Abstract
Environmental DNA (eDNA) sampling has attracted worldwide attention over the past few years as an emerging approach to characterising and monitoring biodiversity, and has become particularly important for species that are rare, elusive or endangered. Most animal studies to date have focused on aquatic taxa; studies on other metazoan taxa, particularly wildlife in terrestrial environments, are scarce, with only a handful utilizing soil sources. We aimed to investigate the use of DNA barcoding from soil eDNA in (1) detecting rare/elusive/threatened species and (2) as a tool to investigate and potentially monitor range distributions. Through extensive eDNA sampling along the west coast of South Africa, we aimed to refine the distributions of four golden mole species thought to occur there, and specifically to determine whether De Winton’s golden mole, Cryptochloris wintoni (IUCN Critically Endangered; Possibly Extinct), is in fact extant or extinct. Sequences were generated for three barcode markers (mtDNA cyt b, 12S and nuclear GHR) using next-generation amplicon sequencing. Tissue samples from four specimens were used to generate reference sequences for species identification, along with available GenBank sequences. We were able to (1) successfully detect all four species in our data, and (2) improve records of the distributions of these species. Furthermore, we uncovered cryptic diversity in Eremitalpa granti. Our data conclusively reveal the presence of the elusive Cryptochloris wintoni and suggest that this species may in fact be widespread, but not necessarily abundant, and certainly less so in areas subjected to mining activities, which continue to pose a threat to the species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Significant loss of biodiversity has been reported in the current era of rapid global environmental change (Díaz et al. 2019; IPBES 2019), and there is increasing concern worldwide about declines in populations of wildlife (Ripple et al. 2014; Estrada et al. 2017; Jia et al. 2018; Saha et al. 2018; Şekercioğlu et al. 2019); Beckett et al. 2023). Many national and international organizations have established biodiversity monitoring strategies to assess and mitigate the impact of this loss (Kurtz et al. 2001; UN 2003; DEAT 2005). Over the past decade, biodiversity research and monitoring has come to rely increasingly on genetic tools, with rapid and relatively affordable DNA sequencing techniques now offering the opportunity to efficiently characterize biodiversity (Corlett 2017; Alexander et al. 2020; Cowart et al. 2020; Ji et al. 2020; Leempoel et al. 2020; Yang and Zhang 2020; Sales et al. 2021). Among these tools, environmental DNA (eDNA) sampling has attracted worldwide attention (Beng and Corlett 2020) and has become particularly important for species that are rare or elusive (Piaggio et al. 2014; Franklin et al. 2019), endangered (Taberlet et al. 1997; Shelton et al. 2019; Takahara et al. 2020), dangerous (Kendall et al. 2009) or otherwise difficult to capture (Constantine et al. 2012).
Environmental DNA (eDNA) is genetic material originating from the hair, skin, urine, faeces, mucous, saliva, sperm, blood or carcasses of organisms that may be present, in a more or less degraded form, in water, soil, or sediments (Andersen et al. 2012; Taberlet et al. 2012; Taberlet et al. 2012; Thomsen and Willerslev 2015; Pedersen et al. 2016; Sigsgaard et al. 2016; Thomsen and Sigsgaard 2019). DNA can persist in the environment for varying periods of time, from hours in temperate waters, to hundreds or thousands of years in cold, dry permafrost (Thomsen and Willerslev 2015). Isolation of this DNA from the environment can facilitate detection of organisms in the absence of obvious signs of the organism’s presence, and provide genetic information that can be used to identify, study and/or monitor species/individuals across time and space without having to catch, handle, or in some cases, even observe them (Waits and Paetkau 2005; Schwartz et al. 2007; Beja-Pereira et al. 2009).
The application of eDNA has the potential to revolutionize conservation science and practice in many ways (see Beng and Corlett (2020) for a review). Environmental DNA techniques are efficient and relatively cheap and simple, non-destructive and non-invasive, and can be highly effective at detecting rare, cryptic, and elusive species, even at relatively low densities (Carvalho et al. 2019; Franklin et al. 2019; Shelton et al. 2019; Takahara et al. 2020). Recent studies have employed eDNA barcoding in detecting invasive, rare, and cryptic wildlife species, map their distributions, and design management strategies (Levi et al. 2019; Qu and Stewart 2019; Reinhardt et al. 2019). A few have evaluated the efficiency of eDNA versus conventional surveys in detecting elusive species, and eDNA has typically proven comparable or more successful at accurately detecting target species (Deiner et al. 2017; Evans et al. 2017; Leempoel et al. 2020). However, most metazoan eDNA studies have focused on aquatic taxa, especially fishes and amphibians (Beauclerc et al. 2019; Deutschmann et al. 2019). Studies on other metazoan taxa, particularly wildlife in terrestrial environments, are scarce (Beng and Corlett 2020), with some making use of aquatic eDNA sources to investigate terrestrial species (Ushio et al. 2017), and only a handful utilizing soil sources (Andersen et al. 2012; Leempoel et al. 2020).
One particularly interesting group of elusive terrestrial animals that have been extremely challenging to study and/or monitor are the golden moles of sub-Saharan Africa. Golden moles represent a family (Chrysochloridae) of highly threatened small mammals: ten of the 21 species are listed as threatened on the IUCN Red List (1 Critically Endangered, 5 Endangered and 4 Vulnerable), and a further three are listed as Data Deficient (IUCN 2022). These subterranean insectivores are dependent on soft soils for burrowing, and are severely threatened by human activities, such as mining, urbanization and agricultural development. Golden moles are notoriously obscure and understudied, mostly due to the challenges associated with finding them, trapping them and/or observing their subterranean behaviour in the wild. Effective conservation of these animals largely depends on the taxonomic delineation of species, as well as critical information about the distributional ranges and population abundance of the various taxa (Mynhardt et al. 2015; Taylor et al. 2018; Mynhardt et al. 2020; IUCN 2022). Distribution and abundance data for most species are largely lacking (IUCN 2022), and the taxonomy of the family is not well understood (Asher et al. 2010; Taylor et al. 2018).
The first character-based phylogenies of chrysochlorids were based on hyoid shape (Bronner 1991), chromosome morphology (Bronner 1995a), and craniodental anatomy (Bronner 1995b), but these were riddled with uncertainties. A more robust phylogenetic estimate for the family, based on 145 morphological characters from the cranium, dentition, and skeleton, combined with approximately 700–900 bases from exon 10 of the nuclear Growth Hormone Receptor (GHR) gene for 18 of the group’s 21 recognized species, was presented in 2010 (Asher et al. 2010). While this study provided substantial insight into the evolutionary history of golden moles, some uncertainties remained, particularly pertaining to the chrysochlorid root placement, and relationships among unsampled taxa. A new phylogenetic estimate for golden moles and tenrecs is currently underway (Bronner et al. 2023), and will help to resolve some of the taxonomic uncertainties.
