S., Castro-Giner, F., Szczerba, B. M., Vetter, M., Landin, J., Scherrer, R., et al. (2019). Circulating tumor cell clustering shapes DNA methylation to enable metastasis seeding. Cell, 176(1-2), 98–112. https://doi.org/10.1016/j.cell.2018.11.046
Aceto, N., Bardia, A., Miyamoto, D. T., Donaldson, M. C., Wittner, B. S., Spencer, J. A., et al. (2014). Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell, 158(5), 1110–1122. https://doi.org/10.1016/j.cell.2014.07.013
Reiter, J. G., Hung, W. T., Lee, I. H., Nagpal, S., Giunta, P., Degner, S., et al. (2020). Lymph node metastases develop through a wider evolutionary bottleneck than distant metastases. Nature Genetics, 52(7), 692–700. https://doi.org/10.1038/s41588-020-0633-2
Priestley, P., Baber, J., Lolkema, M. P., Steeghs, N., de Bruijn, E., Shale, C., et al. (2019). Pan-cancer whole-genome analyses of metastatic solid tumours. Nature, 575(7781), 210–216. https://doi.org/10.1038/s41586-019-1689-y
Bian, S., Hou, Y., Zhou, X., Li, X., Yong, J., Wang, Y., et al. (2018). Single-cell multiomics sequencing and analyses of human colorectal cancer. Science, 362(6418), 1060–1063. https://doi.org/10.1126/science.aao3791
Reiter, J. G., Makohon-Moore, A. P., Gerold, J. M., Heyde, A., Attiyeh, M. A., Kohutek, Z. A., et al. (2018). Minimal functional driver gene heterogeneity among untreated metastases. Science, 361(6406), 1033–1037. https://doi.org/10.1126/science.aat7171
Chen, H. N., Shu, Y., Liao, F., Liao, X., Zhang, H., Qin, Y., et al. (2021). Genomic evolution and diverse models of systemic metastases in colorectal cancer. Gut, 71(2), 322–332. https://doi.org/10.1136/gutjnl-2020-323703
Ishaque, N., Abba, M. L., Hauser, C., Patil, N., Paramasivam, N., Huebschmann, D., et al. (2018). Whole genome sequencing puts forward hypotheses on metastasis evolution and therapy in colorectal cancer. Nature Communications, 9(1), 4782. https://doi.org/10.1038/s41467-018-07041-z
Alves, J. M., Prado-Lopez, S., Tomas, L., Valecha, M., Estevez-Gomez, N., Alvarino, P., et al. (2022). Clonality and timing of relapsing colorectal cancer metastasis revealed through whole-genome single-cell sequencing. Cancer Letters, 543, 215767. https://doi.org/10.1016/j.canlet.2022.215767
van de Haar, J., Hoes, L. R., Roepman, P., Lolkema, M. P., Verheul, H. M. W., Gelderblom, H., et al. (2021). Limited evolution of the actionable metastatic cancer genome under therapeutic pressure. Nature Medicine, 27(9), 1553–1563. https://doi.org/10.1038/s41591-021-01448-w
Ahronian, L. G., Sennott, E. M., Van Allen, E. M., Wagle, N., Kwak, E. L., Faris, J. E., et al. (2015). Clinical acquired resistance to RAF inhibitor combinations in BRAF-mutant colorectal cancer through MAPK pathway alterations. Cancer Discovery, 5(4), 358–367. https://doi.org/10.1158/2159-8290.CD-14-1518
Bertotti, A., Papp, E., Jones, S., Adleff, V., Anagnostou, V., Lupo, B., et al. (2015). The genomic landscape of response to EGFR blockade in colorectal cancer. Nature, 526(7572), 263–267. https://doi.org/10.1038/nature14969
