Abstract
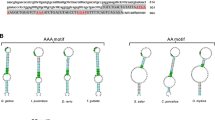
Selenium (Se) plays an essential role in the growth of fish and performs its physiological functions mainly through incorporation into selenoproteins. Our previous studies suggested that the selenoprotein W gene (selenow) is sensitive to changes in dietary Se in rainbow trout. However, the molecular characterization and tissue expression pattern of selenow are still unknown. Here, we revealed the molecular characterization, the tissue expression pattern of rainbow trout selenow and analyzed its response to dietary Se. The open reading frame (ORF) of the selenow gene was composed of 393 base pairs (bp) and encodes a 130-amino-acid protein. The 3′ untranslated region (UTR) was 372 bp with a selenocysteine insertion sequence (SECIS) element. Remarkably, the rainbow trout selenow gene sequence was longer than those reported for mammals and most other fish. A β1-α1-β2-β3-β4-α2 pattern made up the secondary structure of SELENOW. Furthermore, multiple sequence alignment revealed that rainbow trout SELENOW showed a high level of identity with SELENOW from Salmo salar. In addition, the selenow gene was ubiquitously distributed in 13 tissues with various abundances and was predominantly expressed in muscle and brain. Interestingly, dietary Se significantly increased selenow mRNA expression in muscle. Our results highlight the vital role of selenow in rainbow trout muscle response to dietary Se levels and provide a theoretical basis for studies of selenow.







Similar content being viewed by others
References
Aachmann FL, Fomenko DE, Soragni A, Gladyshev VN, Dikiy A (2007) Solution structure of selenoprotein W and NMR analysis of its interaction with 14–3-3 proteins. J Biol Chem 282:37036–37044. https://doi.org/10.1074/jbc.M705410200
Beilstein MA, Vendeland SC, Barofsky E, Jensen ON, Whanger PD (1996) Selenoprotein W of rat muscle binds glutathione and an unknown small molecular weight moiety. J Inorg Biochem 61:117–124. https://doi.org/10.1016/0162-0134(95)00045-3
Chambers I, Frampton J, Goldfarb P, Affara N, McBain W, Harrison PR (1986) The structure of the mouse glutathione peroxidase gene: the selenocysteine in the active site is encoded by the ‘termination’ codon, TGA. EMBO J 5:1221–1227. https://doi.org/10.1002/j.1460-2075.1986.tb04350.x
Chapple CE, Guigó R, Krol A (2009) SECISaln, a web-based tool for the creation of structure-based alignments of eukaryotic SECIS elements. Bioinformatics 25:674–675. https://doi.org/10.1093/bioinformatics/btp020
Chen W, Zhang Z, Dong H, Jiang X (2015) Molecular cloning and sequence analysis of selenoprotein W gene and its mRNA expression patterns in response to metabolic status and cadmium exposure in goldfish, Carassius auratus. Comp Biochem Physiol B 184:1–9. https://doi.org/10.1016/j.cbpb.2015.01.005
Dikiy A, Novoselov SV, Fomenko DE, Sengupta A, Carlson BA, Cerny RL, Ginalski K, Grishin NV, Hatfield DL, Gladyshev VN (2007) SelT, SelW, SelH, and Rdx12: genomics and molecular insights into the functions of selenoproteins of a novel thioredoxin-like family. Biochemistry 46:6871–6882. https://doi.org/10.1021/bi602462q
Dong H, Chen W, Sun C, Sun J, Wang Y, Xie C, Fu Q, Zhu J, Ye J (2017) Identification, characterization of selenoprotein W and its mRNA expression patterns in response to somatostatin 14, cysteamine hydrochloride, 17β-estradiol and a binary mixture of 17β-estradiol and cysteamine hydrochloride in topmouth culter (Erythroculter ilishaeformis). Fish Physiol Biochem 43:115–126. https://doi.org/10.1007/s10695-016-0272-9