De Winton’s golden mole, Cryptochloris wintoni, is a highly elusive species (Critically Endangered; Possibly Extinct; IUCN 2022) recorded only from the type locality at Port Nolloth on the west coast of South Africa (Fig. 1), where it was last seen in 1937, over 80 years ago. The coastal town of Port Nolloth lies in an area of radical habitat transformation by alluvial diamond mining (Bronner and Asher 2016b), which poses a significant threat to golden moles. Cryptochloris wintoni is one of four golden mole species known to occur (or have occurred) on the west coast of southern Africa, alongside the Cape golden mole, Chrysochloris asiatica (Least Concern; IUCN 2022), Grant’s golden mole, Eremitalpa granti (Least Concern; IUCN 2022) and Van Zyl’s golden mole Cryptochloris zyli (Endangered; IUCN 2022; see Fig. 1 for distributions). Chrysoschloris asiatica and E. granti are thought to be relatively abundant and widespread, although little is known about even these species, given the challenges associated with studying their subterranean life (Bronner and Asher 2016a; Maree and Bronner 2016; Taylor et al. 2018). Cryptochloris zyli is known only from the type locality near Lambert's Bay and Groenriviermond (Fig. 1), where it was last seen in 2003 (Bronner and Asher 2016c). Range continuity between these localities cannot be assumed as so little is known about the ecological requirements and tolerances of the species. The estimated extent of occurrence for C. zyli is 4999 km2 with an area of occupancy of 32 km2, although further field assessments are critically needed to obtain more accurate estimates (Bronner and Asher 2016c; IUCN 2022).
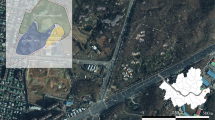
Map of the west coast of southern Africa, depicting our sampling design, as well as the distributions of three of the four golden mole species found along the west coast: Chrysochloris asiatica, Eremitalpa granti (E. g. granti to the South and E. g. namibensis to the North—see inset), and Cryptochloris zyli at Lambert's Bay. Cryptochloris wintoni is known only from the type locality at Port Nolloth, and C. zyli additionally from Groenriviermond. The four tissue samples used in our study were obtained from Lambert’s Bay, Port Nolloth, Papkuilsfontein and Garies. The sampling sites associated with each of the six eDNA sampling regions are colour-coded and indicated on the map, and on separate insets. Region L5 represents a vast stretch of coastline compared to the other regions, in order to encompass the mining areas, where golden mole abundance is substantially lower. Region L6 represents a comparatively small geographic area, focusing on the C. wintoni type locality (Port Nolloth). Since the geographic range of this sample is too small for the scale of the main map, it is indicated within the L4 inset
While C. asiatica is readily identifiable based on external morphology, Cryptochloris spp. may be easily confused with E. granti. Although externally similar, Cryptochloris is well differentiated from E. granti on radiographs based on malleus shape, vertebral count, and length of humeral medial epicondyle (Asher and Avery 2010). Some authors (e.g., Simonetta 1968) have treated C. wintoni as only subspecifically distinct from C. zyli, however these taxa differ consistently in pelage colour and malleus morphology, indicating that they are not conspecific (Meester 1974). Unfortunately C. zyli was one of the three species omitted from the Asher et al. (2010) study, however recent (unpublished) phylogenetic analyses based on both morphological and genetic data support the allocation of the two Cryptochloris taxa to separate species (Bronner and Asher 2016b).
The highly threatened, elusive and poorly understood golden moles on the west coast of South Africa may be considered flagship species for these coastal dune ecosystems, which are being severely impacted by habitat transformation due to ongoing mining and agricultural activities and associated residential developments. Furthermore, they are emblematic of the challenges associated with detecting and monitoring elusive terrestrial wildlife and the potential for eDNA applications to overcome these challenges. In this study, we aimed to evaluate the use of DNA barcoding from environmental DNA (eDNA) collected from soil (1) in detecting rare/elusive/threatened species and (2) as a tool to characterise and monitor range distributions of these species. Through extensive eDNA sampling from soil along the west coast from Lambert's Bay northwards to Port Nolloth, we aimed to refine the distributions of the four golden mole species thought to occur there, and specifically to determine whether C. wintoni is extant or extinct.
Materials and methods
Sample collection
Environmental DNA (eDNA) samples were collected in June 2021 from various sites along the west coast of South Africa (Fig. 1; Provincial permit: CN44-29-14184; Section-20 Permit: SDAH-Epi-21021810220). An initial scouting trip was undertaken to identify potentially active sites, through visual identification of sub-surface tunnels, and to obtain sampling permission from the relevant landowners. Soil samples were subsequently collected from these sites, wherever golden mole activity was detected, with the help of a trained scent-detection dog. GPS co-ordinates were recorded at each site. Wherever possible, soil was scraped from the inner lining of sub-surface tunnels, or alternatively as closely as possible to the furrow, in the case of very loose sand, including where possible, soil from inside the tunnel itself, and from varying depths surrounding the tunnel, up to a total of 15ml of soil per sampling site. Soil samples were collected using gloves and sterile equipment.
A total of 49 soil eDNA samples were collected, and grouped into six sequencing libraries, representing distinct geographic regions along the west coast (Fig. 1). Region L1 consisted of 14 soil samples collected from separate/distinct burrow systems from two distinct sites near Lambert's Bay (Fig. 1). Region L2 comprised 3 soil samples from distinct burrow systems near Kleinsee. Region L3 comprised 10 samples from 3 distinct sites near Groenriviermond. Region L4 comprised 9 samples from three sites on the southern outskirts of Port Nolloth, and Region L6 comprised 6 samples from the northern extent of Port Nolloth. Region L5 consisted of 7 samples from three mining sites in the vicinity of Port Nolloth, and northwards beyond Visagiesfontein towards Alexander Bay; this set of samples covers a far greater geographic range than the others, reflecting the relative dearth of golden mole activity found at the mining sites compared to the other sampling sites.
A single pitfall trap was set at Lambert's Bay (the type locality of Cryptochloris zyli) and at Port Nolloth beach (the type locality of C. wintoni). In each case, a single specimen was captured, and non-invasive DNA samples were collected via buccal and anal swabs, using a sterile cotton bud. Two additional samples were obtained opportunistically from Papkuilsfontein and Garies in the Northern Cape, through independent citizens sending in carcasses of golden moles that had allegedly been killed by pets. In these cases, a small toe clipping was taken for downstream DNA extraction. Since golden moles are morphologically conservative, we used external features (body size, pelage colour and foreclaws) only as an initial means of species identification for specimens, but relied on downstream DNA sequence analysis for more definitive identification.
This study was conducted in accordance with the regulations of the University of Pretoria’s Animal Ethics Committee (ethics clearance no. EC053-18). All samples (tissue, swabs and soil) were stored dry in collection tubes at 4 °C until reaching the lab, where they were frozen at − 20 °C until further processing.
Marker selection
For animal studies, eDNA protocols typically target mitochondrial DNA (mtDNA) rather than nuclear DNA, due to the higher mtDNA copy number in cells, and consequently in the environment (Rees et al. 2014). Mitochondrial DNA barcodes commonly employed in animal studies include cytochrome b (cyt b; Piaggio et al. 2014), 12S ribosomal RNA (Ushio et al. 2017; Leempoel et al. 2020) and cytochrome oxidase I (COI; Hebert et al. 2003). Primers are typically designed to target amplicons (amplified PCR products) of less than 150 bp (base pairs; Rees et al. 2014), based on the assumption that smaller fragments are favourable for optimizing the probability of DNA detection due to environmental degradation of DNA. However, this phenomenon is offset by the improved taxonomic resolution provided by longer fragments. Thus, eDNA barcode markers should aim to target genomic regions with adequate sequence variation, while minimising fragment length. Previous studies have shown that 210 bp of the 12S gene is suitable for successful amplification from soil eDNA, while providing sufficient resolution for mammalian species identification (Leempoel et al. 2020), therefore we targeted this marker, among others, in our study.