Missiaglia, E., Jacobs, B., D'Ario, G., Di Narzo, A. F., Soneson, C., Budinska, E., et al. (2014). Distal and proximal colon cancers differ in terms of molecular, pathological, and clinical features. Annals of Oncology, 25(10), 1995–2001. https://doi.org/10.1093/annonc/mdu275
Amri, R., Bordeianou, L. G., Sylla, P., & Berger, D. L. (2015). Variations in metastasis site by primary location in colon cancer. Journal of Gastrointestinal Surgery, 19(8), 1522–1527. https://doi.org/10.1007/s11605-015-2837-9
Riihimaki, M., Hemminki, A., Sundquist, J., & Hemminki, K. (2016). Patterns of metastasis in colon and rectal cancer. Science Reporter, 6, 29765. https://doi.org/10.1038/srep29765
Russolillo, N., Sperti, E., Langella, S., Menonna, F., Allieta, A., Di Maio, M., et al. (2020). Impact of primary tumor location on patterns of recurrence and survival of patients undergoing resection of liver metastases from colon cancer. HPB (Oxford), 22(1), 116–123. https://doi.org/10.1016/j.hpb.2019.05.014
Engstrand, J., Nilsson, H., Stromberg, C., Jonas, E., & Freedman, J. (2018). Colorectal cancer liver metastases - a population-based study on incidence, management and survival. BMC Cancer, 18(1), 78. https://doi.org/10.1186/s12885-017-3925-x
Yaeger, R., Chatila, W. K., Lipsyc, M. D., Hechtman, J. F., Cercek, A., Sanchez-Vega, F., et al. (2018). Clinical sequencing defines the genomic landscape of metastatic colorectal cancer. Cancer Cell, 33(1), 125–136. https://doi.org/10.1016/j.ccell.2017.12.004
Kim, S. C., Park, J. W., Seo, H. Y., Kim, M., Park, J. H., Kim, G. H., et al. (2022). Multifocal organoid capturing of colon cancer reveals pervasive intratumoral heterogenous drug responses. Advanced Sciencev (Weinh), 9(5), e2103360. https://doi.org/10.1002/advs.202103360
Banerjee, S., Zhang, X., Kuang, S., Wang, J., Li, L., Fan, G., et al. (2021). Comparative analysis of clonal evolution among patients with right- and left-sided colon and rectal cancer. Iscience, 24(7), 102718. https://doi.org/10.1016/j.isci.2021.102718
Sobral, D., Martins, M., Kaplan, S., Golkaram, M., Salmans, M., Khan, N., et al. (2022). Genetic and microenvironmental intra-tumor heterogeneity impacts colorectal cancer evolution and metastatic development. Communications Biology, 5(1), 937. https://doi.org/10.1038/s42003-022-03884-x
Joung, J. G., Oh, B. Y., Hong, H. K., Al-Khalidi, H., Al-Alem, F., Lee, H. O., et al. (2017). Tumor heterogeneity predicts metastatic potential in colorectal cancer. Clinical Cancer Research, 23(23), 7209–7216. https://doi.org/10.1158/1078-0432.CCR-17-0306
Malesci, A., Laghi, L., Bianchi, P., Delconte, G., Randolph, A., Torri, V., et al. (2007). Reduced likelihood of metastases in patients with microsatellite-unstable colorectal cancer. Clinical Cancer Research, 13(13), 3831–3839. https://doi.org/10.1158/1078-0432.CCR-07-0366
Li, X., Ramadori, P., Pfister, D., Seehawer, M., Zender, L., & Heikenwalder, M. (2021). The immunological and metabolic landscape in primary and metastatic liver cancer. Nature Reviews Cancer, 21(9), 541–557. https://doi.org/10.1038/s41568-021-00383-9
Berndt, N., Kolbe, E., Gajowski, R., Eckstein, J., Ott, F., Meierhofer, D., et al. (2021). Functional consequences of metabolic zonation in murine livers: Insights for an old Story. Hepatology, 73(2), 795–810. https://doi.org/10.1002/hep.31274