Fletcher JE, Copeland PR, Driscoll DM, Krol A (2001) The selenocysteine incorporation machinery: interactions between the SECIS RNA and the SECIS-binding protein SBP2. RNA 7:1442–1453
Gladyshev VN (2001) Identity, evolution and function of selenoproteins and selenoprotein genes. In: Hatfield DL (ed) Selenium. Springer, Boston
Goody TA, Melcher SE, Norman DG, Lilley DMJ (2004) The kink-turn motif in RNA is dimorphic, and metal ion-dependent. RNA 10:254–264. https://doi.org/10.1261/rna.5176604
Gu QP, Beilstein MA, Barofsky E, Ream W, Whanger PD (1999) Purification, characterization, and glutathione binding to selenoprotein W from monkey muscle. Arch Biochem Biophys 361:25–33. https://doi.org/10.1006/abbi.1998.0949
Guyomard R, Boussaha M, Krieg F, Hervet C, Quillet E (2012) A synthetic rainbow trout linkage map provides new insights into the salmonid whole genome duplication and the conservation of synteny among teleosts. BMC Genet 13:15. https://doi.org/10.1186/1471-2156-13-15
Hoffmann PR, Berry MJ (2005) Selenoprotein synthesis: a unique translational mechanism used by a diverse family of proteins. Thyroid 15:769–775. https://doi.org/10.1089/thy.2005.15.769
Hu B, Liu Y, Wen C-G, Li A-H, Hu X-P, Wu D, Hu X-J, Tao Z-Y (2014) Cloning and expression of selenoprotein W from pearl mussels Cristaria plicata. Comp Biochem Physiol B 167:8–15. https://doi.org/10.1016/j.cbpb.2013.09.008
Jeon YH, Ko KY, Lee JH, Park KJ, Jang JK, Kim IY (2016) Identification of a redox-modulatory interaction between selenoprotein W and 14–3-3 protein. Biochim Biophys Acta 1863:10–18. https://doi.org/10.1016/j.bbamcr.2015.10.006
Kim H, Lee K, Kim JM, Kim MY, Kim JR, Lee HW, Chung YW, Shin HI, Kim T, Park ES, Rho J, Lee SH, Kim N, Lee SY, Choi Y, Jeong D (2021) Selenoprotein W ensures physiological bone remodeling by preventing hyperactivity of osteoclasts. Nat Commun 12:2258. https://doi.org/10.1038/s41467-021-22565-7
Kryukov G, Gladyshev V (2000) Selenium metabolism in zebrafish: multiplicity of selenoprotein genes and expression of a protein containing 17 selenocysteine residues. Genes Cells 5:1049–1060. https://doi.org/10.1046/j.1365-2443.2000.00392.x
Labunskyy VM, Hatfield DL, Gladyshev VN (2014) Selenoproteins: molecular pathways and physiological roles. Physiol Rev 94:739–777. https://doi.org/10.1152/physrev.00039.2013
Li JL, Ruan H, Li HX, Li S, Xu SW, Tang ZX (2011) Molecular cloning, characterization and mRNA expression analysis of a novel selenoprotein: avian selenoprotein W from chicken. Mol Biol Rep 38:4015–4022. https://doi.org/10.1007/s11033-010-0520-5
Liang Y, Lin S, Wang C, Yao H, Zhang Z, Xu SW (2014) Effect of selenium on selenoprotein expression in the adipose tissue of chickens. Biol Trace Elem Res 160:41–48. https://doi.org/10.1007/s12011-014-0024-6
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2 (-Delta Delta C(T)) Method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Low SC, Berry MJ (1996) Knowing when not to stop: selenocysteine incorporation in eukaryotes. Trends Biochem Sci 21:203–208. https://doi.org/10.1016/S0968-0004(96)80016-8
Lv H, Zhou T, Dong C, Kong S, Chen L, Pu F, Li X, Xu P (2020) Genome-wide identification, evolution, and mRNA expression of complement genes in common carp (Cyprinus carpio). Fish Shellfish Immunol 96:190–200. https://doi.org/10.1016/j.fsi.2019.11.032
Marandel L, Kostyniuk DJ, Best C, Forbes JLI, Liu J, Panserat S, Mennigen JA (2019) Pck-ing up steam: widening the salmonid gluconeogenic gene duplication trail. Gene 698:129–140. https://doi.org/10.1016/j.gene.2019.02.079
Martin GW, Harney JW, Berry MJ (1998) Functionality of mutations at conserved nucleotides in eukaryotic SECIS elements is determined by the identity of a single nonconserved nucleotide. RNA 4:65–73
