Abstract
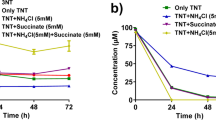
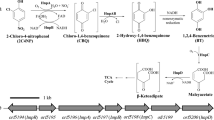
Three bacterial strains utilizing 3-nitrotoluene (3-NT) as a sole source of carbon, nitrogen and energy were isolated from an industrial wastewater treatment plant. Biochemical tests and 16S rDNA sequence analysis revealed that the isolated strains belonged to Diaphorobacter sp. Detailed studies were carried out with Diaphorobacter sp. strain DS2. Degradation of 3-NT by Diaphorobacter sp. strain DS2 was accompanied by the release of nitrite in the culture broth with increase in biomass. Total organic carbon analysis confirmed the extensive mineralization of 3-NT. The strain could degrade 3-methylcatechol, 4-methylcatechol and catechol easily suggesting that the degradation pathway could involve these as possible intermediates. Successful PCR amplification of the oxygenase large subunit and the presence of high activity for catechol 2,3-dioxygenase in the crude cell lysate further confirmed that the degradation of 3-NT occurred through (methyl)catechol intermediates in strain DS2. The strain DS2 was found to degrade other isomers of mononitrotoluene (2-NT and 4-NT) and nitrobenzene as well.





Similar content being viewed by others
References
Ali-Sadat S, Mohan KS, Walia SK (1995) A novel pathway for the biodegradation of 3-nitrotoluene in Pseudomonas putida. FEMS Microbiol Ecol 17:169–176
An D, Gibson DT, Spain JC (1994) Oxidative release of nitrite from 2-nitrotoluene by a three-component enzyme system from Pseudomonas sp. strain JS42. J Bacteriol 176:7462–7467
Arora PK, Sasikala C, Ramana CV (2012) Degradation of chlorinated nitroaromatic compounds. Appl Microbiol Biotechnol 93:2265–2277
Autschul SF, Gish W, Miller W, Myers EM, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Bhattacharya D, Sarma PM, Krishnan S, Mishra S, Lal B (2003) Evaluation of genetic diversity among Pseudomonas citronellolis strains isolated from oily sludge contaminated sites. Appl Environ Microbiol 69:1435–1441
Booth G (2007) Nitro compounds, aromatic. In: Ullmann’s encyclopedia of industrial chemistry. Wiley, New York, pp 1–47. doi:10.1002/14356007.a17_411
Bradford MM (1976) A rapid and sensitive method for the quantization of microgram quantities of protein utilizing the principle of protein–dye binding. Anal Biochem 72:248–254
Cappuccino GJ, Sherman N (1999) Microbiology: a laboratory manual, 4th edn. Addison Wesley, San Francisco
Cerdan P, Wasserfallen A, Rekik M, Timmis KN, Harayama S (1994) Substrate specificity of catechol 2,3-dioxygenase encoded by TOL plasmid pWW0 of Pseudomonas putida and its relationship to cell growth. J Bacteriol 176(19):6074–6081
Cl Kado, Liu ST (1981) Rapid procedure for detection and isolation of large and small plasmids. J Bacteriol 145:1365–1373
Cole JR, Chai B, Farris RJ, Wang Q, Kulam SA, McGarell DM, Garrity GM, Tiedje JM (2005) The Ribosomal Database Project (RDP-II): sequences and tools for high-throughput rRNA analysis. Nucleic Acids Res 1:33
Copley SD (2009) Evolution of efficient pathways for degradation of anthropogenic chemicals. Nat Chem Biol 5:559–566
Dalpozzo R, Bartoli G (2005) Bartoli indole synthesis. Curr Org Chem 9:163–178
Delgado A, Wubbolts MG, Abril MA, Ramos JL (1992) Nitroaromatics are substrates for the TOL plasmid upper-pathway enzymes. Appl Environ Microbiol 58:415–417