Bergers, G., & Fendt, S. M. (2021). The metabolism of cancer cells during metastasis. Nature Reviews Cancer, 21(3), 162–180. https://doi.org/10.1038/s41568-020-00320-2
Bu, P., Chen, K. Y., Xiang, K., Johnson, C., Crown, S. B., Rakhilin, N., et al. (2018). Aldolase B-mediated fructose metabolism drives metabolic reprogramming of colon cancer liver metastasis. Cell Metabolism, 27(6), 1249–1262. https://doi.org/10.1016/j.cmet.2018.04.003
Loo, J. M., Scherl, A., Nguyen, A., Man, F. Y., Weinberg, E., Zeng, Z., et al. (2015). Extracellular metabolic energetics can promote cancer progression. Cell, 160(3), 393–406. https://doi.org/10.1016/j.cell.2014.12.018
Garcia-Bermudez, J., Baudrier, L., La, K., Zhu, X. G., Fidelin, J., Sviderskiy, V. O., et al. (2018). Aspartate is a limiting metabolite for cancer cell proliferation under hypoxia and in tumours. Nature Cell Biology, 20(7), 775–781. https://doi.org/10.1038/s41556-018-0118-z
Yamaguchi, N., Weinberg, E. M., Nguyen, A., Liberti, M. V., Goodarzi, H., Janjigian, Y. Y., et al. (2019). PCK1 and DHODH drive colorectal cancer liver metastatic colonization and hypoxic growth by promoting nucleotide synthesis. Elife, 8, e52135. https://doi.org/10.7554/eLife.52135
Wang, K., Jiang, J., Lei, Y., Zhou, S., Wei, Y., & Huang, C. (2019). Targeting metabolic-redox circuits for cancer therapy. Trends in Biochemical Sciences, 44(5), 401–414. https://doi.org/10.1016/j.tibs.2019.01.001
Gao, W., Chen, L., Ma, Z., Du, Z., Zhao, Z., Hu, Z., et al. (2013). Isolation and phenotypic characterization of colorectal cancer stem cells with organ-specific metastatic potential. Gastroenterology, 145(3), 636–646. https://doi.org/10.1053/j.gastro.2013.05.049
Wu, Z., Wei, D., Gao, W., Xu, Y., Hu, Z., Ma, Z., et al. (2015). TPO-induced metabolic reprogramming drives liver metastasis of colorectal cancer CD110+ tumor-initiating Cells. Cell Stem Cell, 17(1), 47–59. https://doi.org/10.1016/j.stem.2015.05.016
Nguyen, A., Loo, J. M., Mital, R., Weinberg, E. M., Man, F. Y., Zeng, Z., et al. (2016). PKLR promotes colorectal cancer liver colonization through induction of glutathione synthesis. The Journal of Clinical Investigation, 126(2), 681–694. https://doi.org/10.1172/JCI83587
Anastasiou, D., Poulogiannis, G., Asara, J. M., Boxer, M. B., Jiang, J. K., Shen, M., et al. (2011). Inhibition of pyruvate kinase M2 by reactive oxygen species contributes to cellular antioxidant responses. Science, 334(6060), 1278–1283. https://doi.org/10.1126/science.1211485
Heymann, F., & Tacke, F. (2016). Immunology in the liver--from homeostasis to disease. Nature Reviews. Gastroenterology & Hepatology, 13(2), 88–110. https://doi.org/10.1038/nrgastro.2015.200
Crispe, I. N. (2014). Immune tolerance in liver disease. Hepatology, 60(6), 2109–2117. https://doi.org/10.1002/hep.27254
O'Leary, K. (2021). Liver metastases cultivate an immune desert. Nature Reviews. Cancer, 21(3), 143. https://doi.org/10.1038/s41568-021-00338-0
Canellas-Socias, A., Cortina, C., Hernando-Momblona, X., Palomo-Ponce, S., Mulholland, E. J., Turon, G., et al. (2022). Metastatic recurrence in colorectal cancer arises from residual EMP1(+) cells. Nature, 611(7936), 603–613. https://doi.org/10.1038/s41586-022-05402-9
