Abstract
Purpose
Hepatocellular carcinoma (HCC) is the uncontrolled growth of hepatocytes which results in nearly 5 million deaths worldwide. Specific strategies have been developed to treat HCC, including surgery, chemotherapy and radiotherapy. But, the effective disease dealing requires synergistic collaboration with other approaches, which often results in moderate to severe side effects during and after the treatment period. Therefore, the focus is now shifting to explore and retrieve those plant-based products that could be utilized to treat HCC with maximum efficacy without causing any side effects. Strigolactones (SL) are compounds of plant origin derived from Striga lutea responsible for controlling the branching pattern of stem and have reported anti-cancerous activity by promoting apoptosis at micromolar concentrations. However, little work has been done concerning determining the pharmacogenomic effect of strigolactones on HCC.
Methods
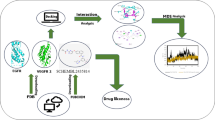
Current work focuses on comparing therapeutic efficiencies of SL analogs against core targets of HCC using network pharmacology approach, pharmacokinetics analysis, gene ontogeny, functional enrichment analysis, molecular docking and Molecular Dynamics simulation.
Results
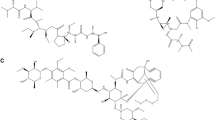
Drug-target prediction and functional enrichment analysis showed that HDAC1 and HDAC2 are the core proteins involved in hepatocellular carcinoma that strigolactone analogs can target. Consequently, results from molecular docking and MD simulation analyses report that among all the SL analogs strigol, epistrigol and nijmegen1 can turn out to be most effective in downregulating the expression of HDAC1, HDAC2 and CYP19A.
Conclusion
Strigol, epistrigol and nijmegen1 could be used as potential inhibitors against HCC and can be further validated through in vitro/in vivo studies.
















Similar content being viewed by others
Data availability
Supplementary information or data can be obtained from the author on request.
References
Bhal SK (2007) LogP—Making sense of the value. Advanced Chemistry Development Toronto, pp 1–4
Bhattacharyya GS, Govind Babu K, Malhotra H, Ranade AA, Murshed S, Datta D (2013) Hepatocellular carcinoma in India. Chinese Clin Oncol 2:3–7. https://doi.org/10.3978/j.issn.2304-3865.2013.09.05
Cheatham TEIII, Miller JL, Fox T, Darden TA, Kollman PA (1995) Molecular dynamics simulations on solvated biomolecular systems: the particle mesh Ewald method leads to stable trajectories of DNA, RNA, and proteins. J Am Chem Soc 117:4193–4194. https://doi.org/10.1021/ja00119a045
Cheng F, Li W, Zhou Y, Wu JSZ, Liu G, Lee PW, Tang Y (2012) admetSAR: a comprehensive source and free tool for assessment of chemical ADMET properties. J Chem Inf Model 52:3099–3105. https://doi.org/10.1021/ci300367a
Choudhary P, Gupta S, Shukla R, Gupta A, Pahal S (2021) Regulation of neuronal repair and regeneration through inhibition of oligodendrocyte myelin glycoprotein (OMgp). J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2021.1997820
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:42717. https://doi.org/10.1038/srep42717
Dennis G, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA (2003) DAVID: database for annotation, visualization, and integrated discovery. Genome Biol 4:R60. https://doi.org/10.1186/gb-2003-4-9-r60
dos Santos JL (2015) Pan-Assay Interference Compounds (PAINS): warning signs in biochemical-pharmacological evaluations. Biochem Pharmacol Open Access. https://doi.org/10.4172/2167-0501.1000e173
Ertl P, Schuffenhauer A (2009) Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions. J Cheminform 1:8. https://doi.org/10.1186/1758-2946-1-8