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the boot strap. Evolution 39:783–791
Ferraro DJ, Gakhar L, Ramaswamy S (2005) Rieske business: structure-function of Rieske non-heme oxygenase. Biochem Biophys Res Commun 338:175–190
Haigler BE, Spain JC (1993) Biodegradation of 4-nitrotoluene by Pseudomonas sp. strain 4NT. Appl Environ Microbiol 59:2239–2243
Haigler BE, Wallace WH, Spain JC (1994) Biodegradation of 2-nitrotoluene by Pseudomonas sp. strain JS 42. Appl Environ Microbiol 60:3466–3469
Harayama S, Rekik M (1990) The meta cleavage operon of TOL degradative plasmid pWW0 comprises 13 genes. Mol Gen Genet 221:113–120
He Z, Spain JC (1991) Comparison of downstream pathways for degradation of nitrobenzene by Pseudomonas pseudoalcaligenes JS45 (2-aminophenol pathway) and by Comamonas sp. JS765 (catechol pathway). Arch Microbiol 171:309–316
Johnson GR, Jain RK, Spain JC (2002) Origins of the 2,4-dinitrotoluene pathway. J Bacteriol 184:4219–4232
Ju KS, Parales RE (2010) Nitroaromatic compounds: from synthesis to biodegradation. Microbiol Mol Biol Rev 74:250–272
Ju KS, Parales RE (2011) Evolution of a new bacterial pathway for 4-nitrotoluene degradation. Mol Microbiol 82(2):355–364
Kanekar P, Doudpure P, Sarnaik S (2003) Biodegradation of nitroexplosives. Indian J Exp Biol 41:991–1001
Kivissar M (2009) Degaradation of nitroaromatic compounds: a model to study evolution of metabolic pathways. Mol Microbiol 74:777–781
Kivissar M (2011) Evolution of catabolic pathways and their regulatory systems in synthetic nitroaromatic compounds degrading bacteria. Mol Microbiol 82(2):265–268
Klecka GM, Gibson DT (1981) Inhibition of catechol 2,3-dioxygenase from Pseudomonas putida by 3-chlorocatechol. Appl Environ Microbiol 41:1159–1165
Kulkarni M, Chaudhari A (2007) Microbial remediation of nitro-aromatic compounds: an overview. J Environ Manag 85:496–512
Lague D (2005) China blames oil company for benzene spill in river. The New York Times, New York
Lee KS, Parales JV, Friemann R, Parales RE (2005) Active site residues controlling substrate specificity in 2-nitrotoluene dioxygenase from Acdovorax sp. strain JS42. J Ind Microbiol Biotechnol 32:465–473
Lessner DJ, Johnson GR, Parales RE, Spain JC, Gibson DT (2002) Molecular characterization and substrate specificity of nitrobenzene dioxygenase from Comamonas sp. strain JS765. Appl Environ Microbiol 68:634–641
Liu H, Wang SJ, Zhang JJ, Dai H, Tang H, Zhou NY (2011) Patchwork assembly of nag-like nitroarene dioxygenase genes and the 3-chlorocatechol degradation cluster for evolution of the 2-chloronitrobenzene catabolism pathway in Pseudomonas stutzeri ZWLR2-1. Appl Environ Microbiol 77:4547–4552
Mulla SI, Hoskeri RS, Shouche YS, Ninnekar HZ (2010) Biodegradation of 2-nitrotoluene by Micrococcus sp. strain SMN-1. Biodegradation 22:95–102
Nishino SF, Spain JC (1995) Oxidative pathway for the biodegradation of nitrobenzene by Comamonas sp. strain JS765. Appl Environ Microbiol 61:2308–2313
Nishino SF, Spain JC (2004) Catabolism of nitroaromatic compounds. In: Rams JL (ed) Pseudomonas, vol 3. Kluwer Academic/Plenum Publishers, New York, pp 575–608
Nishino N, Atkinson R, Arey J (2008) Formation of nitro products from the gas-phase OH radical-initiated reactions of toluene, naphthalene and biphenyl: effect of NO2 concentration. Environ Sci Technol 42:9203–9209
