Abstract
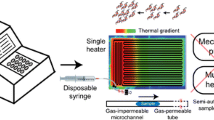
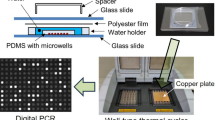
Polymerase chain reaction (PCR) has been considered as the gold standard for detecting nucleic acids. The simple PCR system is of great significance for medical applications in remote areas, especially for the developing countries. Herein, we proposed a low-cost self-assembled platform for microchamber PCR. The working principle is rotating the chamber PCR microfluidic chip between two heaters with fixed temperature to solve the problem of low temperature variation rate. The system consists of two temperature controllers, a screw slide rail, a chamber array microfluidic chip and a self-built software. Such a system can be constructed at a cost of about US$60. The micro chamber PCR can be finished by rotating the microfluidic chip between two heaters with fixed temperature. Results demonstrated that the sensitivity of the temperature controller is 0.1℃. The relative error of the duration for the microfluidic chip was 0.02 s. Finally, we successfully finished amplification of the target gene of Porphyromonas gingivalis in the chamber PCR microfluidic chip within 35 min and on-site detection of its PCR products by fluorescence. The chip consisted of 3200 cylindrical chambers. The volume of reagent in each volume is as low as 0.628 nL. This work provides an effective method to reduce the amplification time required for micro chamber PCR.







Similar content being viewed by others
References
S.E. Cavanaugh, A.S. Bathrick, Forensic science international: Genetics. 32,40 – 9(2018)
S. Chen, Y. Sun, F. Fan, S. Chen, Y. Zhang, Y. Zhang et al., TRAC Trends Anal. Chem. 157,116737(2022)
X. Cui, L. Wu, Y. Wu, J.H. Zhang, Q. Zhao, F.X. Jing et al., Anal. Chim. Acta 1107,127 – 34(2020)
J.S. Farrar, C.T. Wittwer, Clin Chem. 61,145 – 53(2015)
F. Gouriet Fdr, Fenollar, J.-Y. Patrice, M. Drancourt, D. Raoult, J. Clin. Microbiol. 43, 4993–5002 (2005)
K. Hosokawa, H. Ohmori, Analytical Sciences. 39,2067-74(2023)
Y. Huang, Z. Gao, C. Ma, Y. Sun, Y. Huang, C. Jia et al., Analyst (2023)
M.U. Kopp, A.J. Mello, A. Manz, Science. 280, 1046–1048 (1998)
M. Krishnan, V.M. Ugaz, M.A. Burns, Science. 298, 793 (2002)
S.H. Lee, S.-Y. Ruan, S.-C. Pan, T.-F. Lee, J.-Y. Chien, P.-R. Hsueh, J. Microbiol. Immunol. Infect. 52, 920–928 (2019)
Z. Li, Y. Zhao, D. Zhang, S. Zhuang, Y. Yamaguchi, Sens. Actuators B 230,779 – 84(2016)
Z. Li, R. Ju, S. Sekine, D. Zhang, S. Zhuang, Y. Yamaguchi, Lab. On a Chip. 19, 2663–2668 (2019)
Z. Li, J. Liu, P. Wang, C. Tao, L. Zheng, S. Sekine et al., Lab. On a Chip. 21, 3159–3164 (2021)
Z. Li, Y. Wang, Z. Gao, S. Sekine, Q. You, S. Zhuang et al., Anal. Chim. Acta 1251,340995(2023)
C. Liu, W. Eschen, L. Loetgering, D.S. Penagos Molina, R. Klas, A. Iliou et al., PhotoniX. 4, 1–15 (2023)
T.C. Merkel, V.I. Bondar, K. Nagai, B.D. Freeman, I. Pinnau, J. Polym. Sci. Pol. Phys. 38,415 – 34(2000)
Y. Ning, X. Cui, C. Yang, F. Jing, X. Bian, L. Yi et al., Anal. Chim. Acta. 1055, 65–73 (2019)
T. Notomi, H. Okayama, H. Masubuchi, T. Yonekawa, K. Watanabe, N. Amino et al., Nucleic Acids Res. 28, e63 (2000)
J. Shen, J. Zheng, Z. Li, Y. Liu, F. Jing, X. Wan et al., Lab. On a Chip. 21, 3742–3747 (2021)
Q. Song, Y.B. Gao, Q.Y. Zhu, Q.C. Tian, B.W. Yu, B.F. Song et al., Biomed. Microdevices. 17, 64 (2015)
T. Suo, X. Liu, J. Feng, M. Guo, W. Hu, D. Guo et al., Emerg. Microbes Infections. 9, 1259–1268 (2020)
H. Wang, Y.-L. Zhang, D.-D. Han, W. Wang, H.-B. Sun, PhotoniX. 2, 1–13 (2021)
L. Xu, H. Lee, D. Jetta, K.W. Oh, Lab. On a Chip. 15, 3962–3979 (2015)
B. Yang, P. Wang, Z. Li, C. Tao, Q. You, S. Sekine et al., Lab. On a Chip. 22, 733–737 (2022)
Q.Y. Zhu, L. Qiu, B.W. Yu, Y.N. Xu, Y.B. Gao, T.T. Pan et al., Lab. On a Chip. 14, 1176–1185 (2014)
Q.Y. Zhu, Y.N. Xu, L. Qiu, C.C. Ma, B.W. Yu, Q. Song et al., Lab. On a Chip. 17, 1655–1665 (2017)
Acknowledgements
This work was supported by Science and Technology Commission of Shanghai Municipality, China (No.18441900400 and No.19ZR1477500). It was also supported by Excellent Young Talents support plan in Colleges of Anhui Province Grant (gxgnfx 2021167).
Author information
Authors and Affiliations
Contributions
Z. L: Project administration, Supervision, Funding acquisition, Writing; X. M, X. W, X. Y: Methodology, Data acquisition; Z. Z and B. Y: Methodology, Periodontal pathogen extraction; D. Z: Project administration, Supervision; Y. Y: Project administration, Supervision, Review &editing; J. Y and Y. Z: Data analysis, Review &editing.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have declared no conflict of interest.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Z., Ma, X., Zhang, Z. et al. A rapid and low-cost platform for detection of bacterial based on microchamber PCR microfluidic chip. Biomed Microdevices 26, 20 (2024). https://doi.org/10.1007/s10544-024-00699-x
Accepted:
Published:
DOI: https://doi.org/10.1007/s10544-024-00699-x