Three barcode markers were selected for this study: mitochondrial cytochrome b (cyt b) and 12S, and nuclear GHR intron 9 (Table 1). The markers were chosen based on availability of reference golden mole sequences (cyt b and GHR) and reputability for use in eDNA studies (12S). We used universal mammalian primers for amplification of a 354 bp cyt b fragment, and golden mole-specific 12S and GHR primers were designed using the Primer 3 web interface (Rozen and Skaletzky 2000) using the reference Chrysochloris asiatica genome (ChrAsi1.0; WGS: AMDV00000000.1; Genbank Assembly: GCA_000296735.1) and testing the proposed primers on alignments of all available reference golden mole sequences (12S: 16 of 21 species, including 3 of 4 target species; GHR: all 21 species).
DNA extraction and PCR amplification
All DNA extractions were performed in a standard molecular genetics laboratory at the University of Pretoria. Since we did not have access to a dedicated eDNA laboratory, special care was taken to prevent contamination of samples. Extractions and PCRs were both spatially and temporally separated.
Genomic DNA was extracted from soil samples using a NucleoSpin Soil Kit (Macherey–Nagel, Düren, Germany), and from tissue and swab samples using a DNeasy Blood & Tissue Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. DNA quantity and quality were assessed using gel electrophoresis and QubitTM dsDNA HS and BR Assay Kits with the Qubit 2.0 Fluorometer (Invitrogen, USA).
PCR reactions were performed for each sample and consisted of 50–100 ng DNA, 1 × amplification buffer, 2.5 mM MgCl2, 200 μM of each dNTP (Promega, Johannesburg, South Africa), 0.4 μM of each primer and 1 U Supertherm Taq polymerase (Southern Cross Biotechnology, Cape Town, South Africa). A negative control, using ddH20 instead of DNA, was run with each batch of samples, and PCR products were only retained for samples if the negative control was clear (free of contamination). The cycling parameters for the PCR involved an initial denaturation step of 4 min at 94 °C, followed by 25 cycles of 30 s at 94 °C, 30 s at the optimal annealing temperature for each marker (cyt b: 54 °C; 12S and GHR: 60 °C), and 20 s at 72 °C, and a final extension of 30 min at 72 °C. The quantity and quality of PCR products were assessed using gel electrophoresis and QubitTM dsDNA HS and BR Assay Kits with the Qubit 2.0 Fluorometer (Invitrogen, USA).
DNA sequencing
Cytochrome b PCR amplicons obtained from tissue samples were bi-directionally sequenced for species identification using a BigDye Cycle Sequencing Kit (Applied Biosystems, Foster City, CA, USA) and an automated sequencer (ABI 3500xL Genetic Analyser, Applied Biosystems). PCR amplicons obtained from eDNA samples were sequenced by NGS amplicon sequencing using 600 base chemistry on the Ion Torrent S5 platform at the Central Analytical Facility (CAF) at Stellenbosch University.
For each region, or sequencing library, amplicons from all samples were pooled separately for each barcode marker. Purification of the pooled PCR products was performed using the Agencourt AMPure XP protocol (Beckman Coulter). This was followed by equimolar pooling of the three markers for each library using a QubitTM dsDNA HS Assay Kit with the Qubit 2.0 Fluorometer (Invitrogen, USA) to quantify DNA concentrations. The six pooled regional samples (libraries) were subsequently sent to CAF for library preparation and sequencing on a single sequencing chip. Ideally, negative controls would have been sequenced to provide an idea of background contamination, however the pooling strategy employed in this experiment prevented this. We decided instead to apply a stringent quality and read length filter to remove any contamination artefacts. Since we were more interested in the predominant mammalian species present (which we expected to be golden moles, as samples were taken from golden mole burrows), we were not too concerned about potentially failing to detect other rare species by over-filtering the data.
Bioinformatics
The raw sequence data were quality filtered in Geneious Prime v. 2021.2 (Biomatters Ltd.), using the BBduk plugin, with quality filter set to 30 to remove poor quality reads and size filter set to remove reads shorter than 100 bp. The de novo assembler was used to cluster reads into contigs, or operational taxonomic units (OTUs) using custom sensitivity settings, and the contigs were arranged according to depth of coverage, up to 1000 contigs per regional sample. An initial BLAST search, using the blastn function in Geneious Prime, was used to capture all golden mole OTUs (contigs) for each eDNA sample, for each of the three barcode markers. These data were exported as hit tables to .tsv files, which were subsequently filtered to remove bacterial sequences and reflect only hits with > 90% sequence identity.
Phylogenetic and phylogeographic analyses
All available golden mole reference sequences were downloaded from GenBank and aligned, first with all Sanger-sequences from sampled individuals, and then with all captured contigs from the six eDNA samples. Alignments were built separately for each barcode marker using MEGA X v.11.0.11 (Tamura et al. 2021). MEGA software was further used for model selection and subsequent phylogenetic analysis. Maximum Likelihood analysis was conducted for each marker, using the optimal BIC (Bayesian Information Criterion) model, BioNJ/NJ starting tree and 1000 bootstrap iterations. To account for missing data, we used partial deletion with a threshold of 50%, thus separate analyses were conducted for the Sanger sequenced samples and the eDNA contigs, since shorter reads among the eDNA contigs could potentially reduce the overall resolution of the tree. For the Sanger sequenced samples, all available golden mole reference sequences were included in the analyses, so as to provide context regarding each marker’s taxonomic resolution within the family Chrysochloridae.
Statistical parsimony (TCS) haplotype networks were constructed using PopArt (Clement et al. 2000; Leigh and Bryant 2015). Cytochrome b sequence alignments were grouped into four partitions to prevent data loss resulting from missing data in a few samples/contigs. Partition A represents the full 354 bp sequence: Partition B: nt164-329 (165 bp); Partition C: nt222-331 (109 bp) and Partition D: nt217-354 (137 bp). Although there is overlap in nucleotide position between the partitions, each contig was only assigned to one of the four partitions, so that the same contig would not be reflected multiple times across the four partitions, in order to facilitate haplotype counting.
Results
Bioinformatics and preliminary species identification
The raw eDNA sequence data contained an average of 473,294 reads per regional sample. The BBduk function removed on average 279,759 reads (59%) and retained on average 193,535 reads (41%), on the basis of read length and quality (Table S1). This is a large reduction in the data, but we preferred to filter thoroughly to minimize persistence of putative sequencing errors, even at the risk of data loss (see Materials and Methods–DNA Sequencing for the rationale behind this).
De novo assembly produced contigs with coverage ranging from 60,599× to 157,600× in contig 1 (mean of 121,419× across all 6 samples) and mean coverage of 85× in contig 100 and 8× in contig 1000. We provide a comprehensive list of hits from the BLAST searches conducted in Geneious (Table S2; note that contig numbers do not correspond to those in Fig. 3 and 4, as the BLAST searches were re-run after analyses were conducted, in order to accommodate newly available GenBank reference sequences). These data should be interpreted with caution, due to the possibility of contaminants, and incorrect BLAST hits, owing to the paucity of reference sequences for some taxa.
