Abstract
Purpose
Radiation therapy (RT) for triple-negative breast cancer (TNBC) treatment is currently delivered in the adjuvant setting and is under investigation as a booster of neoadjuvant treatments. However, TNBC radioresistance remains an obstacle, so new biomarkers are needed to select patients for any integration of RT in the TNBC therapy sequence. MicroRNAs (miRs) are important regulators of gene expression, involved in cancer response to ionizing radiation (IR) and assessable by tumor tissue or liquid biopsy. This systematic review aimed to evaluate the relationships between miRs and response to radiation in TNBC, as well as their potential predictive and prognostic values.
Methods
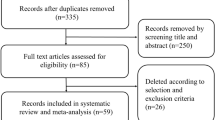
A thorough review of studies related to miRs and RT in TNBC was performed on PubMed, EMBASE, and Web of Science. We searched for original English articles that involved dysregulation of miRs in response to IR on TNBC-related preclinical and clinical studies. After a rigorous selection, 44 studies were chosen for further analysis.
Results
Thirty-five miRs were identified to be TNBC related, out of which 21 were downregulated, 13 upregulated, and 2 had a double-side expression in this cancer. Expression modulation of many of these miRs is radiosensitizing, among which miR-7, -27a, -34a, -122, and let-7 are most studied, still only in experimental models. The miRs reported as most influencing/reflecting TNBC response to IR are miR-7, -27a, -155, -205, -211, and -221, whereas miR-21, -33a, -139-5p, and -210 are associated with TNBC patient outcome after RT.
Conclusion
miRs are emerging biomarkers and radiosensitizers in TNBC, worth further investigation. Dynamic assessment of circulating miRs could improve monitoring and TNBC RT efficacy, which are of particular interest in the neoadjuvant and the high-risk patients’ settings.


Similar content being viewed by others
Abbreviations
- APBI:
-
Accelerated partial breast irradiation
- BC:
-
Breast cancer
- CSC:
-
Cancer stem cells
- DDR:
-
DNA damage repair
- DFS:
-
Disease-free survival
- DMFS:
-
Distant metastasis-free survival
- EMT:
-
Epithelial–mesenchymal transition
- ER:
-
Estrogen receptor
- FFPE:
-
Formalin-fixed paraffin-embedded
- IR:
-
Ionizing radiation
- HR:
-
Homologous recombination
- miR:
-
MicroRNA
- NACT:
-
Neoadjuvant chemotherapy
- OS:
-
Overall survival
- pCR:
-
Pathological complete response
- PTEN:
-
Phosphatase and tensin homolog
- RT:
-
Radiation therapy
- S1P:
-
Sphingosine-1-phosphatase
- TCGA:
-
The Cancer Genome Atlas
- TME:
-
Tumor microenvironment
- TNBC:
-
Triple-negative breast cancer
References
Siegel RL, Miller KD, Jemal A (2020) Cancer statistics. CA Cancer J Clin 70:7–30. https://doi.org/10.3322/caac.21590
Perou CM, Sørlie T, Eisen MB et al (2000) Molecular portraits of human breast tumours. Nature 406:747–752. https://doi.org/10.1038/35021093
Jones T, Neboori H, Wu H et al (2013) Are breast cancer subtypes prognostic for nodal involvement and associated with clinicopathologic features at presentation in early-stage breast cancer? Ann Surg Oncol 20:2866–2872. https://doi.org/10.1245/s10434-013-2994-6
Lin NU, Vanderplas A, Hughes ME et al (2012) Clinicopathologic features, patterns of recurrence, and survival among women with triple-negative breast cancer in the National Comprehensive Cancer Network. Cancer 118:5463–5472. https://doi.org/10.1002/cncr.27581