Finch A, Pillans P (2014) P-glycoprotein and its role in drug-drug interactions. Aust Prescr 37:137–139. https://doi.org/10.18773/austprescr.2014.050
Franz M, Rodriguez H, Lopes C, Zuberi K, Montojo J, Bader GD, Morris Q (2018) GeneMANIA update 2018. Nucleic Acids Res 46:60–64. https://doi.org/10.1093/nar/gky311
Galicia-Moreno M, Silva-Gomez JA, Lucano-Landeros S, Santos A, Monroy-Ramirez HC, Armendariz-Borunda J (2021) Liver cancer: therapeutic challenges and the importance of experimental models. Can J Gastroenterol Hepatol 2021:1–10. https://doi.org/10.1155/2021/8837811
Hamosh A (2004) Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res 33:514–517. https://doi.org/10.1093/nar/gki033
Hasan MN, Razvi SS, Choudhry H, Hassan MA et al (2019) Gene ontology and expression studies of strigolactone analogues on a hepatocellular carcinoma cell line. Anal Cell Pathol 2019:1–10. https://doi.org/10.1155/2019/1598182
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472. https://doi.org/10.1002/(SICI)1096-987X(199709)18:12%3c1463::AID-JCC4%3e3.0.CO;2-H
Hirshman SP (1983) Steepest-descent moment method for three-dimensional magnetohydrodynamic equilibria. Phys Fluids 26:3553. https://doi.org/10.1063/1.864116
Huang J, MacKerell AD (2013) CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J Comput Chem 34:2135–2145. https://doi.org/10.1002/jcc.23354
Kim S, Chen J, Cheng T, Gindulyte A et al (2021) PubChem in 2021: new data content and improved web interfaces. Nucleic Acids Res 49:1388–1395. https://doi.org/10.1093/nar/gkaa971
Kumar A, Acharya SK, Singh SP, Arora A et al (2020) 2019 update of Indian National Association for study of the liver consensus on prevention, diagnosis, and management of hepatocellular carcinoma in India: the Puri II recommendations. J Clin Exp Hepatol 10:43–80. https://doi.org/10.1016/j.jceh.2019.09.007
Liu Z, Guo F, Wang Y, Li C et al (2016) BATMAN-TCM: a bioinformatics analysis tool for molecular mechanism of traditional Chinese medicine. Sci Rep 6:21146. https://doi.org/10.1038/srep21146
Lu Z (2011) PubMed and beyond: a survey of web tools for searching biomedical literature. Database 2011:036–036. https://doi.org/10.1093/database/baq036
Lurje I, Czigany Z, Bednarsch J, Roderburg C, Isfort P, Neumann UP, Lurje G (2019) Treatment strategies for hepatocellular carcinoma – a multidisciplinary approach. Int J Mol Sci 20:1465. https://doi.org/10.3390/ijms20061465
Merle P, Trepo C (2009) Molecular mechanisms underlying hepatocellular carcinoma. Viruses 1:852–872. https://doi.org/10.3390/v1030852
Mishra D, Panda G, Kumar P, Singh S (2017) Novel drug delivery system for herbal formulation in cancer treatment. World J Pharm Res 6:341–353. https://doi.org/10.20959/wjpr201715-10093
Mishra S, Rastogi SK, Singh S, Panwar SL, Shrivash MK, Misra K (2019) Controlling pathogenesis in Candida albicans by targeting Efg1 and Glyoxylate pathway through naturally occurring polyphenols. Mol Biol Rep 46:5805–5820. https://doi.org/10.1007/s11033-019-05014-z
Mishra S, Singh S (2017) Identification of inhibitors against metastasis protein “Survivin:” in silico discovery using virtual screening and molecular docking studies. Pharmacogn Mag 13:S742. https://doi.org/10.4103/pm.pm_178_17
Mishra S, Tripathi R, Singh S (2016) Crosstalk of proteins, miRNAs involved in metastatic and epithelial–mesenchymal transition pathways. Front Life Sci 9:323–346. https://doi.org/10.1080/21553769.2016.1256843
Morris G, Goodsell D, Pique M, et al (2012) User guide autodock version 4.2. Autom Docking Flex Ligands to Flex Recept AutoGrid
Oliveros JC (2007) VENNY. An interactive tool for comparing lists with Venn Diagrams. https://bioinfogp.cnb.csic.es/tools/venny/index.html
