Abstract
Background:
Extensive efforts have been undertaken to discover genes relevant for breast cancer prognosis. Yet, in current opinion, with little overlap in findings. We aimed to reanalyze molecular prediction of breast cancer recurrence.
Methods:
From 44 published gene lists relevant for breast cancer prognosis, we extracted 374 genes, which, besides other quality criteria, are recorded at least twice. From eight published microarray datasets, a single dataset of 1,067 breast cancer patients was created, using transformation to ‘probability of expression’ scale. For recurrence analysis, the Cox proportional hazards model was applied.
Results:
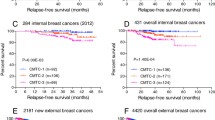
The 374 genes, termed ‘374 Gene Set’, are highly enriched in cell cycle genes. The ‘374 Gene Set’ is significantly associated with breast cancer recurrence (p = 2 × 10−12, log-rank test) in the meta set of 1,067 patients, showing an estimated Hazard Ratio of recurrence for the ‘poor’ prognosis group compared to the ‘good’ prognosis group of 2.03 (95% confidence interval, 1.66–2.48). Notably, the ‘374 Gene Set’ is significantly associated with recurrence in untreated patients. In multivariate analysis, including the standard histopathological parameters, only tumor size and the ‘374 Gene Set’ remain independent predictors of recurrence. External validation further confirmed the prognostic relevance of the gene set (253 patients, p = 0.001, log-rank test).
Conclusions:
The ‘374 Gene Set’ comprises a molecular basis of metastatic breast cancer progression. Starting from this gene set it might be possible to construct a clinically relevant classifier, which then again needs to be validated.




Similar content being viewed by others
References
Goldhirsch A, Wood WC, Gelber RD et al (2003) Meeting highlights: updated international expert consensus on the primary therapy of early breast cancer. J Clin Oncol 21:3357–3365
Eifel P, Axelson JA, Costa J et al (2001) National institutes of health consensus development conference statement: adjuvant therapy for breast cancer, November 1–3, 2000. J Natl Cancer Inst 93:979–989
van’t Veer LJ, Dai H, van de Vijver MJ et al (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415:530–536
Sorlie T, Perou CM, Tibshirani R et al (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98:10869–10874
Wang Y, Klijn JG, Zhang Y et al (2005) Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet 365:671–679
Paik S, Shak S, Tang G et al (2004) A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 351:2817–2826
Ioannidis JP (2005) Why most published research findings are false. PLoS Med 2:e124
Gruvberger SK, Ringner M, Eden P et al (2003) Expression profiling to predict outcome in breast cancer: the influence of sample selection. Breast Cancer Res 5:23–26
Michiels S, Koscielny S, Hill C (2005) Prediction of cancer outcome with microarrays: a multiple random validation strategy. Lancet 365:488–492
Lahad JP, Mills GB, Coombes KR (2005) Stem cell-ness: a “magic marker” for cancer. J Clin Invest 115:1463–1467
Zhang B, Schmoyer D, Kirov S et al (2004) GOTree Machine (GOTM): a web-based platform for interpreting sets of interesting genes using Gene Ontology hierarchies. BMC Bioinformatics 5:16
Kent WJ, Sugnet CW, Furey TS et al (2002) The human genome browser at UCSC. Genome Res 12:996–1006
Sotiriou C, Wirapati P, Loi S et al (2006) Gene expression profiling in breast cancer: understanding the molecular basis of histologic grade to improve prognosis. J Natl Cancer Inst 98:262–272
van de Vijver MJ, He YD, van’t Veer LJ et al (2002) A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 347:1999–2009
Ma XJ, Wang Z, Ryan PD et al (2004) A two-gene expression ratio predicts clinical outcome in breast cancer patients treated with tamoxifen. Cancer Cell 5:607–616
Shen R, Ghosh D, Chinnaiyan AM (2004) Prognostic meta-signature of breast cancer developed by two-stage mixture modeling of microarray data. BMC Genomics 5:94
Scharpf R, Garrett ES, Hu J, et al (2003) Statistical modeling and visualization of molecular profiles in cancer. Biotechniques Suppl:22–29
Ivshina AV, George J, Senko O et al (2006) Genetic reclassification of histologic grade delineates new clinical subtypes of breast cancer. Cancer Res 66:10292–10301
de Hoon MJ, Imoto S, Nolan J et al (2004) Open source clustering software. Bioinformatics 20:1453–1454
Abba MC, Hu Y, Sun H et al (2005) Gene expression signature of estrogen receptor alpha status in breast cancer. BMC Genomics 6:37