Cortazar P, Zhang L, Untch M et al (2014) Pathological complete response and long-term clinical benefit in breast cancer: the CTNeoBC pooled analysis. Lancet 384:164–172. https://doi.org/10.1016/S0140-6736(13)62422-8
Liedtke C, Mazouni C, Hess KR et al (2008) Response to neoadjuvant therapy and long-term survival in patients with triple-negative breast cancer. JCO 26:1275–1281. https://doi.org/10.1200/JCO.2007.14.4147
Loibl S, O’Shaughnessy J, Untch M et al (2018) Addition of the PARP inhibitor veliparib plus carboplatin or carboplatin alone to standard neoadjuvant chemotherapy in triple-negative breast cancer (BrighTNess): a randomised, phase 3 trial. Lancet Oncol 19:497–509. https://doi.org/10.1016/S1470-2045(18)30111-6
Schmid P, Adams S, Rugo HS et al (2018) Atezolizumab and Nab-paclitaxel in advanced triple-negative breast cancer. N Engl J Med 379:2108–2121. https://doi.org/10.1056/NEJMoa1809615
Corradini S, Krug D, Meattini I et al (2019) Preoperative radiotherapy: A paradigm shift in the treatment of breast cancer? A review of literature. Crit Rev Oncol Hematol 141:102–111. https://doi.org/10.1016/j.critrevonc.2019.06.003
Ahmed M, Jozsa F, Douek M (2021) A systematic review of neo-adjuvant radiotherapy in the treatment of breast cancer. Ecancermedicalscience 15:1175. https://doi.org/10.3332/ecancer.2021.1175
Weichselbaum RR, Liang H, Deng L, Fu Y-X (2017) Radiotherapy and immunotherapy: a beneficial liaison? Nat Rev Clin Oncol 14:365–379. https://doi.org/10.1038/nrclinonc.2016.211
Rodríguez-Ruiz ME, Vanpouille-Box C, Melero I et al (2018) Immunological mechanisms responsible for radiation-induced abscopal effect. Trends Immunol 39:644–655. https://doi.org/10.1016/j.it.2018.06.001
Mohammadi C, Gholamzadeh Khoei S, Fayazi N et al (2021) miRNA as promising theragnostic biomarkers for predicting radioresistance in cancer: A systematic review and meta-analysis. Crit Rev Oncol Hematol 157:103183. https://doi.org/10.1016/j.critrevonc.2020.103183
Kahraman M, Röske A, Laufer T et al (2018) MicroRNA in diagnosis and therapy monitoring of early-stage triple-negative breast cancer. Sci Rep 8:11584. https://doi.org/10.1038/s41598-018-29917-2
Ambros V (2004) The functions of animal microRNAs. Nature 431:350–355. https://doi.org/10.1038/nature02871
Bernichon E, Vallard A, Wang Q et al (2017) Genomic alterations and radioresistance in breast cancer: an analysis of the ProfiLER protocol. Ann Oncol 28:2773–2779. https://doi.org/10.1093/annonc/mdx488
Gee HE, Buffa FM, Harris AL et al (2015) MicroRNA-Related DNA Repair/cell-cycle genes independently associated with relapse after radiation therapy for early breast cancer. Int J Radiat Oncol Biol Phys 93:1104–1114. https://doi.org/10.1016/j.ijrobp.2015.08.046
Magbanua MJM, Swigart LB, Wu H-T et al (2021) Circulating tumor DNA in neoadjuvant-treated breast cancer reflects response and survival. Ann Oncol 32:229–239. https://doi.org/10.1016/j.annonc.2020.11.007
Wang H, Yee D (2019) I-SPY 2: a neoadjuvant adaptive clinical trial designed to improve outcomes in high-risk breast cancer. Curr Breast Cancer Rep 11:303–310. https://doi.org/10.1007/s12609-019-00334-2
Tomczak K, Czerwińska P, Wiznerowicz M (2015) The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol (Pozn) 19:A68–A77. https://doi.org/10.5114/wo.2014.47136