Yu, J., Green, M. D., Li, S., Sun, Y., Journey, S. N., Choi, J. E., et al. (2021). Liver metastasis restrains immunotherapy efficacy via macrophage-mediated T cell elimination. Nature Medicine, 27(1), 152–164. https://doi.org/10.1038/s41591-020-1131-x
Song, J., Song, H., Wei, H., Sun, R., Tian, Z., & Peng, H. (2022). Requirement of RORalpha for maintenance and antitumor immunity of liver-resident natural killer cells/ILC1s. Hepatology, 75(5), 1181–1193. https://doi.org/10.1002/hep.32147
Tian, S., Chu, Y., Hu, J., Ding, X., Liu, Z., Fu, D., et al. (2022). Tumour-associated neutrophils secrete AGR2 to promote colorectal cancer metastasis via its receptor CD98hc-xCT. Gut, 71(12), 2489–2501. https://doi.org/10.1136/gutjnl-2021-325137
Wang, H., Zhang, B., Li, R., Chen, J., Xu, G., Zhu, Y., et al. (2022). KIAA1199 drives immune suppression to promote colorectal cancer liver metastasis by modulating neutrophil infiltration. Hepatology, 76(4), 967–981. https://doi.org/10.1002/hep.32383
Canellas-Socias, A., Cortina, C., Hernando-Momblona, X., Palomo-Ponce, S., Mulholland, E. J., Turon, G., et al. (2022). Metastatic recurrence in colorectal cancer arises from residual EMP1<sup>+</sup> cells. Nature, 611(7936), 603–613. https://doi.org/10.1038/s41586-022-05402-9
Gorelik, L., & Flavell, R. A. (2000). Abrogation of TGFbeta signaling in T cells leads to spontaneous T cell differentiation and autoimmune disease. Immunity, 12(2), 171–181. https://doi.org/10.1016/s1074-7613(00)80170-3
Tauriello, D. V. F., Palomo-Ponce, S., Stork, D., Berenguer-Llergo, A., Badia-Ramentol, J., Iglesias, M., et al. (2018). TGFbeta drives immune evasion in genetically reconstituted colon cancer metastasis. Nature, 554(7693), 538–543. https://doi.org/10.1038/nature25492
Ma, C., Han, M., Heinrich, B., Fu, Q., Zhang, Q., Sandhu, M., et al. (2018). Gut microbiome-mediated bile acid metabolism regulates liver cancer via NKT cells. Science, 360(6391). https://doi.org/10.1126/science.aan5931
Bertocchi, A., Carloni, S., Ravenda, P. S., Bertalot, G., Spadoni, I., Lo Cascio, A., et al. (2021). Gut vascular barrier impairment leads to intestinal bacteria dissemination and colorectal cancer metastasis to liver. Cancer Cell, 39(5), 708–724. https://doi.org/10.1016/j.ccell.2021.03.004
Bullman, S., Pedamallu, C. S., Sicinska, E., Clancy, T. E., Zhang, X., Cai, D., et al. (2017). Analysis of fusobacterium persistence and antibiotic response in colorectal cancer. Science, 358(6369), 1443–1448. https://doi.org/10.1126/science.aal5240
Spadoni, I., Zagato, E., Bertocchi, A., Paolinelli, R., Hot, E., Di Sabatino, A., et al. (2015). A gut-vascular barrier controls the systemic dissemination of bacteria. Science, 350(6262), 830–834. https://doi.org/10.1126/science.aad0135
Taieb, J., Svrcek, M., Cohen, R., Basile, D., Tougeron, D., & Phelip, J. M. (2022). Deficient mismatch repair/microsatellite unstable colorectal cancer: Diagnosis, prognosis and treatment. European Journal of Cancer, 175, 136–157. https://doi.org/10.1016/j.ejca.2022.07.020
