Abstract
Purpose
Breast cancer (BC) is a heterogeneous disorder, with variable response to systemic chemotherapy. Likewise, BC shows highly complex immune activation patterns, only in part reflecting classical histopathological subtyping. Schlafen-11 (SLFN11) is a nuclear protein we independently described as causal factor of sensitivity to DNA damaging agents (DDA) in cancer cell line models. SLFN11 has been reported as a predictive biomarker for DDA and PARP inhibitors in human neoplasms. SLFN11 has been implicated in several immune processes such as thymocyte maturation and antiviral response through the activation of interferon signaling pathway, suggesting its potential relevance as a link between immunity and cancer. In the present work, we investigated the transcriptional landscape of SLFN11, its potential prognostic value, and the clinico-pathological associations with its variability in BC.
Methods
We assessed SLFN11 determinants in a gene expression meta-set of 5061 breast cancer patients annotated with clinical data and multigene signatures.
Results
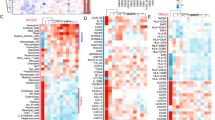
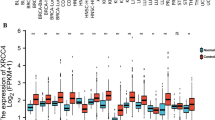
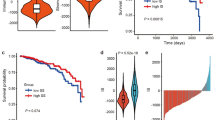
We found that 537 transcripts are highly correlated with SLFN11, identifying “immune response”, “lymphocyte activation”, and “T cell activation” as top Gene Ontology processes. We established a strong association of SLFN11 with stromal signatures of basal-like phenotype and response to chemotherapy in estrogen receptor negative (ER-) BC. We identified a distinct subgroup of patients, characterized by high SLFN11 levels, ER- status, basal-like phenotype, immune activation, and younger age. Finally, we observed an independent positive predictive role for SLFN11 in BC.
Conclusions
Our findings are suggestive of a relevant role for SLFN11 in BC and its immune and molecular variability.




Similar content being viewed by others
Change history
12 July 2019
In the original publication of the article, the funding information was incorrectly published. The corrected funding statement is given in this correction article.
Abbreviations
- BC:
-
Breast cancer
- DDA:
-
DNA damaging agents
- DFS:
-
Disease-free survival
- ER:
-
Estrogen receptor
- HT:
-
Hormone treatment
- ICR:
-
Immunological constant of rejection
- MCA:
-
Multiple correspondence analysis
- SLFN11:
-
Schlafen-11
- TNBC:
-
Triple-negative breast cancer
References
Ferlay J, Soerjomataram I, Dikshit RES, Mathers C, Rebelo M, Parkin DM, Forman D, Bray F (2015) Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer 136:E359–E386. https://doi.org/10.1002/ijc.29210
Perou CM, Sørlie T, Eisen MB et al (2000) Molecular portraits of human breast tumours. Nature 406:747–752. https://doi.org/10.1038/35021093
Sotiriou C, Pusztai L (2009) Gene-expression signatures in breast cancer. N Engl J Med 360:790–800. https://doi.org/10.1056/NEJMra0801289
Hart CD, Sanna G, Siclari O, Biganzoli L, Di Leo A (2015) Defining optimal duration and predicting benefit from chemotherapy in patients with luminal-like subtypes. Breast 24(Suppl 2):S136–S142. https://doi.org/10.1016/j.breast.2015.07.033
Barretina J, Caponigro G, Stransky N et al (2012) The cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483:603–607. https://doi.org/10.1038/nature11003
Zoppoli G, Regairaz M, Leo E, Reinhold WC, Varma S, Ballestrero A, Doroshow JH, Pommier Y (2012) Putative DNA/RNA helicase Schlafen-11 (SLFN11) sensitizes cancer cells to DNA-damaging agents. Proc Natl Acad Sci USA 109:15030–15035. https://doi.org/10.1073/pnas.1205943109
Katsoulidis E, Carayol N, Woodard J et al (2009) Role of Schlafen 2 (SLFN2) in the generation of interferon alpha-induced growth inhibitory responses. J Biol Chem 284:25051–25064. https://doi.org/10.1074/jbc.M109.030445
Geserick P, Kaiser F, Klemm U, Kaufmann SH, Zerrahn J (2004) Modulation of T cell development and activation by novel members of the Schlafen (slfn) gene family harbouring an RNA helicase-like motif. Int Immunol 16:1535–1548. https://doi.org/10.1093/intimm/dxh155
Li M, Kao E, Gao X et al (2012) Codon-usage-based inhibition of HIV protein synthesis by human schlafen 11. Nature 491:125–128. https://doi.org/10.1038/nature11433
Mavrommatis E, Fish EN, Platanias LC (2013) The Schlafen family of proteins and their regulation by interferons. J Interferon Cytokine Res 33:206–210. https://doi.org/10.1089/jir.2012.0133