Welsh J (2013) Chapter 40 - Animal Models for Studying Prevention and Treatment of Breast Cancer. In: Conn PM (ed) Animal Models for the Study of Human Disease. Academic Press, Boston, pp 997–1018
Wang B, Wang H, Yang Z (2012) MiR-122 inhibits cell proliferation and tumorigenesis of breast cancer by targeting IGF1R. PLoS ONE 7:e47053. https://doi.org/10.1371/journal.pone.0047053
Perez-Añorve IX, la Rosa CHG-D, Soto-Reyes E et al (2019) New insights into radioresistance in breast cancer identify a dual function of miR-122 as a tumor suppressor and oncomiR. Mol Oncol 13:1249–1267. https://doi.org/10.1002/1878-0261.12483
Sun Q, Liu T, Yuan Y et al (2015) MiR-200c inhibits autophagy and enhances radiosensitivity in breast cancer cells by targeting UBQLN1. Int J Cancer 136:1003–1012. https://doi.org/10.1002/ijc.29065
Su Y, Shih P-H, Lee W-H et al (2017) Antrodia cinnamomea sensitizes radio-/chemo-therapy of cancer stem-like cells by modulating microRNA expression. J Ethnopharmacol 207:47–56. https://doi.org/10.1016/j.jep.2017.06.004
Masoudi-Khoram N, Abdolmaleki P, Hosseinkhan N et al (2020) Differential miRNAs expression pattern of irradiated breast cancer cell lines is correlated with radiation sensitivity. Sci Rep 10:9054. https://doi.org/10.1038/s41598-020-65680-z
Wang B, Zheng J, Li R et al (2019) Long noncoding RNA LINC02582 acts downstream of miR-200c to promote radioresistance through CHK1 in breast cancer cells. Cell Death Dis 10:1–15. https://doi.org/10.1038/s41419-019-1996-0
Zhang P, Wang L, Rodriguez-Aguayo C et al (2014) miR-205 acts as a tumour radiosensitizer by targeting ZEB1 and Ubc13. Nat Commun 5:5671. https://doi.org/10.1038/ncomms6671
Liang Z, Ahn J, Guo D et al (2013) MicroRNA-302 replacement therapy sensitizes breast cancer cells to ionizing radiation. Pharm Res 30:1008–1016. https://doi.org/10.1007/s11095-012-0936-9
Anastasov N, Hirmer E, Klenner M et al (2020) MEK1 Inhibitor combined with irradiation reduces migration of breast cancer cells including miR-221 and ZEB1 EMT Marker Expression. Cancers 12:3760. https://doi.org/10.3390/cancers12123760
Luo J, Chen J, He L (2015) mir-129–5p attenuates irradiation-induced autophagy and decreases radioresistance of breast cancer cells by targeting HMGB1. Med Sci Monit 21:4122–4129. https://doi.org/10.12659/msm.896661
Koo T, Cho BJ, Kim DH et al (2017) MicroRNA-200c increases radiosensitivity of human cancer cells with activated EGFR-associated signaling. Oncotarget 8:65457–65468. https://doi.org/10.18632/oncotarget.18924
Lee KM, Choi EJ, Kim IA (2011) microRNA-7 increases radiosensitivity of human cancer cells with activated EGFR-associated signaling. Radiother Oncol 101:171–176. https://doi.org/10.1016/j.radonc.2011.05.050
Wu J, Sun Z, Sun H, Li Y (2018) MicroRNA-27a promotes tumorigenesis via targeting AKT in triple negative breast cancer. Mol Med Rep 17:562–570. https://doi.org/10.3892/mmr.2017.7886
Yu L, Yang Y, Hou J et al (2015) MicroRNA-144 affects radiotherapy sensitivity by promoting proliferation, migration and invasion of breast cancer cells. Oncol Rep 34:1845–1852. https://doi.org/10.3892/or.2015.4173
Huang X, Taeb S, Jahangiri S et al (2013) miRNA-95 mediates radioresistance in tumors by targeting the sphingolipid phosphatase SGPP1. Cancer Res 73:6972–6986. https://doi.org/10.1158/0008-5472.CAN-13-1657