Initial BLAST searches detected all four golden mole species thought to occur on the west coast among each of the six regions analysed. In addition, some contigs showed highest similarity to Amblysomus golden moles, but since these represent species from previous projects in our lab, with distribution ranges well outside our study area, it is possible that these may represent contaminant sequences. This hypothesis is supported by our finding of Bryde’s whale (Balaenoptera brydei; Table S2: L2 and L4), riverine rabbit (Bunolagus monticularis; L2, L4) mole-rats (Cryptomys hottentotus; L2, L3, L4, L5), and eland (Tragelaphus oryx; L1 and L4) sequences in our data, also species from other projects concurrently running in our lab. Additional species that were detected with high confidence in our data include human, dog, chicken, Lomi's blind legless skink (Typhlosaurus lomii; L4), wild turkey (Meleagris gallopavo; L5) and yellow-spotted rock hyrax (Heterohyrax brucei; L6).
BLAST searches for cyt b sequences generated for the four golden mole specimens from Lambert's Bay, Papkuilsfontein, Garies and Port Nolloth showed highest similarity to Eremitalpa granti (98, 51% sequence identity), Chrysochloris asiatica (97,49%), E. granti (90,39%) and Cryptochloris wintoni (98,88%), respectively.
Maximum likelihood phylogenetic analysis
Maximum likelihood analysis of cyt b grouped both the Lambert′s Bay sample and the Garies sample with Eremitalpa granti (Fig. 2a, blue shading); the Lambert's Bay sample grouped closely with E. granti granti (95% bootstrap support) and the Garies sample as sister to these (with weak bootstrap support), to the exclusion of the only other E. granti subspecies, E. granti namibensis. The Papkuilsfontein sample grouped with Chrysochloris asiatica (Fig. 2a, yellow shading), and yet appears to be somewhat divergent from reference sequences (genetic distance to nearest C. asiatica AJ428944.1 = 0.0302). The sample from Port Nolloth groups with Cryptochloris wintoni, sister to C. zyli (Fig. 2a, green shading).
Maximum Likelihood bootstrap consensus trees of sampled specimens as inferred by the Tamura-Nei model for cyt b (a) and the General Time Reversible model for 12S (b). Bootstrap values above 50% are shown at nodes. West coast taxa are indicated by coloured shading: Chrysochloris asiatica: yellow; Eremitalpa granti: blue; Cryptochloris spp. (C. wintoni and C. zyli): green
Analysis of the 12S marker corroborated the placement of the samples from Garies with stronger support (99%) for the relationships within E. granti (Fig. 2b, blue shading). The placement of the Port Nolloth sample was also corroborated as sister to C. zyli in the absence of a reference sequence for C. wintoni (Fig. 2b, blue shading). The other two samples—Lambert's Bay and Papkuilsfontein—were not sequenced for 12S, as their species identity had already been confidently established through cyt b analysis.
In the eDNA, Eremitalpa granti was detected in regions L2 and L4 (cyt b and 12S; Fig. S1a and b) and in L3, L5 and L6 (cyt b only; Fig. S1a) and Cryptochloris wintoni in regions L1 through L6 (cyt b; Fig. S1a) and L2 through L4 (12S; Fig. S1b). Chrysochloris asiatica was detected in L2 through L4 (cyt b; Fig. S1a), but was not detected with 12S (see Discussion for comparison of relative performance of markers). Resolution at the GHR marker was poor, and many samples failed to provide golden mole sequences at all, partly due to poor amplification success. Only regions L1, L4, L5 and L6 contained golden mole sequences, but species identification for these was problematic due to weak marker resolution. Thus, this marker was omitted from further analyses.
Haplotype network phylogeographic analysis
TCS network analysis revealed that the cyt b barcode detected one E. granti granti haplotype in Region L1 (Fig. 3b), three in Region L2 (Fig. 3a), three in Region L3 (Fig. 3a), two in Region L4 (Fig. 3c), four in Region L5 (Fig. 3a) and two in Region L6 (Fig. 3b). Thus E. granti granti was detected at each sampling site, and represents the most frequently detected taxon in our cyt b dataset (Fig. 5). Furthermore, the Garies specimen is grouped within Eremitalpa but is clearly distinct from both E. g. granti and E. g. namibensis (Fig. 3a). Chysochloris asiatica was not detected in Region L1, but two haplotypes were detected in Region L2 (Fig. 3a), one in Region L3 (Fig. 3a) and two in Region L4 (Fig. 3c).
TCS haplotype networks representing the evolutionary relationships among the 47 golden mole haplotypes detected across four partitions (A–D) of our cyt b dataset. The dataset comprises eDNA samples (contigs from six sampling regions along the west coast, tissue samples from live specimens (denoted as locality_ “sample”) and GenBank reference sequences. Circle sizes reflect the number of samples sharing the same haplotype; mutational steps between haplotypes are indicated by cross-hatching, and missing haplotypes are denoted by black dots. Colours correspond to eDNA sampling regions as presented in Fig. 1, with tissue samples shown in grey and reference sequences in white. Background shading roughly denotes broader taxonomic groups: Chrysochloris asiatica: yellow; Eremitalpa granti: blue; Cryptochloris spp. (C. wintoni and C. zyli): green
The only available reference Cryptochloris sequences (C. zyli KM388924.1 and C. wintoni KM388925.1) were included in the analysis, but the C. wintoni reference sequence contains missing data, and could therefore only be included in partition B (Fig. 3b). However, since the specimen from Port Nolloth was already confidently assigned to C. wintoni (98.88% sequence similarity as mentioned above), we treated this as a C. wintoni reference across the other partitions. A single Cryptochloris haplotype was found in Region L1, (Fig. 3b), three haplotypes in Region L2 (Fig. 3a–c), three in Region L3 (Fig. 3a, d), three in Region L4 (Fig. 3b, c), five in Region L5 (Fig. 3a–d) and three in Region L6 (Fig. 3a, c, d). Thus, C. wintoni was detected at each of the six sampling sites (Fig. 5). However, partition D did not provide sufficient resolution to distinguish C. wintoni from C. zyli, therefore this particular haplotype, detected in regions L3, L5 and L6, remains unresolved and is denoted as “Cryptochloris sp.” (Fig. 5).
The 12S barcode detected one E. granti granti haplotype in Region L2 and three in Region L4 (Fig. 4), corroborating cyt b detection of this taxon at Kleinsee and Port Nolloth (Fig. 5). The 12S marker did not detect Chrysochloris asiatica in any of the eDNA samples. Cryptochloris was the most frequently detected taxon in this dataset, with one haplotype detected in Region L2, three in Region L3, and three in Region L4, again corroborating cyt b findings. A further three haplotypes were detected in Regions L2 and L4, which were grouped within a Chrysochloris-Cryptochloris clade in ML analysis (66% bootstrap support; Fig. S1b), but which appear to be divergent from both taxonomic groups, and could not be confidently assigned to either (Fig. 4).
TCS haplotype networks representing the evolutionary relationships among the 20 golden mole 12S haplotypes detected in our dataset. The dataset comprises eDNA samples (contigs from six sampling regions along the west coast, tissue samples from live specimens (denoted as locality_ “sample”) and GenBank reference sequences. Circle sizes reflect the number of samples sharing the same haplotype; mutational steps between haplotypes are indicated by cross-hatching, and missing haplotypes are denoted by black dots. Colours correspond to eDNA sampling regions as presented in Fig. 1, with tissue samples shown in grey and reference sequences in white. Background shading roughly denotes broader taxonomic groups: Chrysochloris asiatica: yellow; Eremitalpa granti): blue; Cryptochloris spp. (C. wintoni and C. zyli): green
Map of the west coast of southern Africa, depicting the golden mole species detected at each of six sampling regions (L1-6). Pie chart sizes reflect the number of cyt b haplotypes detected in each regional sample, and coloured wedges reflect the proportion of haplotypes representing each of the four species. Since some haplotypes could not be confidently assigned to either C. zyli or C. wintoni, the fourth species category is denoted as “Cryptochloris sp.”
