Abstract
Objective
Superoxide dismutase (SOD) enzyme has implications in modulating the cell’s redox state. The study aims to explore the host genetic factors that limit the heterologous expression of a thermostable SOD from Potentilla atrosanguinea (Pa-SOD) in E. coli.
Results
It was observed that the heterologous expression of Pa-SOD in E. coli did not exhibit any enhancement after 1 h of induction. This led to the alteration in cell morphology and an increase in the doubling time of E. coli cells expressing Pa-SOD. Label-free quantification and MALDI-TOF/TOF-MS/MS analysis suggested differential expression of 81 proteins, of which 77 proteins were found to be downregulated and 4 were found to be upregulated in Pa-SOD expressing cells as compared to uninduced E. coli cells. Functional analysis of downregulated proteins shows involvement in molecular function, biological process, and were the part of a cellular component. The STRING database revealed interaction of an essential autoregulatory protein, RNase E with other proteins involved in biosynthetic processes, protein biosynthesis and folding, and cell division. Further, validation of RNase E protein revealed upregulation of rne at transcript level and downregulation of RNase E at protein level as compared to uninduced cells.
Conclusions
The observations suggested the operation of multifaceted mechanisms with a key role of RNase E that regulated the expression of Pa-SOD at the physiological and molecular level. Since Pa-SOD has commercial applications, identification and manipulation of these networked genetic factors could lead to improvement of host strain for large-scale production of biologically active Pa-SOD and other heterologous proteins.



Similar content being viewed by others
References
Adams DW, Wu LJ, Errington J (2014) Cell cycle regulation by the bacterial nucleoid. Curr Opin Microbiol 22:94–101
Addinall SG, Lutkenhaus J (1996) FtsA is localized to the septum in an FtsZ-dependent manner. J Bacteriol 178:7167–7172
Allard STM, Giraud MF, Naismith JH (2001) Epimerases: structure, function and mechanism. Cell Mol Life Sci 58:1650–1665
Anderson OR, Rogerson A, Hannah F (1996) A description of the testate amoeba Ovulina parva gen. nov., sp. nov. from coastal marine sediments. J Mar Biol Ass UK 76:851–865
Arraiano CM, Andrade JM, Domingues S, Guinote IB, Malecki M, Matos RG et al (2010) The critical role of RNA processing and degradation in the control of gene expression. FEMS Microbiol Rev 34:883–923
Bafana A, Dutt S, Kumar S, Ahuja PS (2011) Superoxide dismutase: an industrial perspective. Crit Rev Biotechnol 31:65–76
Baneyx F, Mujacic M (2004) Recombinant protein folding and misfolding in Escherichia coli. Nat Biotechnol 22:1399–1408
Baumgardt K, Charoenpanich P, McIntosh M, Schikora A, Stein E, Thalmann S et al (2014) RNase E affects the expression of the acyl-homoserine lactone synthase gene sinI in Sinorhizobium meliloti. J Bacteriol 196:1435–1447
Bhardwaj PK, Sahoo R, Kumar S, Ahuja PS (2011) A gene encoding autoclavable super-oxide dismutase and its expression in E. coli. US patent 7888088B2.
Bouvier J, Richaud C, Richaud F, Patte JC, Stragier P (1984) Nucleotide sequence and expression of the Escherichia coli dapB gene. J Biol Chem 259:14829–14834
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analyt Biochem 72:248–254
Campbell TL, Brown ED (2002) Characterization of the depletion of 2-C-methyl-D-erythritol-2, 4-cyclodiphosphate synthase in Escherichia coli and Bacillus subtilis. J bacterial 184:5609–5918
Carrier MJ, Nugent ME, Tacon WCA, Primrose SB (1983) High expression of cloned genes in E. coli and its consequences. Trends Biotechnol 1:109–113
Chou CP (2007) Engineering cell physiology to enhance recombinant protein production in Escherichia coli. Appl Microbiol Biotechnol 76:521–532
Deutscher MP (2015) How bacterial cells keep ribonucleases under control. FEMS Microbiol Rev 39:350–361
Dewar SJ, Dorazi R (2000) Control of division gene expression in Escherichia coli. FEMS Microbiol Lett 187:1–7
Dri AM, Rouviere-Yaniv J, Moreau PL (1991) Inhibition of cell division in hupA hupB mutant bacteria lacking HU protein. J Bacteriol 173:2852–2863
El-Hajj ZW, Newman EB (2015) An Escherichia coli mutant that makes exceptionally long cells. J Bacteriol 197:1507–1514
Ghora BK, Apirion D (1978) Structural analysis and in vitro processing to p5 rRNA of a 9S RNA molecule isolated from an rne mutant of E. coli. Cell 15:1055–1066
Glick BR (1995) Metabolic load and heterogenous gene expression. Biotechnol Adv 13:247–261
Grabherr R, Nilsson E, Striedner G, Bayer K (2002) Stabilizing plasmid copy number to improve recombinant protein production. Biotechnol Bioeng 77:142–147
Huang S, Deutscher MP (1992) Sequence and transcriptional analysis of the Escherichia coli rnt gene encoding RNase T. J Biol Chem 267:25609–25613
Hughes D (2016) Using the power of genetic suppressors to probe the essential functions of RNase E. Curr Genet 62:53–57
Kang L, Shaw AC, Xu D, Xia W, Zhang J, Deng J et al (2011) Upregulation of MetC is essential for D-alanine-independent growth of an alr/dadX-deficient Escherichia coli strain. J bacterial 193:1098–1106
