Abstract
Purpose
BRCA1/2 genes are the two main genes associated with hereditary breast cancers (BC). In the present study, we explore clinical and molecular characteristics of BRCA-associated BC in relation to estrogen receptor (ER) status.
Methods
Three BC databases (DB) were evaluated: (i) Hadassah oncogenetics (n = 4826); (ii) METABRIC (n = 1980), and (iii) Nick-Zainal (n = 560). We evaluated age at diagnosis in BRCA positive (BP) and BRCA negative (BN) patients, and tested for mutational signature differences in cohort iii. mRNA differential expression analysis (DEA) and pathway analysis were performed in cohort ii.
Results
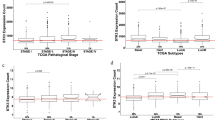
Age at diagnosis was lower in BP vs. BN tumors in all cohorts in the ER- group, and only in cohort i for the ER + group. Signature 3 was universal in BP BC, whereas several signatures were associated with ER status. Pathway analysis was performed between BP&BN, and was significant only in ER- tumors: the major activated pathways involved cancer-related processes and were highly significant. The most significant pathway was estrogen-mediated S-phase entry and the most activated upstream regulator was ERBB2.
Conclusion
Signature 3 was universal for all BP BC, while other signatures were associated with ER status. ER + BP& BN show similar genomic characteristics, ER- BP differed markedly from BN. This suggests that the initial carcinogenic process is universal for all BRCA carriers, but further insults lead to the development of two genomically distinct subtypes ER− and ER + . This may shed light on possible mechanisms involved in BP and carry preventive and therapeutic implications.



Similar content being viewed by others
Data availability
We analyzed published available databases.
Code availability
Not applicable.
References
Bleyer A, Welch HG (2012) Effect of three decades of screening mammography on breast-cancer incidence. N Engl J Med 367:1998–2005
Valencia OM et al (2017) The role of genetic testing in patients with breast cancer: a review. JAMA Surg 152:589–594
Alexandrov LB et al (2020) The repertoire of mutational signatures in human cancer. Nature 578:94–101
Polak P et al (2017) A mutational signature reveals alterations underlying deficient homologous recombination repair in breast cancer. Nat Genet 49:1476–1486
Maxwell KN et al (2017) BRCA locus-specific loss of heterozygosity in germline BRCA1 and BRCA2 carriers. Nat Commun. https://doi.org/10.1038/s41467-017-00388-9
Wang Z, Zhang J, Zhang Y, Deng Q, Liang H (2018) Expression and mutations of BRCA in breast cancer and ovarian cancer: evidence from bioinformatics analyses. Int J Mol Med 42:3542–3550
Hedenfalk I et al (2001) Gene-expression profiles in hereditary breast cancer. N Engl J Med 344:539–548
Wiggins GAR, Walker LC, Pearson JF (2020) Genome-wide gene expression analyses of BRCA1- and BRCA2-associated breast and ovarian tumours. Cancers (Basel) 12:1–17
Akbari V, Kallhor M, Akbari MT (2019) Transcriptome mining of non-BRCA1/A2 and BRCA1/A2 familial breast cancer. J Cell Biochem 120:575–583
Hartmann LC, Lindor NM (2016) The role of risk-reducing surgery in hereditary breast and ovarian cancer. N Engl J Med 374:454–468
Robson M et al (2017) Olaparib for metastatic breast cancer in patients with a germline BRCA mutation. N Engl J Med. https://doi.org/10.1056/NEJMoa1706450
Lord CJ, Ashworth A (2017) PARP inhibitors: synthetic lethality in the clinic. Science 355:1152–1158
Kedmi A et al (2022) Genetic anticipation of breast cancer among BRCA1/BRCA2 mutation carriers: a retrospective study. Int J Gynaecol Obstet. https://doi.org/10.1002/IJGO.14179
Curtis C et al (2012) The genomic and transcriptomic architecture of 2000 breast tumours reveals novel subgroups. Nature 486:346–352
Nik-Zainal S et al (2016) Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534:47–54
Team, R. C. R: A Language and Environment for Statistical Computing. (2018).