Bonnaud S, Niaudet C, Legoux F et al (2010) Sphingosine-1-Phosphate Activates the AKT pathway to protect small intestines from radiation-induced endothelial apoptosis. Cancer Res 70:9905–9915
Levine AJ, Hu W, Feng Z (2006) The P53 pathway: what questions remain to be explored? Cell Death Differ 13:1027–1036. https://doi.org/10.1038/sj.cdd.4401910
Wong MYW, Yu Y, Walsh WR, Yang J-L (2011) microRNA-34 family and treatment of cancers with mutant or wild-type p53 (Review). Int J Oncol 38:1189–1195. https://doi.org/10.3892/ijo.2011.970
He X, He L, Hannon GJ (2007) The guardian’s little helper: microRNAs in the p53 tumor suppressor network. Cancer Res 67:11099–11101. https://doi.org/10.1158/0008-5472.CAN-07-2672
Lodygin D, Tarasov V, Epanchintsev A et al (2008) Inactivation of miR-34a by aberrant CpG methylation in multiple types of cancer. Cell Cycle 7:2591–2600. https://doi.org/10.4161/cc.7.16.6533
Kato M, Paranjape T, Ullrich R et al (2009) The mir-34 microRNA is required for the DNA damage response in vivo in C. elegans and in vitro in human breast cancer cells. Oncogene 28:2419–2424. https://doi.org/10.1038/onc.2009.106
Klein HL (2008) The consequences of Rad51 overexpression for normal and tumor cells. DNA Repair (Amst) 7:686–693. https://doi.org/10.1016/j.dnarep.2007.12.008
Bayraktar R, Van Roosbroeck K (2018) miR-155 in cancer drug resistance and as target for miRNA-based therapeutics. Cancer Metastasis Rev 37:33–44. https://doi.org/10.1007/s10555-017-9724-7
Gasparini P, Lovat F, Fassan M et al (2014) Protective role of miR-155 in breast cancer through RAD51 targeting impairs homologous recombination after irradiation. PNAS 111:4536–4541. https://doi.org/10.1073/pnas.1402604111
Olea-Flores M, Zuñiga-Eulogio MD, Mendoza-Catalán MA et al (2019) Extracellular-signal regulated kinase: a central molecule driving epithelial-mesenchymal transition in cancer. Int J Mol Sci. https://doi.org/10.3390/ijms20122885
Tripathi K, Garg M (2018) Mechanistic regulation of epithelial-to-mesenchymal transition through RAS signaling pathway and therapeutic implications in human cancer. J Cell Commun Signal 12:513–527. https://doi.org/10.1007/s12079-017-0441-3
Shah MY, Calin GA (2011) MicroRNAs miR-221 and miR-222: a new level of regulation in aggressive breast cancer. Genome Med 3:56. https://doi.org/10.1186/gm272
Howe EN, Cochrane DR, Richer JK (2011) Targets of miR-200c mediate suppression of cell motility and anoikis resistance. Breast Cancer Res 13:R45. https://doi.org/10.1186/bcr2867
Lin J, Liu C, Gao F et al (2013) miR-200c enhances radiosensitivity of human breast cancer cells. J Cell Biochem 114:606–615. https://doi.org/10.1002/jcb.24398
Brunner TB, Kunz-Schughart LA, Grosse-Gehling P, Baumann M (2012) Cancer stem cells as a predictive factor in radiotherapy. Semin Radiat Oncol 22:151–174. https://doi.org/10.1016/j.semradonc.2011.12.003
Lytle NK, Barber AG, Reya T (2018) Stem cell fate in cancer growth, progression and therapy resistance. Nat Rev Cancer 18:669–680. https://doi.org/10.1038/s41568-018-0056-x
Griñán-Lisón C, Olivares-Urbano MA, Jiménez G et al (2020) miRNAs as radio-response biomarkers for breast cancer stem cells. Mol Oncol 14:556–570. https://doi.org/10.1002/1878-0261.12635