Pahal S, Gupta A, Choudhary P, Chaudhary A, Singh S (2021) Network pharmacological evaluation of Withania somnifera bioactive phytochemicals for identifying novel potential inhibitors against neurodegenerative disorder. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2021.1951355
Perlman L, Gottlieb A, Atias N, Ruppin E, Sharan R (2011) Combining drug and gene similarity measures for drug-target elucidation. J Comput Biol 18:133–145. https://doi.org/10.1089/cmb.2010.0213
Raza A (2014) Hepatocellular carcinoma review: current treatment, and evidence-based medicine. World J Gastroenterol 20:4115. https://doi.org/10.3748/wjg.v20.i15.4115
Richon VM (2006) Cancer biology: mechanism of antitumour action of vorinostat (suberoylanilide hydroxamic acid), a novel histone deacetylase inhibitor. Br J Cancer 95:2–6. https://doi.org/10.1038/sj.bjc.6603463
Safran M, Dalah I, Alexander J et al (2010) GeneCards Version 3: the human gene integrator. Database 2010:020–020. https://doi.org/10.1093/database/baq020
Schaftenaar G, de Vlieg J (2012) Quantum mechanical polar surface area. J Comput Aided Mol Des 26:311–318. https://doi.org/10.1007/s10822-012-9557-y
Schüttelkopf AW, van Aalten DMF (2004) PRODRG : a tool for high-throughput crystallography of protein–ligand complexes. Acta Crystallogr Sect D Biol Crystallogr 60:1355–1363. https://doi.org/10.1107/S0907444904011679
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage T, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Shukla R, Singh S, Singh A, Misra K (2021) Two pronged approach for prevention and therapy of COVID-19 (Sars-CoV-2) by a multi-targeted herbal drug, a component of ayurvedic decoction. Eur J Integr Med 43:101268. https://doi.org/10.1016/j.eujim.2020.101268
Tang Y, Li M, Wang J, Pan Y, Wu F (2015) CytoNCA: a cytoscape plugin for centrality analysis and evaluation of protein interaction networks. Biosystems 127:67–72. https://doi.org/10.1016/j.biosystems.2014.11.005
Vieira TF, Sousa SF (2019) Comparing AutoDock and Vina in ligand/decoy discrimination for virtual screening. Appl Sci 9:4538. https://doi.org/10.3390/app9214538
Wang Y, Bouwmeester HJ (2018) Structural diversity in the strigolactones. J Exp Bot 69:2219–2230. https://doi.org/10.1093/jxb/ery091
Yi Y, Fang Y, Wu K, Liu Y, Zhang W (2020) Comprehensive gene and pathway analysis of cervical cancer progression. Oncol Lett. https://doi.org/10.3892/ol.2020.11439
Zwanenburg B, Pospíšil T, Ćavar Zeljković S (2016) Strigolactones: new plant hormones in action. Planta 243:1311–1326. https://doi.org/10.1007/s00425-015-2455-5
Acknowledgements
Authors are grateful to Director, Indian Institute of Information Technology, Allahabad, India for providing facilities for research and to Ministry of Education, India for providing research fellowship. The computational results reported in this work were performed on the Central Computing Facility of IIIT Allahabad.
Author information
Authors and Affiliations
Contributions
AA performed analysis and written manuscript. SP helped in performed analysis and done proofreading, PC and AG performed MD simulations analysis, SS designed this study.
Corresponding author
Ethics declarations
Confilct of interests
Authors have no conflict of interest.
Ethical approval
Compliance with ethical standard of research.
Consent to participate
This study does not involve any animals or humans as subject for experimentation.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Amod, A., Pahal, S., Choudhary, P. et al. Network pharmacological evaluation of strigolactones efficacy as potential inhibitors against therapeutic targets of hepatocellular carcinoma. Biotechnol Lett 44, 879–900 (2022). https://doi.org/10.1007/s10529-022-03266-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-022-03266-7