Amatschek S, Koenig U, Auer H et al (2004) Tissue-wide expression profiling using cDNA subtraction and microarrays to identify tumor-specific genes. Cancer Res 64:844–856
Beer DG, Kardia SL, Huang CC et al (2002) Gene-expression profiles predict survival of patients with lung adenocarcinoma. Nat Med 8:816–824
Berchuck A, Iversen ES, Lancaster JM et al (2005) Patterns of gene expression that characterize long-term survival in advanced stage serous ovarian cancers. Clin Cancer Res 11:3686–3696
Bertucci F, Nasser V, Granjeaud S et al (2002) Gene expression profiles of poor-prognosis primary breast cancer correlate with survival. Hum Mol Genet 11:863–872
Bieche I, Tozlu S, Girault I et al (2004) Identification of a three-gene expression signature of poor-prognosis breast carcinoma. Mol Cancer 3:37
Chang HY, Nuyten DS, Sneddon JB et al (2005) Robustness, scalability, and integration of a wound-response gene expression signature in predicting breast cancer survival. Proc Natl Acad Sci USA 102:3738–3743
Glinsky GV, Berezovska O, Glinskii AB (2005) Microarray analysis identifies a death-from-cancer signature predicting therapy failure in patients with multiple types of cancer. J Clin Invest 115:1503–1521
Glinsky GV, Higashiyama T, Glinskii AB (2004) Classification of human breast cancer using gene expression profiling as a component of the survival predictor algorithm. Clin Cancer Res 10:2272–2283
Glinsky GV, Glinskii AB, Stephenson AJ et al (2004) Gene expression profiling predicts clinical outcome of prostate cancer. J Clin Invest 113:913–923
Huang E, Cheng SH, Dressman H, et al (2003) Gene expression predictors of breast cancer outcomes. Lancet 361:1590–1596
Iwao K, Matoba R, Ueno N et al (2002) Molecular classification of primary breast tumors possessing distinct prognostic properties. Hum Mol Genet 11:199–206
Jacquemier J, Ginestier C, Rougemont J et al (2005) Protein expression profiling identifies subclasses of breast cancer and predicts prognosis. Cancer Res 65:767–779
Jones C, Mackay A, Grigoriadis A et al (2004) Expression profiling of purified normal human luminal and myoepithelial breast cells: identification of novel prognostic markers for breast cancer. Cancer Res 64:3037–3045
Korkola JE, DeVries S, Fridlyand J et al (2003) Differentiation of lobular versus ductal breast carcinomas by expression microarray analysis. Cancer Res 63:7167–7175
Ma XJ, Salunga R, Tuggle JT et al (2003) Gene expression profiles of human breast cancer progression. Proc Natl Acad Sci USA 100:5974–5979
Makretsov NA, Huntsman DG, Nielsen TO et al (2004) Hierarchical clustering analysis of tissue microarray immunostaining data identifies prognostically significant groups of breast carcinoma. Clin Cancer Res 10:6143–6151
Nutt CL, Mani DR, Betensky RA et al (2003) Gene expression-based classification of malignant gliomas correlates better with survival than histological classification. Cancer Res 63:1602–1607
Onda M, Emi M, Nagai H et al (2004) Gene expression patterns as marker for 5-year postoperative prognosis of primary breast cancers. J Cancer Res Clin Oncol 130:537–545
Pomeroy SL, Tamayo P, Gaasenbeek M et al (2002) Prediction of central nervous system embryonal tumour outcome based on gene expression. Nature 415:436–442
Pusztai L, Ayers M, Stec J et al (2003) Gene expression profiles obtained from fine-needle aspirations of breast cancer reliably identify routine prognostic markers and reveal large-scale molecular differences between estrogen-negative and estrogen-positive tumors. Clin Cancer Res 9:2406–2415
Ramaswamy S, Ross KN, Lander ES et al (2003) A molecular signature of metastasis in primary solid tumors. Nat Genet 33:49–54
Rhodes DR, Yu J, Shanker K et al (2004) Large-scale meta-analysis of cancer microarray data identifies common transcriptional profiles of neoplastic transformation and progression. Proc Natl Acad Sci USA 101:9309–9314
Rosenwald A, Wright G, Chan WC et al (2002) The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N Engl J Med 346:1937–1947
Sotiriou C, Neo SY, McShane LM et al (2003) Breast cancer classification and prognosis based on gene expression profiles from a population-based study. Proc Natl Acad Sci USA 100:10393–10398
West M, Blanchette C, Dressman H et al (2001) Predicting the clinical status of human breast cancer by using gene expression profiles. Proc Natl Acad Sci USA 98:11462–11467