Sun H, Ding C, Zhang H, Gao J (2016) Let-7 miRNAs sensitize breast cancer stem cells to radiation-induced repression through inhibition of the cyclin D1/Akt1/Wnt1 signaling pathway. Mol Med Rep 14:3285–3292. https://doi.org/10.3892/mmr.2016.5656
Wang L, Yuan C, Lv K et al (2013) Lin28 mediates radiation resistance of breast cancer cells via regulation of caspase, H2A.X and Let-7 Signaling. PLoS ONE 8:e67373. https://doi.org/10.1371/journal.pone.0067373
Many AM, Brown AMC (2014) Both canonical and non-canonical Wnt signaling independently promote stem cell growth in mammospheres. PLoS ONE 9:e101800. https://doi.org/10.1371/journal.pone.0101800
Troschel FM, Böhly N, Borrmann K et al (2018) miR-142-3p attenuates breast cancer stem cell characteristics and decreases radioresistance in vitro. Tumour Biol 40:1010428318791887. https://doi.org/10.1177/1010428318791887
Smith AG, Macleod KF (2019) Autophagy, cancer stem cells and drug resistance. J Pathol 247:708–718. https://doi.org/10.1002/path.5222
Yi H, Liang B, Jia J et al (2013) Differential roles of miR-199a-5p in radiation-induced autophagy in breast cancer cells. FEBS Lett 587:436–443. https://doi.org/10.1016/j.febslet.2012.12.027
Metheetrairut C, Adams BD, Nallur S et al (2017) cel-mir-237 and its homologue, hsa-miR-125b, modulate the cellular response to ionizing radiation. Oncogene 36:512–524. https://doi.org/10.1038/onc.2016.222
Jang MH, Kim HJ, Gwak JM et al (2017) Prognostic value of microRNA-9 and microRNA-155 expression in triple-negative breast cancer. Hum Pathol 68:69–78. https://doi.org/10.1016/j.humpath.2017.08.026
Kim ES, Choi YE, Hwang SJ et al (2016) IL-4, a direct target of miR-340/429, is involved in radiation-induced aggressive tumor behavior in human carcinoma cells. Oncotarget 7:86836–86856. https://doi.org/10.18632/oncotarget.13561
Wolfe AR, Bambhroliya A, Reddy JP et al (2016) MiR-33a decreases radiation sensitivity to high-density lipoprotein in breast cancer. Int J Radiat Oncol Biol Phys 95:791–799. https://doi.org/10.1016/j.ijrobp.2016.01.025
Pajic M, Froio D, Daly S et al (2018) miR-139-5p Modulates radiotherapy resistance in breast cancer by repressing multiple gene networks of DNA Repair and ROS Defense. Cancer Res 78:501–515. https://doi.org/10.1158/0008-5472.CAN-16-3105
Anastasov N, Höfig I, Vasconcellos IG et al (2012) Radiation resistance due to high expression of miR-21 and G2/M checkpoint arrest in breast cancer cells. Radiat Oncol 7:206. https://doi.org/10.1186/1748-717X-7-206
Li Y, Liang Y, Sang Y et al (2018) MiR-770 suppresses the chemo-resistance and metastasis of triple negative breast cancer via direct targeting of STMN1. Cell Death Dis 9:1–12. https://doi.org/10.1038/s41419-017-0030-7
Bouchie A (2013) First microRNA mimic enters clinic. Nat Biotechnol 31:577–577. https://doi.org/10.1038/nbt0713-577
Kristen AV, Ajroud-Driss S, Conceição I et al (2018) Patisiran, an RNAi therapeutic for the treatment of hereditary transthyretin-mediated amyloidosis. Neurodegenerative Disease Management 9:5–23. https://doi.org/10.2217/nmt-2018-0033
Hanna J, Hossain GS, Kocerha J (2019) The Potential for microRNA Therapeutics and Clinical Research. Front Genet. https://doi.org/10.3389/fgene.2019.00478
Janssen HLA, Reesink HW, Lawitz EJ et al (2013) Treatment of HCV infection by targeting microRNA. N Engl J Med 368:1685–1694. https://doi.org/10.1056/NEJMoa1209026