The GHR marker detected only a single golden mole haplotype in the eDNA samples, in Regions L1, L4, L5 and L6. This haplotype is shared with C. asiatica references and the Papkuilsfontein sample. However, resolution at this marker was poor, therefore, taxonomic designations could not be assigned on the basis of this marker.
Discussion
We investigated the use of DNA barcoding from environmental DNA (eDNA) collected from soil in detecting and investigating the distributions of rare/elusive species. We used tissue samples from trapped and opportunistically obtained specimens, along with GenBank sequences as references for taxonomic designation in our study. PCR amplicons for three barcode markers from soil eDNA samples collected at various sampling sites along the west coast of southern Africa were pooled into 6 sequencing libraries, or “regional samples” (see Methods – Sample Collection for details). Pooling of samples is common practice in eDNA studies, and generally facilitates reduced sequencing costs, by reducing the number of unique identifiers, which are required for each sample. Pooling may be achieved through direct mixing of soil samples (Taberlet et al. 2012), or pooling of PCR amplicons (Andersen et al. 2012; Leempoel et al. 2020). In our study we used the latter approach, involving separate PCR reactions for each sample and each barcode marker, which may be more labour intensive, but allows equimolar pooling of the various barcode amplicons, resulting in relatively even abundance in the sequencing reaction. It is important to note that due to the pooling of samples from various different sites into a single regional sample, the occurrence of multiple species in a single sample does not necessary indicate sympatry of these species, since a single species may have been detected at the first site, another at a second site and yet another at a third site, and when pooled into a single sample, we are not able to determine which specific sampling site each species was detected in. This sampling design was aimed at maximising detection of all golden mole species across the west coast, while testing the effectiveness of the eDNA sampling technique, and broadly refining the distributions of the various taxa.
Given the limited availability of reference sequences for golden moles, we targeted three genes (cyt b, 12S and GHR) as barcode markers for our study. As expected, the two mitochondrial markers far outperformed the nuclear intron in terms of both amplification success and taxonomic resolution. This is not surprising, given the relative abundance of mtDNA in animal cells, and consequently in the environment (Rees et al. 2014), compared to nuclear DNA. We were able to detect golden mole GHR sequences in Regions L1, L4, L5 and L6, but species assignment was problematic due to weak marker resolution. Thus, we were able to demonstrate successful amplification of mammalian nuclear eDNA from soil in our study, but the nuclear intron marker did not provide sufficient taxonomic resolution. Despite equimolar pooling of amplicons, we observed a significant bias in representation of mtDNA markers, and therefore in future studies it will be advisable to refrain from pooling nuclear and mtDNA fragments together for amplicon sequencing. Furthermore, cyt b outperformed 12S in the detection of golden mole species; all four species were detected in our data, whereas 12S failed to detect Chrysochloris asiatica in any of the samples. However, 12S provided good resolution when amplification was successful; we detected at least six Cryptochloris 12S haplotypes in our data, compared to only three Cryptochloris cyt b haplotypes. In contrast we detected at least nine Eremitalpa granti cyt b haplotypes, compared to only four 12S haplotypes. Taken together, both mtDNA markers performed comparatively well in terms of taxonomic resolution, and the failure of 12S to detect C. asiatica in our data may be explained by primer specificity and poor amplification success in this species.
Due to high mtDNA sequence similarity between C. wintoni and C. zyli, in some cases our relatively short barcode markers failed to distinguish between these sister species. Thus, we cannot confirm whether or not C. zyli was detected in our dataset. Unresolved Cryptochloris sequences were detected in Regions L3, L5 and L6, thus C. zyli may well be extant at these sites (Groenriviermond, Port Nolloth and Visagiesfontein). It is plausible that the two Cryptochloris species in fact represent a single species, however more data (additional samples and more sequence data) will be required to test this hypothesis. Limited evidence currently supports their allocation to separate species (Meester 1974; Bronner and Asher 2016b).
Our data have conclusively revealed the presence of the elusive Cryptochloris wintoni (IUCN Red List Critically Endangered; Possibly Extinct) on the west coast of southern Africa. Moreover, our data suggest that this species may be widespread in the area, ranging from Lambert's Bay in the south to Visagiesfontein (beyond Port Nolloth) in the north; we detected C. wintoni mtDNA haplotypes at each of our six sampling sites (Fig. 5). However, this cannot be taken to mean that the species is abundant across this distribution. On the contrary, we found relatively few cyt b haplotypes; no more than two in any given partition (Fig. 3). Generally, this was comparable to the one or two haplotypes detected for C. asiatica, but fewer than in E. granti (up to 9 haplotypes; Fig. 3). Although 12S detected six Cryptochloris haplotypes across the six regional samples, relative to only four for E. granti and none for C. asiatica, resolution at this marker was insufficient to distinguish C. wintoni from C. zyli, and these six haplotypes may represent both Cryptochloris species (Fig. 4).
The 12S barcode failed to detect Chrysochloris asiatica, however cyt b detected three to five C. asiatica haplotypes in Regions L2 through L4, i.e. Groenriviermond, Kleinsee and Port Nolloth (Fig. 4, Fig. 5), and additionally at Papkuilsfontein. These sites all fall within the known range of the species, however, we were surprised that C. asiatica was not detected at Lambert's Bay (L1), since this species is thought to be increasingly abundant further south towards Cape Town (Bronner and Asher 2016a; Taylor et al. 2018). Our relatively low detection of this species may be due to a sampling bias favouring the coastal dune habitats, which are more characteristic of Eremitalpa and Cryptochloris species. Cape golden moles are known to inhabit a wide range of habitat types, depending on soil friability and availability of invertebrate prey (Bronner and Asher 2016a; Taylor et al. 2018), and we suspect that the apparent sympatry of the four golden mole species along the west coast may be explained by subtle ecological niche preferences.
Eremitalpa granti was detected at all six sites, and additionally at Garies, all within the species’ known distribution (Fig. 5). Specifically, 9–13 E. g. granti haplotypes were detected in the eDNA samples across the first three cyt b partitions, and four from the 12S barcode. Eremitalpa g. namibensis was not detected in our eDNA samples, however TCS analysis for both cyt b and 12S placed the Garies sample within E. granti, but distinct from both E. granti subspecies. This may indicate that the sample from Garies represents a third cryptic subspecies. It is also interesting to note the high level of sequence divergence between the two E. granti subspecies, relative to the level of divergence between the two Cryptochloris species, indicating that E. g. granti and E. g. namibensis likely represent distinct species, in which case the Garies sample could represent a third cryptic species. Once again, more data (additional samples representing a broader geographic range and/or more sequence data) will be required to test this hypothesis, and to establish the taxonomic placement of the Garies sample.
Conclusion
Our study demonstrates the use of DNA barcoding from environmental DNA (eDNA) collected from soil (1) in detecting rare/elusive species and (2) as a tool to investigate and potentially monitor their distributions. Through extensive eDNA sampling along the west coast from Lambert's Bay northwards to Port Nolloth, we have not only rediscovered the “lost” De Winton’s golden mole, moreover we demonstrate that this species may be widespread along the west coast, albeit in low abundance. Given that this species is listed as critically endangered (IUCN 2022) and occurs in an area under high threat of habitat transformation by alluvial diamond mining (Bronner and Asher 2016b), it will be important to gather additional data to (1) resolve the potential taxonomic uncertainty within Cryptochloris, (2) further refine the distribution and investigate abundance throughout the distribution, (3) determine presence vs. absence in protected areas, as well as areas under particularly high threat, and (4) assess the viability of sub-populations, and potential for connectivity between them.
