Abstract
Purpose
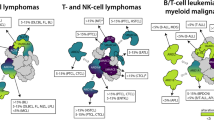
Chromatin remodeling plays an essential role in regulating transcriptional networks and timing of gene expression. Chromatin remodelers such as SWItch/Sucrose Non-Fermentable (SWI/SNF) harbor many protein components, with the catalytic subunit providing ATPase activity to displace histones along or from the DNA molecules, and associated subunits ensuring tissue specificity and transcriptional or co-transcriptional activities. Mutations in several of the SWI/SNF subunits have been linked to cancer. Here, we investigate between SMARCD3/Baf60c expression and hormone-positive (ER+) breast cancer.
Methods
The level of SMARCD3 was detected by immunohistochemistry in breast cancer patient samples, and expression levels of SMARCD1, SMARCD2, and SMARCD3 were investigated using publicly available datasets from large cohorts of breast cancer patients. Using molecular biology and microscopy, we interrogated the cellular consequences of lower SMARCD3 expression.
Results
Lower proliferation rates were observed in SMARCD3-depleted cells, which reflects a failure of the cell cycle progression and an increase in endoreplication. In the absence of SMARCD3, p21 accumulates in cells, but does not halt the cell cycle, and DNA damage accumulates and remains unrepaired.
Conclusion
Taken together, our data begin to explain why ER+ breast cancer patients with low-SMARCD3 expressing tumors exhibit reduced survival rates compared to patients expressing normal or higher levels of SMARCD3. SMARCD3 might act as a tumor suppressor through regulation of cell cycle checkpoints and could be a reliable and specific breast cancer prognostic biomarker.




Similar content being viewed by others
References
Tang L, Nogales E, Ciferri C (2010) Structure and function of SWI/SNF chromatin remodeling complexes and mechanistic implications for transcription. Prog Biophys Mol Biol 102(2–3):122–128
Winston F, Carlson M (1992) Yeast SNF/SWI transcriptional activators and the SPT/SIN chromatin connection. Trends Genet 8(11):387–391
Clapier CR, Cairns BR (2009) The biology of chromatin remodeling complexes. Annu Rev Biochem 78:273–304
Kadoch C, Crabtree GR (2015) Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci Adv 1(5):e1500447
Dutta A, Sardiu M, Gogol M, Gilmore J, Zhang D, Florens L, Abmayr SM, Washburn MP, Workman JL (2017) Composition and function of mutant Swi/Snf complexes. Cell Rep 18(9):2124–2134
Sen P, Luo J, Hada A, Hailu SG, Dechassa ML, Persinger J, Brahma S, Paul S, Ranish J, Bartholomew B (2017) Loss of Snf5 induces formation of an aberrant SWI/SNF complex. Cell Rep 18(9):2135–2147
Wang X, Lee RS, Alver BH, Haswell JR, Wang S, Mieczkowski J, Drier Y, Gillespie SM, Archer TC, Wu JN et al (2017) SMARCB1-mediated SWI/SNF complex function is essential for enhancer regulation. Nat Genet 49(2):289–295
Goljanek-Whysall K, Mok GF, Fahad Alrefaei A, Kennerley N, Wheeler GN, Munsterberg A (2014) myomiR-dependent switching of BAF60 variant incorporation into Brg1 chromatin remodeling complexes during embryo myogenesis. Development 141(17):3378–3387
Witzel M, Petersheim D, Fan Y, Bahrami E, Racek T, Rohlfs M, Puchalka J, Mertes C, Gagneur J, Ziegenhain C et al (2017) Chromatin-remodeling factor SMARCD2 regulates transcriptional networks controlling differentiation of neutrophil granulocytes. Nat Genet 49(5):742–752
Debril MB, Gelman L, Fayard E, Annicotte JS, Rocchi S, Auwerx J (2004) Transcription factors and nuclear receptors interact with the SWI/SNF complex through the BAF60c subunit. J Biol Chem 279(16):16677–16686
Flajollet S, Lefebvre B, Cudejko C, Staels B, Lefebvre P (2007) The core component of the mammalian SWI/SNF complex SMARCD3/BAF60c is a coactivator for the nuclear retinoic acid receptor. Mol Cell Endocrinol 270(1–2):23–32