Bustos O, Naik S, Ayers G, Casola C, Perez-Lamigueiro MA, Chippindale PT, Pritham EJ, de la Casa-Esperón E (2009) Evolution of the Schlafen genes, a gene family associated with embryonic lethality, meiotic drive, immune processes and orthopoxvirus virulence. Gene 447:1–11. https://doi.org/10.1016/j.gene.2009.07.006
Schwarz DA, Katayama CD, Hedrick SM (1998) Schlafen, a new family of growth regulatory genes that affect thymocyte development. Immunity 9:657–668
Neumann B, Zhao L, Murphy K, Gonda TJ (2008) Subcellular localization of the Schlafen protein family. Biochem Biophys Res Commun 370:62–66. https://doi.org/10.1016/j.bbrc.2008.03.032
Murai J, Tang S-W, Leo E et al (2018) SLFN11 blocks stressed replication forks independently of ATR. Mol Cell 69:371–384. https://doi.org/10.1016/j.molcel.2018.01.012
Mu Y, Lou J, Srivastava M, Srivastava M, Zhao B, Feng XH, Liu T, Chen J, Huang J (2016) SLFN11 inhibits checkpoint maintenance and homologous recombination repair. EMBO Rep 17:94–109. https://doi.org/10.15252/embr.201540964
Tang S-W, Thomas A, Murai J, Trepel JB, Bates SE, Rajapakse VN, Pommier Y (2018) Overcoming resistance to DNA targeted agents by epigenetic activation of Schlafen 11 (SLFN11) expression with class I histone deacetylase inhibitors. Clin Cancer Res. https://doi.org/10.1158/1078-0432.ccr-17-0443
Tang S-W, Bilke S, Cao L et al (2015) SLFN11 is a transcriptional target of EWS-FLI1 and a determinant of drug response in ewing sarcoma. Clin Cancer Res 21:4184–4193. https://doi.org/10.1158/1078-0432.CCR-14-2112
Tian L, Song S, Liu X et al (2014) Schlafen-11 sensitizes colorectal carcinoma cells to irinotecan. Anticancer Drugs 25:1175–1181. https://doi.org/10.1097/CAD.0000000000000151
Nogales V, Reinhold WC, Varma S et al (2016) Epigenetic inactivation of the putative DNA/RNA helicase SLFN11 in human cancer confers resistance to platinum drugs. Oncotarget 7:3084–3097. https://doi.org/10.18632/oncotarget.6413
He T, Zhang M, Zheng R et al (2017) Methylation of SLFN11is a marker of poor prognosis and cisplatin resistance in colorectal cancer. Epigenomics 9:849–862. https://doi.org/10.2217/epi-2017-0019
Pietanza MC, Waqar SN, Krug LM et al (2018) Randomized, double-blind, phase II study of temozolomide in combination with either veliparib or placebo in patients with relapsed-sensitive or refractory small-cell lung cancer. J Clin Oncol. https://doi.org/10.1200/jco.2018.77.7672
Haibe-Kains B, Desmedt C, Loi S et al (2012) A three-gene model to robustly identify breast cancer molecular subtypes. J Natl Cancer Inst 104:311–325. https://doi.org/10.1093/jnci/djr545
Cancer Genome Atlas Network (2012) Comprehensive molecular portraits of human breast tumours. Nature 490:61–70. https://doi.org/10.1038/nature11412
Curtis C, Shah SP, Chin S-F et al (2012) The genomic and transcriptomic architecture of 2000 breast tumours reveals novel subgroups. Nature 486:346–352. https://doi.org/10.1038/nature10983
Dedeurwaerder S, Desmedt C, Calonne E et al (2011) DNA methylation profiling reveals a predominant immune component in breast cancers. EMBO Mol Med 3:726–741. https://doi.org/10.1002/emmm.201100801
Hendrickx W, Simeone I, Anjum S et al (2017) Identification of genetic determinants of breast cancer immune phenotypes by integrative genome-scale analysis. OncoImmunology 6:e1253654. https://doi.org/10.1080/2162402X.2016.1253654
Gendoo DMA, Ratanasirigulchai N, Schröder MS et al (2016) Genefu: an R/bioconductor package for computation of gene expression-based signatures in breast cancer. Bioinformatics 32:1097–1099. https://doi.org/10.1093/bioinformatics/btv693
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57. https://doi.org/10.1038/nprot.2008.211
Prat A, Perou CM (2010) Deconstructing the molecular portraits of breast cancer. Mol Oncol 5:5–23. https://doi.org/10.1016/j.molonc.2010.11.003
Desmedt C, Zoppoli G, Gundem G et al (2016) Genomic characterization of primary invasive lobular breast cancer. J Clin Oncol 34:1872–1881. https://doi.org/10.1200/JCO.2015.64.0334
Ignatiadis M, Singhal SK, Desmedt C et al (2012) Gene modules and response to neoadjuvant chemotherapy in breast cancer subtypes: a pooled analysis. J Clin Oncol 30:1996–2004. https://doi.org/10.1200/JCO.2011.39.5624
Desmedt C, Haibe-Kains B, Wirapati P et al (2008) Biological processes associated with breast cancer clinical outcome depend on the molecular subtypes. Clin Cancer Res 14:5158–5165. https://doi.org/10.1158/1078-0432.CCR-07-4756
