Abstract
Objectives
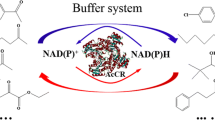
Alteration of the cofactor specificity of acrylyl-CoA reductase (AcuI) catalyzing the NAD(P)H-dependent reduction of acrylyl-CoA to propionyl-CoA is often desirable for designing of artificial metabolic pathways of various appointments.
Results
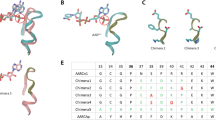
Several variants of AcuIs from Escherichia coli K-12 with multiple amino acid substitutions to alter the cofactor preference were obtained by site directed mutagenesis and the modified enzymes as His6-tagged proteins were characterized. The simultaneous substitutions of arginine-180, arginine-198 and serine-178 residues by alanine in the enzyme pocket sequence as well as other amino acid changes decreased both NADPH- and NADH-dependent activities in comparison to the wild-type enzyme. The replacement of serine-156 by glutamic acid decreased NADPH-dependent activity at least 7000-fold but NADH-dependent activity only by threefold. The replacement of serine-156 by aspartic acid decreased NADPH-dependent activity 70-fold with fair preservation of activity and specificity to NADH.
Conclusions
These results demonstrated a relevance of Asp156 in the interaction of AcuI from E. coli K-12 with NADH as a coenzyme. These findings may provide reference information for shifting coenzyme specificity of acrylyl-CoA reductases.



Similar content being viewed by others
References
Asao M, Alber BE (2013) Acrylyl-Coenzyme A reductase, an enzyme involved in the assimilation of 3-hydroxypropionate by Rhodobacter sphaeroides. J Bacteriol 195:4716–4725
Berg IA, Kockelkorn D, Ramos-Vera WH, Say RF, Zarzycki J, Hügler M, Alber BE, Fuchs G (2010) Autotrophic carbon fixation in archaea. Nat Rev Microbiol 8:447–460
Curson AR, Burns OJ, Voget S, Daniel R, Todd JD, McInnis K, Wexler M, Johnston AW (2014) Screening of metagenomic and genomic libraries reveals three classes of bacterial enzymes that overcome the toxicity of acrylate. PLoS ONE 9:e97660
Herrmann G, Selmer T, Jessen HJ, Gokarn RR, Selifonova O, Gort SJ, Buckel W (2005) Two beta-alanyl-CoA:ammonia lyases in Clostridium propionicum. FEBS J 272:813–821
Hetzel M, Brock M, Selmer T, Pierik AJ, Golding BT, Buckel W (2003) Acryloyl-CoA reductase from Clostridium propionicum an enzyme complex of propionyl-CoA dehydrogenase and electron-transferring flavoprotein. Eur J Biochem 270:902–910
Kandasamy V, Vaidyanathan H, Djurdjevic I, Jayamani E, Ramachandran KB, Buckel W, Jayaraman G, Ramalingam S (2013) Engineering Escherichia coli with acrylate pathway genes for propionic acid synthesis and its impact on mixed-acid fermentation. Appl Microbiol Biotechnol 97:1191–1200
Kuchta RD, Abeles RH (1985) Lactate reduction in Clostridium propionicum. Purification and properties of lactyl-CoA dehydratase. J Biol Chem 260:13181–13189
Meng H, Liu P, Sun H, Cai Z, Zhou J, Lin J, Li Y (2016) Engineering a d-lactate dehydrogenase that can super-efficiently utilize NADPH and NADH as cofactors. Sci Rep 6:24887
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera: a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Schneider K, Asao M, Carter MS, Alber BE (2012) Rhodobacter sphaeroides uses a reductive route via propionyl coenzyme A to assimilate 3-hydroxypropionate. J Bacteriol 194:225–232
Seo JK, Kim S-W, Kim MH, Upadhaya SD, Kam DK, Ha JK (2010) Direct-fed microbials for ruminant animals. Asian-Aust J Anim Sci 23:1657–1667
Sullivan MJ, Curson AR, Shearer N, Todd JD, Green RT, Johnston AW (2011) Unusual regulation of a leaderless operon involved in the catabolism of dimethylsulfoniopropionate in Rhodobacter sphaeroides. PLoS ONE 6:e15972
Sulzenbacher G, Roig-Zamboni V, Pagot F, Grisel S, Salomoni A, Valencia C, Campanacci V, Vincentelli R, Tegoni M, Eklund H, Cambillau C (2004) Structure of Escherichia coli YhdH, a putative quinone oxidoreductase. Acta Crystallogr D Biol Crystallogr 60:1855–1862
Todd JD, Curson ARJ, Sullivan MJ, Kirkwood M, Johnston AW (2012) The Ruegeria pomeroyi acuI gene has a role in DMSP catabolism and resembles yhdH of E. coli and other bacteria in conferring resistance to acrylate. PLoS ONE 7:e35947
Vallejo AN, Pogulis RJ, Pease LR (2008) PCR mutagenesis by overlap extension and gene SOE. Cold Spring Harbor Protocols 2: pdb.prot4861
Wang P, Cao HY, Chen X-L, Li CY, Li PY, Zhang XY, Qin QL, Todd JD, Zhang YZ (2017) Mechanistic insight into acrylate metabolism and detoxification in marine dimethylsulfoniopropionate-catabolizing bacteria. Mol Microbiol 105:674–688
Acknowledgements
This work was supported by Russian Foundation for Basic Research, project 18-29-07044 MK
Author information
Authors and Affiliations
Contributions
ASR conceived and designed the experiments, analyzed, collected and interpreted all the data obtained by other authors, obtained point mutations of the acuI gene. SYB determined the activity of mutant forms of the AcuI enzyme. IIM performed the part of the work related to the synthesis and chromatographic purification of acrylyl CoA. ONR performed the part of the work related to modeling the tertiary structure of the mutated AcuI enzyme. VNK is responsible for all parts of the work, resolving issues related to the accuracy and integrity of the work. All authors take part in the final approval of the version of the article for publication.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflicts of interest with the current work or its publication.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Reshetnikov, A.S., But, S.Y., Rozova, O.N. et al. Alteration of cofactor specificity of the acrylyl-CoA reductase from Escherichia coli. Biotechnol Lett 43, 1421–1427 (2021). https://doi.org/10.1007/s10529-021-03130-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-021-03130-0