van Zandwijk N, Pavlakis N, Kao SC et al (2017) Safety and activity of microRNA-loaded minicells in patients with recurrent malignant pleural mesothelioma: a first-in-man, phase 1, open-label, dose-escalation study. Lancet Oncol 18:1386–1396. https://doi.org/10.1016/S1470-2045(17)30621-6
Jin X, Chen Y, Chen H et al (2017) Evaluation of Tumor-Derived Exosomal miRNA as potential diagnostic biomarkers for early-stage non-small cell lung cancer using next-generation sequencing. Clin Cancer Res 23:5311–5319. https://doi.org/10.1158/1078-0432.CCR-17-0577
Schwarzenbach H, Nishida N, Calin GA, Pantel K (2014) Clinical relevance of circulating cell-free microRNAs in cancer. Nat Rev Clin Oncol 11:145–156. https://doi.org/10.1038/nrclinonc.2014.5
Zampetaki A, Mayr M (2012) Analytical challenges and technical limitations in assessing circulating MiRNAs. Thromb Haemost 108:592–598. https://doi.org/10.1160/TH12-02-0097
Faraldi M, Gomarasca M, Sansoni V et al (2019) Normalization strategies differently affect circulating miRNA profile associated with the training status. Sci Rep 9:1584. https://doi.org/10.1038/s41598-019-38505-x
Zhao H, Shen J, Medico L et al (2010) A pilot study of circulating miRNAs as potential biomarkers of early stage breast cancer. PLoS ONE 5:e13735. https://doi.org/10.1371/journal.pone.0013735
Meder B, Backes C, Haas J et al (2014) Influence of the confounding factors age and sex on microRNA profiles from peripheral blood. Clin Chem 60:1200–1208. https://doi.org/10.1373/clinchem.2014.224238
Chiarantini L, Cerasi A, Fraternale A et al (2005) Comparison of novel delivery systems for antisense peptide nucleic acids. J Control Release 109:24–36. https://doi.org/10.1016/j.jconrel.2005.09.013
Meng Z, Lu M (2017) RNA Interference-Induced Innate Immunity, Off-Target Effect, or Immune Adjuvant? Front Immunol 8:331. https://doi.org/10.3389/fimmu.2017.00331
Wang Z (2011) The guideline of the design and validation of MiRNA mimics. Methods Mol Biol 676:211–223. https://doi.org/10.1007/978-1-60761-863-8_15
Mei Z, Su T, Ye J et al (2015) The miR-15 family enhances the radiosensitivity of breast cancer cells by targeting G2 checkpoints. Radiat Res 183:196–207. https://doi.org/10.1667/RR13784.1
Leung C-M, Chen T-W, Li S-C et al (2014) MicroRNA expression profiles in human breast cancer cells after multifraction and single-dose radiation treatment. Oncol Rep 31:2147–2156. https://doi.org/10.3892/or.2014.3089
Radulovic V, Heider T, Richter S et al (2017) Differential response of normal and transformed mammary epithelial cells to combined treatment of anti-miR-21 and radiation. Int J Radiat Biol 93:361–372. https://doi.org/10.1080/09553002.2016.1266057
Zhang X, Li Y, Wang D, Wei X (2017) miR-22 suppresses tumorigenesis and improves radiosensitivity of breast cancer cells by targeting Sirt1. Biol Res 50:27. https://doi.org/10.1186/s40659-017-0133-8
Ren Y, Fu F, Han J (2015) MiR-27a modulates radiosensitivity of triple-negative breast cancer (TNBC) cells by targeting CDC27. Med Sci Monit 21:1297–1303. https://doi.org/10.12659/MSM.893974
Lai Y, Chen Y, Lin Y, Ye L (2018) Down-regulation of LncRNA CCAT1 enhances radiosensitivity via regulating miR-148b in breast cancer. Cell Biol Int 42:227–236. https://doi.org/10.1002/cbin.10890