Matsumura S, Ikawa Y, Inoue T (2003) Biochemical characterization of the kink-turn RNA motif. Nucleic Acids Res 31:5544–5551. https://doi.org/10.1093/nar/gkg760
Rayman MP (2000) The importance of selenium to human health. Lancet 356(9225):233–241. https://doi.org/10.1016/S0140-6736(00)02490-9
Sunde RA (2018) Selenium regulation of selenoprotein enzyme activity and transcripts in a pilot study with founder strains from the collaborative cross. PLoS ONE 13:e0191449. https://doi.org/10.1371/journal.pone.0191449
Vendeland SC, Beilstein MA, Yeh JY, Ream W, Whanger PD (1995) Rat skeletal muscle selenoprotein W: cDNA clone and mRNA modulation by dietary selenium. Proc Natl Acad Sci USA 92:8749–8753. https://doi.org/10.1073/pnas.92.19.8749
Walczak R, Carbon P, Krol A (1998) An essential non-Watson-Crick base pair motif in 3’UTR to mediate selenoprotein translation. RNA 4:74–84
Wang L, Zhang XZ, Wu L, Liu Q, Zhang DF, Yin JJ (2018) Expression of selenoprotein genes in muscle is crucial for the growth of rainbow trout (Oncorhynchus mykiss) fed diets supplemented with selenium yeast. Aquaculture 492:82–90. https://doi.org/10.1016/j.aquaculture.2018.03.054
Whanger PD (2000) Selenoprotein W: a review. CMLS. Cell Mol Life Sci 57:1846–1852. https://doi.org/10.1007/pl00000666
Whanger PD (2009) Selenoprotein expression and function-selenoprotein W. Biochim Biophys Acta 1790:1448–1452. https://doi.org/10.1016/j.bbagen.2009.05.010
Xu X, Zhang DG, Zhao T, Xu YH, Luo Z (2020) Characterization and expression analysis of seven selenoprotein genes in yellow catfish Pelteobagrus fulvidraco to dietary selenium levels. J Trace Elem Med Biol 62:126600. https://doi.org/10.1016/j.jtemb.2020.126600
Yeh JY, Ou BR, Gu QP, Whanger P (1998) Influence of gender on selenoprotein w, glutathione peroxidase and selenium in tissues of rats*. Comp Biochem Physiol B 119:151–155. https://doi.org/10.1016/S0305-0491(97)00298-8
Yu D, Li JL, Zhang J, Gao X, Xu SW (2011) Effects of dietary selenium on selenoprotein W gene expression in the chicken immune organs. Biol Trace Elem Res 144:678–687. https://doi.org/10.1007/s12011-011-9062-5
Yu D, Zhang Z, Yao H, Li S, Xu SW (2015) The role of selenoprotein W in inflammatory injury in chicken immune tissues and cultured splenic lymphocyte. Biometals 28:75–87. https://doi.org/10.1007/s10534-014-9804-x
Zhang F, Teng ZL, Wang L, Wang L, Huang TT, Zhang XZ (2021) Dietary selenium deficiency and excess accelerate ubiquitin-mediated protein degradation in the muscle of rainbow trout (Oncorhynchus mykiss) via Akt/FoxO3a and NF-κB Signaling pathways. Biol Trace Elem Res. https://doi.org/10.1007/s12011-021-02726-x
Acknowledgements
This work was financially supported by the National Key R&D Program (Grant Number: 2019YFD0900303) and the Fundamental Research Funds for the Central Universities (Grant Numbers: 2662021SPPY001 and 11900022414).
Author information
Authors and Affiliations
Contributions
CL: conceptualization, investigation, formal analysis, writing-original draft. FZ, ZT and GZ: methodology, formal analysis, writing—review & editing. YY, PX, XH, LW, FY and ZY: methodology, formal analysis. XZ: writing-review & editing, supervision, project administration, funding acquisition. All the authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liao, C., Zhang, F., Teng, Z. et al. Molecular characterization and expression analysis of selenoprotein W gene in rainbow trout (Oncorhynchus mykiss) with dietary selenium levels. Biometals 35, 1359–1370 (2022). https://doi.org/10.1007/s10534-022-00451-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10534-022-00451-z