Acevedo N, Reinius LE, Vitezic M, Fortino V, Soderhall C, Honkanen H, Veijola R, Simell O, Toppari J, Ilonen J et al (2015) Age-associated DNA methylation changes in immune genes, histone modifiers and chromatin remodeling factors within 5 years after birth in human blood leukocytes. Clin Epigenet 7:34
Sohni A, Mulas F, Ferrazzi F, Luttun A, Bellazzi R, Huylebroeck D, Ekker SC, Verfaillie CM (2012) TGFbeta1-induced Baf60c regulates both smooth muscle cell commitment and quiescence. PLoS ONE 7(10):e47629
Stirzaker C, Zotenko E, Song JZ, Qu W, Nair SS, Locke WJ, Stone A, Armstong NJ, Robinson MD, Dobrovic A et al (2015) Methylome sequencing in triple-negative breast cancer reveals distinct methylation clusters with prognostic value. Nat Commun 6:5899
Gregoire JM, Fleury L, Salazar-Cardozo C, Alby F, Masson V, Arimondo PB, Ausseil F (2016) Identification of epigenetic factors regulating the mesenchyme to epithelium transition by RNA interference screening in breast cancer cells. BMC Cancer 16:700
Lee RS, Roberts CW (2013) Linking the SWI/SNF complex to prostate cancer. Nat Genet 45(11):1268–1269
Wu Q, Madany P, Akech J, Dobson JR, Douthwright S, Browne G, Colby JL, Winter GE, Bradner JE, Pratap J et al (2015) The SWI/SNF ATPases are required for triple negative breast cancer cell proliferation. J Cell Physiol 230(11):2683–2694
Helming KC, Wang X, Wilson BG, Vazquez F, Haswell JR, Manchester HE, Kim Y, Kryukov GV, Ghandi M, Aguirre AJ et al (2014) ARID1B is a specific vulnerability in ARID1A-mutant cancers. Nat Med 20(3):251–254
Snell CE, Gough M, Middleton K, Hsieh M, Furnas L, Seidl B, Gibbons K, Pyke C, Shannon C, Woodward N et al (2017) Absent progesterone receptor expression in the lymph node metastases of ER-positive, HER2-negative breast cancer is associated with relapse on tamoxifen. J Clin Pathol 70(11):954–960
Sung P, Klein H (2006) Mechanism of homologous recombination: mediators and helicases take on regulatory functions. Nat Rev Mol Cell Biol 7(10):739–750
Goodarzi AA, Jeggo P, Lobrich M (2010) The influence of heterochromatin on DNA double strand break repair: getting thestrong, silent type to relax. DNA Repair (Amst) 9(12):1273–1282
Watanabe R, Ui A, Kanno S, Ogiwara H, Nagase T, Kohno T, Yasui A (2014) SWI/SNF factors required for cellular resistance to DNA damage include ARID1A and ARID1B and show interdependent protein stability. Cancer Res 74(9):2465–2475
Qi W, Wang R, Chen H, Wang X, Xiao T, Boldogh I, Ba X, Han L, Zeng X (2015) BRG1 promotes the repair of DNA double-strand breaks by facilitating the replacement of RPA with RAD51. J Cell Sci 128(2):317–330
Velez-Cruz R, Manickavinayaham S, Biswas AK, Clary RW, Premkumar T, Cole F, Johnson DG (2016) RB localizes to DNA double-strand breaks and promotes DNA end resection and homologous recombination through the recruitment of BRG1. Genes Dev 30(22):2500–2512
Shen J, Xiao Z, Wu WK, Wang MH, To KF, Chen Y, Yang W, Li MS, Shin VY, Tong JH et al (2015) Epigenetic silencing of miR-490-3p reactivates the chromatin remodeler SMARCD1 to promote Helicobacter pylori-induced gastric carcinogenesis. Cancer Res 75(4):754–765
de Castro RO, Previato L, Goitea V, Felberg A, Guiraldelli MF, Filiberti A, Pezza RJ (2017) The chromatin-remodeling subunit Baf200 promotes homology-directed DNA repair and regulates distinct chromatin-remodeling complexes. J Biol Chem 292(20):8459–8471
Peng G, Chun-Jen Lin C, Mo W, Dai H, Park YY, Kim SM, Peng Y, Mo Q, Siwko S, Hu R et al (2014) Genome-wide transcriptome profiling of homologous recombination DNA repair. Nat Commun 5:3361
He L, Chen Y, Feng J, Sun W, Li S, Ou M, Tang L (2017) Cellular senescence regulated by SWI/SNF complex subunits through p53/p21 and p16/pRB pathway. Int J Biochem Cell Biol 90:29–37
Hendricks KB, Shanahan F, Lees E (2004) Role for BRG1 in cell cycle control and tumor suppression. Mol Cell Biol 24(1):362–376