The sampling design for the current study was aimed at maximising detection of all golden mole species across the west coast, while testing the effectiveness of the eDNA sampling technique, and broadly refining the distributions of the various taxa. We detected Eremitalpa granti and Cryptochloris wintoni in all six regional samples, each consisting of around 10 pooled samples from 2 to 5 proximate sampling sites. Unresolved Cryptochloris, potentially including C. zyli was detected in Regions L3, L5 and L6 (Kleinsee, Port Nolloth and Visagiesfontein), and Chrysochloris asiatica was detected in Regions L2 through L4 (Groenriviermond, Kleinsee and Port Nolloth). Thus, we were able to broadly investigate the distributions of the four golden mole species on the west coast, however, fine-scale sampling, targeting specific habitats or ecological niches, along with unpooled amplicon sequencing will be required to further refine the distributions of these taxa. In the meantime, regional and species-specific conservation action is both critical and urgent, in order to protect not only the threatened golden mole populations this study has revealed, but also the highly threatened west coast dune ecosystems they inhabit.
Data availability
The data generated and analysed in the current study are available on GenBank (accession numbers: OP279338-OP279343). Additional data generated but not analysed in the current study may be found in the supplementary information, and/or upon reasonable request from the authors.
References
Alexander JB, Bunce M, White N, Wilkinson SP, Adam AA, Berry T, Stat M, Thomas L, Newman SJ, Dugal L (2020) Development of a multi-assay approach for monitoring coral diversity using eDNA metabarcoding. Coral Reefs 39:159–171
Andersen K, Bird KL, Rasmussen M, Haile J, Breuning-Madsen H, Kjær KH, Orlando L, Gilbert MTP, Willerslev E (2012) Meta-barcoding of ‘dirt’ DNA from soil reflects vertebrate biodiversity. Mol Ecol 21:1966–1979
Asher RJ, Avery DM (2010) New golden moles (Afrotheria, Chrysochloridae) from the early pliocene of South Africa. Palaeontol Electron 13:12
Asher RJ, Maree S, Bronner G, Bennett NC, Bloomer P, Czechowski P, Meyer M, Hofreiter M (2010) A phylogenetic estimate for golden moles (Mammalia, Afrotheria, Chrysochloridae). BMC Evol Biol 10:69
Avenant N (2011) The potential utility of rodents and other small mammals as indicators of ecosystem “integrity” of South African grasslands. Wildl Res 38:626–639
Basset Y, Cizek L, Cuénoud P, Didham RK, Guilhaumon F, Missa O, Novotny V, Ødegaard F, Roslin T, Schmidl J (2012) Arthropod diversity in a tropical forest. Science 338:1481–1484
Beauclerc K, Wozney K, Smith C, Wilson C (2019) Development of quantitative PCR primers and probes for environmental DNA detection of amphibians in Ontario. Cons Gen Resour 11:43–46
Beckett H, Hansen OK, von der Heyden S, Midgley GF (2023) A natural terminal Pleistocene decline of African penguin populations enhances their anthropogenic extinction risk. African Journal of Marine Science 45:57–62
Beja-Pereira A, Oliveira R, Alves PC, Schwartz MK, Luikart G (2009) Advancing ecological understandings through technological transformations in noninvasive genetics. Mol Ecol Res 9:1279–1301
Beng KC, Corlett RT (2020) Applications of environmental DNA (eDNA) in ecology and conservation: opportunities, challenges and prospects. Biodivers Conserv 29:2089–2121
Bronner GN (1991) Comparative hyoid morphology of nine chrysochlorid species (Mammalia: Chrysochloridae). Annals of the Transvaal Museum 35:295–311
Bronner GN (1995a) Cytogenetic properties of nine species of golden moles (Insectivora: Chrysochloridae). J Mammal 76:957–971
Bronner GN, Asher R (2016a) A conservation assessment of Chrysochloris asiatica. In: Child MFRL, Do Linh San E, Raimondo D, Davies-Mostert HT (eds) The Red List of Mammals of South Africa. South African National Biodiversity Institute and Endangered Wildlife Trust., Swaziland and Lesotho South Africa
Bronner GN, Asher R (2016b) A conservation assessment of Cryptochloris wintoni. In: Child MFRL, Do Linh San E, Raimondo D, Davies-Mostert HT (eds) The Red List of Mammals of South Africa. South African National Biodiversity Institute and Endangered Wildlife Trust, Swaziland and Lesotho South Africa
Bronner GN, Asher R (2016c) A conservation assessment of Cryptochloris zyli. In: Child MFRL, Do Linh San E, Raimondo D, Davies-Mostert HT (eds) The Red List of Mammals of South Africa. South African National Biodiversity Institute and Endangered Wildlife Trust, Swaziland and Lesotho South Africa
Bronner GN, Mynhardt S, Bennett NC, Cohen L, Crumpton N, Hofreiter M, Arnold P, Asher RJ (2023) Phylogenetic history of golden moles and tenrecs (Mammalia: Afrotheria). Zool J. https://doi.org/10.1093/zoolinnean/zlad121
Bronner GN (1995b) Systematic revision of the Golden mole genera: Amblysomus, Chlorotalpa & Chrysochloris, University of Natal, Durban
Carvalho S, Aylagas E, Villalobos R, Kattan Y, Berumen M, Pearman JK (2019) Beyond the visual: using metabarcoding to characterize the hidden reef cryptobiome. Proc R Soc B 286:20182697
Clement M, Posada D, Crandall K (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1660
Constantine R, Jackson JA, Steel D, Baker CS, Brooks L, Burns D, Clapham P, Hauser N, Madon B, Mattila D (2012) Abundance of humpback whales in Oceania using photo-identification and microsatellite genotyping. Mar Ecol Prog Ser 453:249–261
Corlett RT (2017) A bigger toolbox: biotechnology in biodiversity conservation. Trends Biotechnol 35:55–65
Cowart DA, Matabos M, Brandt MI, Marticorena J, Sarrazin J (2020) Exploring environmental dna (edna) to assess biodiversity of hard substratum faunal communities on the lucky strike vent field (mid-atlantic ridge) and investigate recolonization dynamics after an induced disturbance. Front Mar Sci. https://doi.org/10.3389/fmars.2019.00783
DEAT (2005) South Africa's National Biodiversity Strategy and Action Plan. In: Department of Environmental Affairs and Tourism (DEAT) Pretoria
Deiner K, Bik HM, Mächler E, Seymour M, Lacoursière-Roussel A, Altermatt F, Creer S, Bista I, Lodge DM, de Vere N (2017) Environmental DNA metabarcoding: transforming how we survey animal and plant communities. Mol Ecol 26:5872–5895
Deutschmann B, Müller A-K, Hollert H, Brinkmann M (2019) Assessing the fate of brown trout (Salmo trutta) environmental DNA in a natural stream using a sensitive and specific dual-labelled probe. Sci Total Environ 655:321–327