Allison Stewart C, Tong P, Cardnell RJ et al (2017) Dynamic variations in epithelial-to-mesenchymal transition (EMT), ATM, and SLFN11 govern response to PARP inhibitors and cisplatin in small cell lung cancer. Oncotarget. 17:28575–28587. https://doi.org/10.18632/oncotarget.15338
Puck A, Aigner R, Modak M et al (2015) Expression and regulation of Schlafen (SLFN) family members in primary human monocytes, monocyte-derived dendritic cells and T cells. Results Immunol 5:23–32. https://doi.org/10.1016/j.rinim.2015.10.001
Bedognetti D, Hendrickx W, Marincola FM, Miller LD (2015) Prognostic and predictive immune gene signatures in breast cancer. Curr Opin Oncol 27:433–444. https://doi.org/10.1097/CCO.0000000000000234
Denkert C, von Minckwitz G, Brase JC et al (2015) Tumor-infiltrating lymphocytes and response to neoadjuvant chemotherapy with or without carboplatin in human epidermal growth factor receptor 2-positive and triple-negative primary breast cancers. J Clin Oncol 33:983–991. https://doi.org/10.1200/JCO.2014.58.1967
Gingras I, Desmedt C, Ignatiadis M, Sotiriou C (2015) CCR 20th anniversary commentary: gene-expression signature in breast cancer-where did it start and where are we now? Clin Cancer Res 21:4743–4746. https://doi.org/10.1158/1078-0432.CCR-14-3127
Cleator SJ, Powles TJ, Dexter T et al (2006) The effect of the stromal component of breast tumours on prediction of clinical outcome using gene expression microarray analysis. Breast Cancer Res 8:R32. https://doi.org/10.1186/bcr1506
Winslow S, Leandersson K, Edsjö A, Larsson C (2015) Prognostic stromal gene signatures in breast cancer. Breast Cancer Res 17:747. https://doi.org/10.1186/s13058-015-0530-2
Farmer P, Bonnefoi H, Anderle P et al (2009) A stroma-related gene signature predicts resistance to neoadjuvant chemotherapy in breast cancer. Nat Med 15:68–74. https://doi.org/10.1038/nm.1908
Finak G, Bertos N, Pepin F et al (2008) Stromal gene expression predicts clinical outcome in breast cancer. Nat Med 14:518–527. https://doi.org/10.1038/nm1764
Gu-Trantien C, Loi S, Garaud S et al (2013) CD4+ follicular helper T cell infiltration predicts breast cancer survival. J Clin Invest 123:2873–2892. https://doi.org/10.1172/JCI67428
Meissl K, Macho-Maschler S, Müller M, Strobl B (2017) The good and the bad faces of STAT1 in solid tumours. Cytokine 89:12–20. https://doi.org/10.1016/j.cyto.2015.11.011
Bianchini G, Balko JM, Mayer IA et al (2016) Triple-negative breast cancer: challenges and opportunities of a heterogeneous disease. Nat Pub Group 13:674–690. https://doi.org/10.1038/nrclinonc.2016.66
Metzger-Filho O, Sun Z, Viale G et al (2013) Patterns of Recurrence and outcome according to breast cancer subtypes in lymph node-negative disease: results from international breast cancer study group trials VIII and IX. J Clin Oncol 31:3083–3090. https://doi.org/10.1200/JCO.2012.46.1574
Bhargava R, Beriwal S, Dabbs DJ et al (2010) Immunohistochemical surrogate markers of breast cancer molecular classes predicts response to neoadjuvant chemotherapy. Cancer 116:1431–1439. https://doi.org/10.1002/cncr.24876
Nelson DJ, Clark B, Munyard K et al (2017) A review of the importance of immune responses in luminal B breast cancer. OncoImmunology 6:e1282590. https://doi.org/10.1080/2162402X.2017.1282590
Yersal O, Barutca S (2014) Biological subtypes of breast cancer: prognostic and therapeutic implications. World J Clin Oncol 5:412–424. https://doi.org/10.5306/wjco.v5.i3.412
Miller LD, Chou JA, Black MA et al (2016) Immunogenic subtypes of breast cancer delineated by gene classifiers of immune responsiveness. Cancer Immunol Res 4:600–610. https://doi.org/10.1158/2326-6066.CIR-15-0149
Acknowledgements
GZ would like to thank Dr. P. Blandini, MD, for his invaluable scientific insights during all the phases of this project.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Availability of data and material
All raw data used for the generation of the expression set we analyzed are available in GEO under their respective publication IDs. Normalized expression data are available upon request to the Corresponding Author.
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Isnaldi, E., Ferraioli, D., Ferrando, L. et al. Schlafen-11 expression is associated with immune signatures and basal-like phenotype in breast cancer. Breast Cancer Res Treat 177, 335–343 (2019). https://doi.org/10.1007/s10549-019-05313-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-019-05313-w