Ritchie ME et al (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43:e47
IPA, QIAGEN Redwood City. vol. 2016 http://www.qiagen.com/ingenuity.
Chen S, Parmigiani G (2007) Meta-analysis of BRCA1 and BRCA2 penetrance. J Clin Oncol 25:1329–1333
Atchley DP et al (2008) Clinical and pathologic characteristics of patients with BRCA-positive and BRCA-negative breast cancer. J Clin Oncol 26:4282–4288
COSMIC - Signatures of Mutational Processes in Human Cancer. https://cancer.sanger.ac.uk/signatures/signatures_v2/.
Gampenrieder SP et al (2021) Landscape of HER2-low metastatic breast cancer (MBC): results from the Austrian AGMT_MBC-Registry. Breast Cancer Res. https://doi.org/10.1186/s13058-021-01492-x
Zhang H, Katerji H, Turner BM, Audeh W, Hicks DG (2022) HER2-low breast cancers: incidence, HER2 staining patterns, clinicopathologic features, MammaPrint and BluePrint genomic profiles. Mod Pathol 35:1075–1082
Lim E et al (2009) Aberrant luminal progenitors as the candidate target population for basal tumor development in BRCA1 mutation carriers. Nat Med 15:907–913
Liu S et al (2008) BRCA1 regulates human mammary stem/progenitor cell fate. Proc Natl Acad Sci U S A 105:1680–1685
Halpern N et al (2017) Oncotype Dx recurrence score among BRCA1/2 germline mutation carriers with hormone receptors positive breast cancer. Int J Cancer 140:2145–2149
Barnes-Kedar I et al (2018) The yield of full BRCA1/2 genotyping in Israeli high-risk breast/ovarian cancer patients who do not carry the predominant mutations. Breast Cancer Res Treat 172:151–157
Funding
This work was partially supported by the Institutional Women's health foundation and the joint research fund between the Hebrew University faculty of Medicine and Hadassah University Hospital.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interest
The authors have not disclosed any competing interests.
Consent to participate
All patients signed consent to participate in research.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10549_2022_6851_MOESM1_ESM.tif
Supplementary file1 (TIF 245 kb)—Distribution of age at diagnosis grouped by BRCA1/2 and ER status. Histograms shown separately for each of the three databases
10549_2022_6851_MOESM2_ESM.tif
Supplementary file2 (TIF 1411 kb)—Ingenuity Pathway Analysis for ER positive: BRCA positive vs. BRCA negative tumors. Orange – pathways overrepresented in BRCA positive tumors, Blue – pathways underrepresented in BRCA positive tumors. Only pathways for which the direction of the enrichment could be determined are shown
10549_2022_6851_MOESM3_ESM.tif
Supplementary file3 (TIF 9547 kb)—Ingenuity Pathway Analysis for ER-negative tumors after exclusion of Her2 positive tumors BRCA positive vs. BRCA negative tumors. Orange – pathways overrepresented in BRCA positive tumors, Blue – pathways underrepresented in BRCA positive tumors pathways for which the direction of the enrichment could be determined are shown in gray.
10549_2022_6851_MOESM4_ESM.tif
Supplementary file4 (TIF 11149 kb)—Ingenuity Pathway Analysis for ER-positive tumors after exclusion of Her2 positive tumors BRCA positive vs. BRCA negative tumors. Orange – pathways overrepresented in BRCA positive tumors, Blue – pathways underrepresented in BRCA positive tumors pathways for which the direction of the enrichment could be determined are shown in gray.
10549_2022_6851_MOESM5_ESM.tif
Supplementary file5 (TIF 1669 kb)—Number of genetic aberrations for the BRCA and ER status groups. A. single nucleotide substitutions, B. indels, C. rearrangements. The number of aberrations is given in log 10 scale.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rosenberg, S., Devir, M., Kaduri, L. et al. Distinct breast cancer phenotypes in BRCA 1/2 carriers based on ER status. Breast Cancer Res Treat 198, 197–205 (2023). https://doi.org/10.1007/s10549-022-06851-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-022-06851-6