Padda RS, Wang C, Hughes JB, Kutty R, Bennett GN (2003) Mutagenicity of nitroaromatic degradation compounds. Environ Toxicol Chem 22:2293–2297
Parales JV, Kumar A, Parales RE, Gibson DT (1996) Cloning and sequencing of the genes encoding 2-nitrotoluene dioxygenase from Pseudomonas sp. JS42. Gene 181:57–61
Park H-S, Kim H-S (2000) Identification and characterization of the nitrobenzene catabolic plasmids pNB1 and pNB2 in Pseudomonas putida HS12. J Bacteriol 182:573–580
Robertson JB, Spain JC, Haddock JD, Gibson DT (1992) Oxidation of nitrotoluenes by toluene dioxygenase: evidence for a monooxygenase reaction. Appl Environ Microbiol 58:2643–2648
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sambrook J, Rusell DW (2001) Molecular cloning, vol 1, 3rd edn. CSHL Press, New York
Schmidt M, Teitge M, Castillo ME, Brandt T, Dobner B, Langner A (2008) Synthesis and biochemical characterization of new phenothiazines and related drugs as MDR reversal agents. Arch Pharm Chem Life Sci 341:624–638
Shields MS, Montgomery SO, Chapman PJ, Cuskey SM, Pritchard PH (1989) Novel pathway of toluene catabolism in the trichloroethylene-degrading bacterium G4. Appl Environ Microbiol 55:1624–1629
Shin KA, Spain JC (2009) Pathway and evolutionary implications of diphenylamine biodegradation by Burkholderia sp. strain JS667. Appl Environ Microbiol 75:2694–2704
Spiess T, Desiere F, Fischer P, Spain JC, Knackmuss HJ, Lenke H (1998) A new 4-nitrotoluene degradation pathway in a Mycobacterium strain. Appl Environ Microbiol 64:446–452
Struijs J, Stoltenkamp J (1986) Ultimate biodegradation of 2-, 3-, and 4-nitrotoluene. Sci Total Environ 57:161–170
Swaroop S, Sughosh P, Ramanathan G (2009) Biomineralization of N,N-dimethylformamide by Paracoccus sp. strain DMF. J Hazard Mater 171:268–272
Talmage SS, Opresco DM, Maxevel CZ, Welsh CJE, Cretella EM, Keno PH, Daniel FB (1999) Nitroaromatic munition compounds: environmental effects and screening values. Rev Environ Contam Toxicol 161:1–156
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Walia SK, Ali-Sadat S, Chaudhry GR (2003) Influence of nitro group on biotransformation of nitrotoluenes in Pseudomonas putida strain OU83. Pestic Biochem Physiol 76:73–81
Williams WR, Taylor SC, Williams PA (1993) A novel pathway for the catabolism of 4-nitrotoluene by Pseudomonas. J Gen Microbiol 139:1967–1972
Winkler R, Hertweck C (2007) Biosynthesis of nitro compounds. ChemBioChem 8:973–977
Ye J, Singh A, Ward OP (2004) Biodegradation of nitroaromatics and other nitrogen containing xenobiotics. World J Microbiol Biotechnol 20:117–135
Yen JH, Lin KH, Wang YS (2002) Acute lethal toxicity of environmental pollutants to aquatic organisms. Ecotoxicol Environ Saf 52:113–116
Acknowledgments
Deepak Singh thanks IIT Kanpur for a research fellowship. The department of biotechnology (DBT), India is gratefully acknowledged for partial funding of this project. We are thankful to Hindustan Organic Chemicals Limited for providing sludge from their waste water treatment plant.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Singh, D., Ramanathan, G. Biomineralization of 3-nitrotoluene by Diaphorobacter species. Biodegradation 24, 645–655 (2013). https://doi.org/10.1007/s10532-012-9612-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-012-9612-3