Woelfle U, Cloos J, Sauter G et al (2003) Molecular signature associated with bone marrow micrometastasis in human breast cancer. Cancer Res 63:5679–5684
Yu K, Lee CH, Tan PH et al (2004) A molecular signature of the Nottingham prognostic index in breast cancer. Cancer Res 64:2962–2968
Zhu G, Reynolds L, Crnogorac-Jurcevic T et al (2003) Combination of microdissection and microarray analysis to identify gene expression changes between differentially located tumour cells in breast cancer. Oncogene 22:3742–3748
Ahr A, Karn T, Solbach C et al (2002) Identification of high risk breast-cancer patients by gene expression profiling. Lancet 359:131–132
West RB, Nuyten DS, Subramanian S et al (2005) Determination of stromal signatures in breast carcinoma. PLoS Biol 3:e187
Miller DV, Leontovich AA, Lingle WL et al (2004) Utilizing Nottingham Prognostic Index in microarray gene expression profiling of breast carcinomas. Mod Pathol 17:756–764
Van Laere S, Van dA I, Van den Eynden GG et al (2005) Distinct molecular signature of inflammatory breast cancer by cDNA microarray analysis. Breast Cancer Res Treat 93:237–246
Hu Z, Fan C, Oh DS et al (2006) The molecular portraits of breast tumors are conserved across microarray platforms. BMC Genomics 7:96
Dai H, van’t Veer L, Lamb J et al (2005) A cell proliferation signature is a marker of extremely poor outcome in a subpopulation of breast cancer patients. Cancer Res 65:4059–4066
Wang W, Wyckoff JB, Frohlich VC et al (2002) Single cell behavior in metastatic primary mammary tumors correlated with gene expression patterns revealed by molecular profiling. Cancer Res 62:6278–6288
Lang G, Gombert WM, Gould HJ (2005) A transcriptional regulatory element in the coding sequence of the human Bcl-2 gene. Immunology 114:25–36
Ginestier C, Cervera N, Finetti P et al (2006) Prognosis and gene expression profiling of 20q13-amplified breast cancers. Clin Cancer Res 12:4533–4544
Kapp AV, Jeffrey SS, Langerod A et al (2006) Discovery and validation of breast cancer subtypes. BMC Genomics 7:231
Muller HM, Widschwendter A, Fiegl H et al (2003) DNA methylation in serum of breast cancer patients: an independent prognostic marker. Cancer Res 63:7641–7645
Irizarry RA, Warren D, Spencer F et al (2005) Multiple-laboratory comparison of microarray platforms. Nat Methods 2:345–350
Ein-Dor L, Kela I, Getz G et al (2005) Outcome signature genes in breast cancer: is there a unique set? Bioinformatics 21:171–178
Warnat P, Eils R, Brors B (2005) Cross-platform analysis of cancer microarray data improves gene expression based classification of phenotypes. BMC Bioinformatics 6:265
Choi JK, Yu U, Kim S, et al (2003) Combining multiple microarray studies and modeling interstudy variation. Bioinformatics 19(Suppl 1):i84–i90
Acknowledgments
We thank Prof. Fritz Leisch and Prof. Fritz Pittner for constructive advice. This work was supported by FFG, project no. 809596. The sponsor had no role in any part of the study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Martin Lauss, Albert Kriegner, Klemens Vierlinger, and Christa Noehammer are named inventors on a related patent application
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
Summary of the 44 gene lists used in this study (DOC 75 kb)
Supplementary Table 2
All 4475 gene identifiers linked to their corresponding input gene list, input identifiers and output annotation to UniGene Cluster (XLS 1098 kb)
Supplementary Table 3
Genes of the ‘374 Gene Set’ and centroids for the ‘good’ and ‘poor’ prognosis-groups (XLS 104 kb)
Supplementary Table 4
Patient data and grouping of the metaset and external validation set (XLS 362 kb)
Supplementary Figure 1
DAG diagram of the GO terms enriched in the ‘374 Gene Set’ when compared to the human genome (GIF 294 kb)
Supplementary Figure 2
KEGG pathway chart ‘cell cycle’. Genes contained in ‘374 Gene Set’ are red (GIF 87 kb)
Supplementary Figure 3
Kaplan Meier plots of recurrence-free survival (RFS) using 1067 patients that were grouped by the reduced independent gene set. Thirty-three genes remained for grouping the patients and subsequent survival analysis. Numbers at risk are given at last time of event before the fixed timepoints (0,50,100,150,200) (EMF 321 kb)
Rights and permissions
About this article
Cite this article
Lauss, M., Kriegner, A., Vierlinger, K. et al. Consensus genes of the literature to predict breast cancer recurrence. Breast Cancer Res Treat 110, 235–244 (2008). https://doi.org/10.1007/s10549-007-9716-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-007-9716-3