Newman RH, Zhang J (2008) Fucci: street lights on the road to mitosis. Chem Biol 15(2):97–98
Zack TI, Schumacher SE, Carter SL, Cherniack AD, Saksena G, Tabak B, Lawrence MS, Zhsng CZ, Wala J, Mermel CH et al (2013) Pan-cancer patterns of somatic copy number alteration. Nat Genet 45(10):1134–1140
Bultman SJ, Herschkowitz JI, Godfrey V, Gebuhr TC, Yaniv M, Perou CM, Magnuson T (2008) Characterization of mammary tumors from Brg1 heterozygous mice. Oncogene 27(4):460–468
Dykhuizen EC, Hargreaves DC, Miller EL, Cui K, Korshunov A, Kool M, Pfister S, Cho YJ, Zhao K, Crabtree GR (2013) BAF complexes facilitate decatenation of DNA by topoisomerase IIalpha. Nature 497(7451):624–627
Betts JA, Moradi Marjaneh M, Al-Ejeh F, Lim YC, Shi W, Sivakumaran H, Tropee R, Patch AM, Clark MB, Bartonicek N et al (2017) Long noncoding RNAs CUPID1 and CUPID2 mediate breast cancer risk at 11q13 by modulating the response to DNA damage. Am J Hum Genet 101(2):255–266
Carter SL, Cibulskis K, Helman E, McKenna A, Shen H, Zack T, Laird PW, Onofrio RC, Winckler W, Weir BA et al (2012) Absolute quantification of somatic DNA alterations in human cancer. Nat Biotechnol 30(5):413–421
Birkbak NJ, Wang ZC, Kim JY, Eklund AC, Li Q, Tian R, Bowman-Colin C, Li Y, Greene-Colozzi A, Iglehart JD et al (2012) Telomeric allelic imbalance indicates defective DNA repair and sensitivity to DNA-damaging agents. Cancer Discov 2(4):366–375
Popova T, Manie E, Rieunier G, Caux-Moncoutier V, Tirapo C, Dubois T, Delattre O, Sigal-Zafrani B, Bollet M, Longy M et al (2012) Ploidy and large-scale genomic instability consistently identify basal-like breast carcinomas with BRCA1/2 inactivation. Cancer Res 72(21):5454–5462
Abkevich V, Timms KM, Hennessy BT, Potter J, Carey MS, Meyer LA, Smith-McCune K, Broaddus R, Lu KH, Chen J et al (2012) Patterns of genomic loss of heterozygosity predict homologous recombination repair defects in epithelial ovarian cancer. Br J Cancer 107(10):1776–1782
Marquard AM, Eklund AC, Joshi T, Krzystanek M, Favero F, Wang ZC, Richardson AL, Silver DP, Szallasi Z, Birkbak NJ (2015) Pan-cancer analysis of genomic scar signatures associated with homologous recombination deficiency suggests novel indications for existing cancer drugs. Biomark Res 3:9
Funding
ED and PHGD are recipient of National Breast Cancer Fellowships. RT and BDLP are recipient of the Princess Alexandra Research Foundation. CS and MG are supported by funding from Mater Foundation. This work was funded by ECR13-04 (NBCF) and Cancer Council Queensland (CCQ). The Mays cancer center is supported by a NCI Cancer Center Support Core Grant P30 CA054174.
Author information
Authors and Affiliations
Contributions
ED, RT, and PHGD designed research, RT, BDLP, MG. and CS performed research, RT and PHGD. Wrote analysis pipelines, RT, ED, PHGD and CS analyzed data; and ED, PHGD and RT wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
All procedures performed in studies involving human samples were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Research involving human and animal participants
This article does not contain any studies with human participants or animals performed by any of the authors. Samples acquired from US Biomax guarantees informed consent was obtained from all individual participants included in the study. Subsequent staining experimental protocols were approved by the QUT ethics committee. Samples included in the Mater TMA were cleared for use by the Mater hospital ethics committee.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tropée, R., de la Peña Avalos, B., Gough, M. et al. The SWI/SNF subunit SMARCD3 regulates cell cycle progression and predicts survival outcome in ER+ breast cancer. Breast Cancer Res Treat 185, 601–614 (2021). https://doi.org/10.1007/s10549-020-05997-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-020-05997-5