Díaz S, Settele J, Brondízio ES, Ngo HT, Agard J, Arneth A, Balvanera P, Brauman KA, Butchart SH, Chan KM (2019) Pervasive human-driven decline of life on Earth points to the need for transformative change. Science 366:eaax3100
Estrada A, Garber PA, Rylands AB, Roos C, Fernandez-Duque E, Di Fiore A, Nekaris KA-I, Nijman V, Heymann EW, Lambert JE (2017) Impending extinction crisis of the world’s primates: Why primates matter. Science advances 3:e1600946
Evans NT, Shirey PD, Wieringa JG, Mahon AR, Lamberti GA (2017) Comparative cost and effort of fish distribution detection via environmental DNA analysis and electrofishing. Fisheries 42:90–99
Franklin TW, McKelvey KS, Golding JD, Mason DH, Dysthe JC, Pilgrim KL, Squires JR, Aubry KB, Long RA, Greaves SE (2019) Using environmental DNA methods to improve winter surveys for rare carnivores: DNA from snow and improved noninvasive techniques. Biol Conserv 229:50–58
Gomez-Zurita J, Cardoso A, Coronado I, De la Cadena G, Jurado-Rivera JA, Maes J-M, Montelongo T, Nguyen DT, Papadopoulou A (2016) High-throughput biodiversity analysis: rapid assessment of species richness and ecological interactions of Chrysomelidae (Coleoptera) in the tropics. ZooKeys 597:3
Hebert PD, Cywinska A, Ball SL, DeWaard JR (2003) Biological identifications through DNA barcodes. Proc R Soc London Ser B: Biol Sci 270:313–321
IPBES (2019) The global assessment report on biodiversity and ecosystem services of the Intergovernmental Science-Policy Platform on Biodiversity and Ecosystem Services: Summary for policymakers. Díaz S, Settele J, Brondízio ES, Ngo HT, Guèze M, Agard J, Arneth A, Balvanera P, Brauman KA, Butchart SHM, Chan KMA, Garibaldi LA, Ichii K, Liu J, Subramanian SM, Midgley GF, Miloslavich P, Molnár Z, Obura D, Pfaff A, Polasky S, Purvis A, Razzaque J, Reyers B, Chowdhury R, Shin YJ, Visseren-Hamakers IJ, Willis KJ, and Zayas CN (eds.). IPBES secretariat, Bonn, Germany.
Ji Y, Baker CC, Li Y, Popescu VD, Wang Z, Wang J, Wang L, Wu C, Hua C, Yang Z (2020) Large-scale quantification of vertebrate biodiversity in ailaoshan nature reserve from leech iDNA. BioRxiv. https://doi.org/10.1101/2020.02.10.941336
Jia Q, Wang X, Zhang Y, Cao L, Fox AD (2018) Drivers of waterbird communities and their declines on Yangtze River floodplain lakes. Biol Conserv 218:240–246
Kendall KC, Stetz JB, Boulanger J, Macleod AC, Paetkau D, White GC (2009) Demography and genetic structure of a recovering grizzly bear population. J Wildl Manag 73:3–16
Khelifa R (2019) Sensitivity of biodiversity indices to life history stage, habitat type and landscape in Odonata community. Biol Conserv 237:63–69
Kocher TD, Thomas WK, Meyer A, Edwards SV, Pääbo S, Villablanca FX, Wilson AC (1989) Dynamics of mitochondrial DNA evolution in animals: amplification and sequencing with conserved primers. Proc Nat Acad Sci 86:6196–6200
Kurtz JC, Jackson LE, Fisher WS (2001) Strategies for evaluating indicators based on guidelines from the environmental protection agency’s office of research and development. Ecol Indic 1:49–60
Leempoel K, Hebert T, Hadly EA (2020) A comparison of eDNA to camera trapping for assessment of terrestrial mammal diversity. Proc R Soc B 287:20192353
Leigh JW, Bryant D (2015) POPART: full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116
Levi T, Allen JM, Bell D, Joyce J, Russell JR, Tallmon DA, Vulstek SC, Yang C, Yu DW (2019) Environmental DNA for the enumeration and management of Pacific salmon. Mol Ecol Res 19:597–608
Maree S, Bronner GN (2016) A conservation assessment of Eremitalpa granti granti. In: Child MF, Roxburgh LDLSE, Raimondo D, Davies-Mostert HT (eds) The Red List of Mammals of South Africa. South African National Biodiversity Institute and Endangered Wildlife Trust, Swaziland and Lesotho South Africa
Meester JAJ (1974) Family Chrysochloridae. In: Meester J, Setzer HW (eds) The mammals of Africa: an identification manual. Smithsonian Institution Press, Washington D.C, pp 1–7
Mynhardt S, Maree S, Pelser I, Bennett NC, Bronner GN, Wilson JW, Bloomer P (2015) Phylogeography of a morphologically cryptic golden mole assemblage from South-Eastern Africa. PLoS ONE 10:e0144995
Mynhardt S, Bennett NC, Bloomer P (2020) New insights from RADseq data on differentiation in the hottentot golden mole species complex from South Africa. Mol Phylogenet Evol 143:106667
Outhwaite CL, Gregory RD, Chandler RE, Collen B, Isaac NJ (2020) Complex long-term biodiversity change among invertebrates, bryophytes and lichens. Nat Ecol Evol 4:384–392
Ovaskainen O, Moliterno de Camargo U, Somervuo P (2018) Animal sound identifier (ASI): software for automated identification of vocal animals. Ecol Lett 21:1244–1254
Pedersen MW, Ruter A, Schweger C, Friebe H, Staff RA, Kjeldsen KK, Mendoza ML, Beaudoin AB, Zutter C, Larsen NK (2016) Postglacial viability and colonization in North America’s ice-free corridor. Nature 537:45–49
Piaggio AJ, Engeman RM, Hopken MW, Humphrey JS, Keacher KL, Bruce WE, Avery ML (2014) Detecting an elusive invasive species: a diagnostic PCR to detect Burmese python in Florida waters and an assessment of persistence of environmental DNA. Mol Ecol Res 14:374–380
Qu C, Stewart KA (2019) Evaluating monitoring options for conservation: comparing traditional and environmental DNA tools for a critically endangered mammal. Sci Nat 106:1–9
Rajan SC, Athira K, Jaishanker R, Sooraj N, Sarojkumar V (2019) Rapid assessment of biodiversity using acoustic indices. Biodivers Conserv 28:2371–2383
Rees HC, Maddison BC, Middleditch DJ, Patmore JRM, Gough KC (2014) REVIEW: The detection of aquatic animal species using environmental DNA—a review of eDNA as a survey tool in ecology. J Appl Ecol 51:1450–1459
Reinhardt T, van Schingen M, Windisch HS, Nguyen TQ, Ziegler T, Fink P (2019) Monitoring a loss: detection of the semi-aquatic crocodile lizard (Shinisaurus crocodilurus) in inaccessible habitats via environmental DNA. Aquat Conserv: Mar Freshw Ecosyst 29:353–360
Ripple WJ, Estes JA, Beschta RL, Wilmers CC, Ritchie EG, Hebblewhite M, Berger J, Elmhagen B, Letnic M, Nelson MP (2014) Status and ecological effects of the world’s largest carnivores. Science 343:1241484
Rodríguez-Estrella R, Estrada CG, Alvarez-Castañeda ST, Ferrer-Sánchez Y (2019) Comparing individual raptor species and coarse taxonomic groups as biodiversity surrogates in desert ecosystems. Biodivers Conserv 28:1225–1244
Rozen S, Skaletzky H (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386
Saha A, McRae L, Dodd CK Jr, Gadsden H, Hare KM, Lukoschek V, Böhm M (2018) Tracking global population trends: population time-series data and a living planet index for reptiles. J Herpetol 52:259–268