Testa, U., Castelli, G., & Pelosi, E. (2020). Genetic alterations of metastatic colorectal cancer. Biomedicines, 8(10), 414. https://doi.org/10.3390/biomedicines8100414
Mouradov, D., Greenfield, P., Li, S., In, E. J., Storey, C., Sakthianandeswaren, A., et al. (2023). Onco-microbial community profiling identifies clinico-molecular and prognostic subtypes of colorectal cancer. Gastroenterology. https://doi.org/10.1053/j.gastro.2023.03.205
Moreno, E., & Basler, K. (2004). dMyc transforms cells into super-competitors. Cell, 117(1), 117–129. https://doi.org/10.1016/s0092-8674(04)00262-4
Flanagan, D. J., Pentinmikko, N., Luopajarvi, K., Willis, N. J., Gilroy, K., Raven, A. P., et al. (2021). NOTUM from Apc-mutant cells biases clonal competition to initiate cancer. Nature, 594(7863), 430–435. https://doi.org/10.1038/s41586-021-03525-z
Kanada, M., Bachmann, M. H., & Contag, C. H. (2016). Signaling by extracellular vesicles advances cancer hallmarks. Trends in Cancer, 2(2), 84–94. https://doi.org/10.1016/j.trecan.2015.12.005
Moya, I. M., Castaldo, S. A., Van den Mooter, L., Soheily, S., Sansores-Garcia, L., Jacobs, J., et al. (2019). Peritumoral activation of the hippo pathway effectors YAP and TAZ suppresses liver cancer in mice. Science, 366(6468), 1029–1034. https://doi.org/10.1126/science.aaw9886
Lee, J. W., Stone, M. L., Porrett, P. M., Thomas, S. K., Komar, C. A., Li, J. H., et al. (2019). Hepatocytes direct the formation of a pro-metastatic niche in the liver. Nature, 567(7747), 249–252. https://doi.org/10.1038/s41586-019-1004-y
Sethi, V., Kurtom, S., Tarique, M., Lavania, S., Malchiodi, Z., Hellmund, L., et al. (2018). Gut microbiota promotes tumor growth in mice by modulating immune response. Gastroenterology, 155(1), 33–37. https://doi.org/10.1053/j.gastro.2018.04.001
Bhattacharjee, S., Hamberger, F., Ravichandra, A., Miller, M., Nair, A., Affo, S., et al. (2021). Tumor restriction by type I collagen opposes tumor-promoting effects of cancer-associated fibroblasts. The Journal of Clinical Investigation, 131(11). https://doi.org/10.1172/JCI146987
Biffi, G., & Tuveson, D. A. (2021). Diversity and biology of cancer-associated fibroblasts. Physiological Reviews, 101(1), 147–176. https://doi.org/10.1152/physrev.00048.2019
Barbazan, J., & Matic Vignjevic, D. (2019). Cancer associated fibroblasts: Is the force the path to the dark side? Current Opinion in Cell Biology, 56, 71–79. https://doi.org/10.1016/j.ceb.2018.09.002
Qi, M., Fan, S., Huang, M., Pan, J., Li, Y., Miao, Q., et al. (2022). Targeting FAPα-expressing hepatic stellate cells overcomes resistance to antiangiogenics in colorectal cancer liver metastasis models. The Journal of Clinical Investigation, 132. https://doi.org/10.1172/JCI157399
Lampi, M. C., & Reinhart-King, C. A. (2018). Targeting extracellular matrix stiffness to attenuate disease: From molecular mechanisms to clinical trials. Science Translational Medicine, 10(422). https://doi.org/10.1126/scitranslmed.aao0475
Kalli, M., & Stylianopoulos, T. (2018). Defining the role of solid stress and matrix stiffness in cancer cell proliferation and metastasis. Frontiers in Oncology, 8, 55. https://doi.org/10.3389/fonc.2018.00055