Farsinejad S, Rahaie M, Alizadeh AM et al (2016) Expression of the circulating and the tissue microRNAs after surgery, chemotherapy, and radiotherapy in mice mammary tumor. Tumor Biol 37:14225–14234. https://doi.org/10.1007/s13277-016-5292-7
Zhang Z-Q, Cao Z, Liu C et al (2016) MiRNA-Embedded ShRNAs for Radiation-Inducible LGMN knockdown and the antitumor effects on breast cancer. PLoS ONE 11:e0163446. https://doi.org/10.1371/journal.pone.0163446
Bing Z, Tian J, Zhang J et al (2016) An Integrative Model of miRNA and mRNA expression signature for patients of breast invasive carcinoma with radiotherapy prognosis. Cancer Biother Radiopharm 31:253–260. https://doi.org/10.1089/cbr.2016.2059
Moskwa P, Buffa FM, Pan Y et al (2011) miR-182-Mediated Downregulation of BRCA1 Impacts DNA Repair and Sensitivity to PARP Inhibitors. Mol Cell 41:210–220. https://doi.org/10.1016/j.molcel.2010.12.005
Shi R, Wu P, Liu M, et al (2020) <p>Knockdown of lncRNA <em>PCAT6</em> Enhances Radiosensitivity in Triple-Negative Breast Cancer Cells by Regulating<em> miR-185–5p</em>/<em>TPD52</em> Axis</p>. In: OncoTargets and Therapy. https://www.dovepress.com/knockdown-of-lncrna-pcat6-enhances-radiosensitivity-in-triple-negative-peer-reviewed-article-OTT. Accessed 3 Feb 2021
Liu L, Zhu Y, Liu A-M et al (2019) Long noncoding RNA LINC00511 involves in breast cancer recurrence and radioresistance by regulating STXBP4 expression via miR-185. Eur Rev Med Pharmacol Sci 23:7457–7468. https://doi.org/10.26355/eurrev_201909_18855
Hu X, Ding D, Zhang J, Cui J (2019) Knockdown of lncRNA HOTAIR sensitizes breast cancer cells to ionizing radiation through activating miR-218. Biosci Rep. https://doi.org/10.1042/BSR20181038
Fabris L, Berton S, Citron F et al (2016) Radiotherapy-induced miR-223 prevents relapse of breast cancer by targeting the EGF pathway. Oncogene 35:4914–4926. https://doi.org/10.1038/onc.2016.23
Zhang S, Wang B, Xiao H et al (2020) LncRNA HOTAIR enhances breast cancer radioresistance through facilitating HSPA1A expression via sequestering miR-449b-5p. Thoracic Cancer 11:1801–1816. https://doi.org/10.1111/1759-7714.13450
Huang X, Taeb S, Jahangiri S et al (2015) miR-620 promotes tumor radioresistance by targeting 15-hydroxyprostaglandin dehydrogenase (HPGD). Oncotarget 6:22439–22451. https://doi.org/10.18632/oncotarget.4210
Yang B, Kuai F, Chen Z et al (2020) miR-634 decreases the radioresistance of human breast cancer cells by targeting STAT3. Cancer Biother Radiopharm 35:241–248. https://doi.org/10.1089/cbr.2019.3220
Bakhtari N, Mozdarani H, Salimi M, Omranipour R (2021) Association study of miR-22 and miR-335 expression levels and G2 assay related inherent radiosensitivity in peripheral blood of ductal carcinoma breast cancer patients. Neoplasma 68:190–199. https://doi.org/10.4149/neo_2020_200225N185
Acknowledgements
The authors would like to thank Ms. Myrna Perlmutter for her checking in English writing.
Funding
INCa (PROUST Project).
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
To, N.H., Nguyen, H.Q., Thiolat, A. et al. Radiation therapy for triple-negative breast cancer: emerging role of microRNAs as biomarkers and radiosensitivity modifiers. A systematic review. Breast Cancer Res Treat 193, 265–279 (2022). https://doi.org/10.1007/s10549-022-06533-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-022-06533-3