Sales NG, Wangensteen OS, Carvalho DC, Deiner K, Præbel K, Coscia I, McDevitt AD, Mariani S (2021) Space-time dynamics in monitoring neotropical fish communities using eDNA metabarcoding. Sci Total Environ 754:142096
Schwartz MK, Luikart G, Waples RS (2007) Genetic monitoring as a promising tool for conservation and management. Trends Ecol Evol 22:25–33
Şekercioğlu ÇH, Mendenhall CD, Oviedo-Brenes F, Horns JJ, Ehrlich PR, Daily GC (2019) Long-term declines in bird populations in tropical agricultural countryside. Proc Nat Acad Sci 116:9903–9912
Shelton AO, Kelly RP, O’Donnell JL, Park L, Schwenke P, Greene C, Henderson RA, Beamer EM (2019) Environmental DNA provides quantitative estimates of a threatened salmon species. Biol Conserv 237:383–391
Sigsgaard EE, Nielsen IB, Bach SS, Lorenzen ED, Robinson DP, Knudsen SW, Pedersen MW, Al Jaidah M, Orlando L, Willerslev E (2016) Population characteristics of a large whale shark aggregation inferred from seawater environmental DNA. Nat Ecol Evol 1:1–5
Simonetta A (1968) A new golden mole from Somalia with an appendix on the taxonomy of the family Chrysochloridae (Mammlia, Insectivora). Monitore Zoologico Italiana 2:27–55
Steenweg R, Hebblewhite M, Kays R, Ahumada J, Fisher JT, Burton C, Townsend SE, Carbone C, Rowcliffe JM, Whittington J (2017) Scaling-up camera traps: monitoring the planet’s biodiversity with networks of remote sensors. Front Ecol Environ 15:26–34
Stoeckle BC, Kuehn R, Geist J (2016) Environmental DNA as a monitoring tool for the endangered freshwater pearl mussel (Margaritifera margaritifera L.): a substitute for classical monitoring approaches? Aquat Conserv: Mar Freshw Ecosyst 26:1120–1129
Taberlet P, Camarra JJ, Griffin S, Uhres E, Hanotte O, Waits L, Dubois-Paganon C, Burke T, Bouvet J (1997) Noninvasive genetic tracking of the endangered Pyrenean brown bear population. Mol Ecol 6:869–876
Taberlet P, Coissac E, Hajibabaei M, Rieseberg LH (2012) Environmental DNA. Mol Ecol. https://doi.org/10.1111/j.1365-294X.2012.05542.x
Taberlet P, Prud’homme SM, Campione E, Roy J, Miquel C, Shehzad W, Gielly L, Rioux D, Choler P, Clément JC (2012) Soil sampling and isolation of extracellular DNA from large amount of starting material suitable for metabarcoding studies. Mol Ecol 21:1816–1820
Takahara T, Iwai N, Yasumiba K, Igawa T (2020) Comparison of the detection of 3 endangered frog species by eDNA and acoustic surveys across 3 seasons. Freshw Sci 39:18–27
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027
Taylor WA, Mynhardt S, Maree S (2018) Family Chrysochloridae (Golden Moles). Handbook of the Mammals of the World. Lynx Edicions, Cerdanyola del Vallès, pp 180–203
The IUCN Red List of Threatened Species Version 2022–1 [Internet] (2022) http://www.iucnredlist.org Accessed 2022
Thomsen PF, Sigsgaard EE (2019) Environmental DNA metabarcoding of wild flowers reveals diverse communities of terrestrial arthropods. Ecol Evol 9:1665–1679
Thomsen PF, Willerslev E (2015) Environmental DNA—an emerging tool in conservation for monitoring past and present biodiversity. Biol Conserv 183:4–18
UN (2003) United Nations Environment Programme Convention on Biological Diversity SBSTTA. Monitoring and indicators: Designing national-level monitoring programmes and indicators. In: Montreal, United Nations. Canada
Ushio M, Fukuda H, Inoue T, Makoto K, Kishida O, Sato K, Murata K, Nikaido M, Sado T, Sato Y et al (2017) Environmental DNA enables detection of terrestrial mammals from forest pond water. Mol Ecol Res 17:e63–e75
Van Der Heyde M, Bunce M, Wardell-Johnson G, Fernandes K, White NE, Nevill P (2020) Testing multiple substrates for terrestrial biodiversity monitoring using environmental DNA metabarcoding. Mol Ecol Res 20:732–745
Waits LP, Paetkau D (2005) Noninvasive genetic sampling tools for wildlife biologists: a review of applications and recommendations for accurate data collection. J Wildl Manag 69:1419–1433
Yang J, Zhang X (2020) eDNA metabarcoding in zooplankton improves the ecological status assessment of aquatic ecosystems. Environ Int 134:105230
Zhang Y, Pavlovska M, Stoica E, Prekrasna I, Yang J, Slobodnik J, Zhang X, Dykyi E (2020) Holistic pelagic biodiversity monitoring of the Black Sea via eDNA metabarcoding approach: from bacteria to marine mammals. Environ Int 135:105307
Acknowledgements
We thank Alexkor, the Richtersveld Municipality, De Beers and all the landowners involved for granting access to sites. We acknowledge our trusted scent detection dog and team member, Jessie, for assistance in finding golden mole tunnels, and Insauf de Vries for assistance with administrative tasks. We acknowledge the Central Analytical Facility (CAF) at Stellenbosch University (SUN) for high quality amplicon sequencing.
Funding
Open access funding provided by Stellenbosch University. This work was funded in part by Re:wild and in part by the National Research Foundation (NRF) Foundational Biodiversity Information Programme (FBIP Unique grant number 136334; Prof. Paulette Bloomer) and supported by Rand Merchant Bank. We also acknowledge the National Research Foundation (NRF) South African Research Chair Initiative (SARChI) Chair of Behavioural Ecology and Physiology (Prof. Nigel Bennett) for postdoctoral funding (SM). The opinions, findings and conclusions expressed in this publication are those of the authors and the NRF accepts no responsibility in this regard.
Author information
Authors and Affiliations
Contributions
SM and CT designed and conceptualized the project. SM, PB and CT acquired the funding. SM and EM obtained permits and ethical clearance and JPLR acquired landowner consent. CT planned and led the field work and SM, EM, JPLR, IL and CT collected the data. SM analyzed the data and wrote the first draft of the manuscript. PB and IL commented on previous versions of the manuscript and provided advice and assistance throughout the project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by David Hawksworth.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10531_2023_2728_MOESM1_ESM.docx
Fig. S1 Maximum likelihood bootstrap consensus trees of sampled specimens and eDNA contigs for cyt b (a) 12S (b) (DOCX 137 KB)
10531_2023_2728_MOESM2_ESM.pdf
Table S1 Bioinformatic filtering of environmental DNA (eDNA) NGS data using the BBDuk function in geneious prime (PDF 163 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mynhardt, S., Matthew, E., le Roux, J.P. et al. Environmental DNA from soil reveals the presence of a “lost” Afrotherian species. Biodivers Conserv 33, 31–50 (2024). https://doi.org/10.1007/s10531-023-02728-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10531-023-02728-2