Burnier, J. V., Wang, N., Michel, R. P., Hassanain, M., Li, S., Lu, Y., et al. (2011). Type IV collagen-initiated signals provide survival and growth cues required for liver metastasis. Oncogene, 30(35), 3766–3783. https://doi.org/10.1038/onc.2011.89
Nystrom, H., Tavelin, B., Bjorklund, M., Naredi, P., & Sund, M. (2015). Improved tumour marker sensitivity in detecting colorectal liver metastases by combined type IV collagen and CEA measurement. Tumour Biology, 36(12), 9839–9847. https://doi.org/10.1007/s13277-015-3729-z
Shen, Y., Wang, X., Lu, J., Salfenmoser, M., Wirsik, N. M., Schleussner, N., et al. (2020). Reduction of liver metastasis stiffness improves response to bevacizumab in metastatic colorectal cancer. Cancer Cell, 37(6), 800–817. https://doi.org/10.1016/j.ccell.2020.05.005
Liu, L., Mayes, P. A., Eastman, S., Shi, H., Yadavilli, S., Zhang, T., et al. (2015). The BRAF and MEK inhibitors dabrafenib and trametinib: Effects on immune function and in combination with immunomodulatory antibodies targeting PD-1, PD-L1, and CTLA-4. Clinical Cancer Research, 21(7), 1639–1651. https://doi.org/10.1158/1078-0432.CCR-14-2339
Pfirschke, C., Engblom, C., Rickelt, S., Cortez-Retamozo, V., Garris, C., Pucci, F., et al. (2016). Immunogenic chemotherapy sensitizes tumors to checkpoint blockade therapy. Immunity, 44(2), 343–354. https://doi.org/10.1016/j.immuni.2015.11.024
Zhang, L., Zhu, Z., Yan, H., Wang, W., Wu, Z., Zhang, F., et al. (2021). Creatine promotes cancer metastasis through activation of Smad2/3. Cell Metabolism, 33(6), 1111–1123. https://doi.org/10.1016/j.cmet.2021.03.009
Le, D. T., Durham, J. N., Smith, K. N., Wang, H., Bartlett, B. R., Aulakh, L. K., et al. (2017). Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science, 357(6349), 409–413. https://doi.org/10.1126/science.aan6733
Ebert, P. J. R., Cheung, J., Yang, Y., McNamara, E., Hong, R., Moskalenko, M., et al. (2016). MAP kinase inhibition promotes T cell and anti-tumor activity in combination with PD-L1 checkpoint blockade. Immunity, 44(3), 609–621. https://doi.org/10.1016/j.immuni.2016.01.024
Hodi, F. S., Lawrence, D., Lezcano, C., Wu, X., Zhou, J., Sasada, T., et al. (2014). Bevacizumab plus ipilimumab in patients with metastatic melanoma. Cancer Immunology Research, 2(7), 632–642. https://doi.org/10.1158/2326-6066.CIR-14-0053
Funding
This work was supported by (1) National Natural Science Foundation of China (82073246); (2) Sichuan Science and Technology Program (2022JDRC0049, 2022YFS0175); (3) 1.3.5 project for disciplines of excellence, West China Hospital, Sichuan University (ZYGD20006).
Author information
Authors and Affiliations
Contributions
Hai-Ning Chen and Zong-Guang Zhou designed the article, Qiu-Luo Liu and Huijie Zhou performed the literature search and drafted the work, Hai-Ning Chen and Zong-Guang Zhou revised the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, QL., Zhou, H., Zhou, ZG. et al. Colorectal cancer liver metastasis: genomic evolution and crosstalk with the liver microenvironment. Cancer Metastasis Rev 42, 575–587 (2023). https://doi.org/10.1007/s10555-023-10107-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-023-10107-0