Kaur J, Kumar A, Kaur J (2018) Strategies for optimization of heterologous protein expression in E. coli: roadblocks and reinforcements. Int J Biol Macromol 106:803–822
Kumar S, Sahoo R, Ahuja PS (2006) Isozyme of autoclavable superoxide dismutase (SOD), a process for the identification and extraction of the SOD and use of the said SOD in the cosmetic, food and pharmaceutical compositions. US Patent 7037697
Kumar A, Dutt S, Bagler G, Ahuja PS, Kumar S (2012) Engineering a thermo-stable superoxide dismutase functional at sub-zero to >50 °C, which also tolerates autoclaving. Sci Rep 2:387
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lees N, Cozzitorto J, Wainwright N, Testa D (1984) Cloning with tandem gene systems for high level gene expression. Nucleic Acids Res 12:6797–6812
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Mackie GA (2013) RNase E: at the interface of bacterial RNA processing and decay. Nat Rev Microbiol 11:45–57
McCord JM, Fridovich I (1969) Superoxide dismutase an enzymic function for erythrocuprein (hemocuprein). J Biol Chem 244:6049–6055
Miroux B, Walker JE (1996) Over-production of proteins in Escherichia coli: mutant hosts that allow synthesis of some membrane proteins and globular proteins at high levels. J Mol Biol 260:289–298
Mittler R (2017) ROS are good. Trends in Plant Sci 22:11–19
Murashko ON, Lin-Chao S (2017) Escherichia coli responds to environmental changes using enolasic degradosomes and stabilized DicF sRNA to alter cellular morphology. Proc Natl Acad Sci USA 114:E8025–E8034
Neuberger P, Lin HY, Mathiszik B (2003) Metabolic load of recombinant protein production: inhibition of cellular capacities for glucose uptake and respiration after induction of a heterologous gene in Escherichia coli. Biotechnol Bioeng 83:53–64
Ortiz C, Natale P, Cueto L, Vicente M (2015) The keepers of the ring: regulators of FtsZ assembly. FEMS Microbiol Rev 40:57–67
Ow MC, Kushner SR (2002) Initiation of tRNA maturation by RNase E is essential for cell viability in E. coli. Genes Dev 16:1102–1105
Pellizza L, Smal C, Rodrigo G, Arán M (2018) Codon usage clusters correlation: towards protein solubility prediction in heterologous expression systems in E. coli. Sci Rep 8:10618
Perez-Riverol Y, Csordas A, Bai J, Bernal-Llinares M, Hewapathirana S, Kundu DJ et al (2019) The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res 47:D442–450
Rabinowitch HD, Fridovich I (1983) Superoxide radicals, superoxide dismutases and oxygen toxicity in plants. Photochem Photobiol 37:679–690
Rai M, Padh H (2001) Expression systems for production of heterologous proteins. Curr Sci 80:1121–1128
Rosano GL, Ceccarelli EA (2014) Recombinant protein expression in Escherichia coli: advances and challenges. Front Microbiol 5:172
Sahdev S, Khattar SK, Saini KS (2008) Production of active eukaryotic proteins through bacterial expression systems: a review of the existing biotechnology strategies. Mol Cell Biochem 307:249–264
Schein CH (1989) Production of soluble recombinant proteins in bacteria. Nat Biotechnol 7:141–1149
Sharma S, Ray S, Moiyadi A, Sridhar E, Srivastava S (2014) Quantitative proteomic analysis of meningiomas for the identification of surrogate protein markers. Sci Rep 4:7140
Shojaosadati SA, Varedi Kolaei SM, Babaeipour V, Farnoud AM (2008) Recent advances in high cell density cultivation for production of recombinant protein. Iran J Biotechnol 6:63–84
Singh D, Chang SJ, Lin P-H, Averina OV, Kaberdin VR, Lin-Chao S (2009) Regulation of ribonuclease E activity by the L4 ribosomal protein of Escherichia coli. Proc Natl Acad Sci USA 106:864–869
Singh V, Singh B, Joshi R, Jaju P, Pati PK (2017) Changes in the leaf proteome profile of Withania somnifera (L.) Dunal in response to Alternaria alternata infection. PLoS ONE 12:e0178924
Sánchez-Gorostiaga A, Palacios P, Martínez-Arteaga R, Sánchez M, Casanova M, Vicente M (2016) Life without division: physiology of Escherichia coli FtsZ-deprived filaments. M Bio 7:e01620–e1716
Yogavel M, Mishra PC, Gill J, Bhardwaj PK, Dutt S, Kumar S, Ahuja PS, Sharma A (2008) Structure of a superoxide dismutase and implications for copper-ion chelation. Acta Crystallogr D Biol Crystallogr 64:892–901
Acknowledgements
SG is thankful to Indian council of Medical Research for Junior and Senior Research Fellowships. Authors gratefully acknowledge Dr Sue Lin-Chao, Institute of Molecular Biology, Academia Sinica, Taipei, Taiwan for providing RNase E antibodies as a gift. Financial support from CSIR network projects (Grant No. BSC0109, BSC0209) and in-house grant (MLP201) is duly acknowledged. The manuscript represents CSIR-IHBT communication number 4148.
Supporting Information
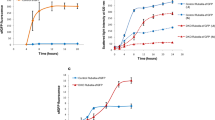
Fig. S1 Effect of Pa-SOD expression on growth of E. coli at 37 °C and 28 °C. Graphs on left represent E. coli growth curve at 37 °C while graphs on right represent E. coli growth curve at 28 °C a, b Growth curve of E. coli M15 strain harboring empty pQE30 vector (control). c, d Growth curve of E. coli M15 strain harboring recombinant pQE-Pa-SOD. e, f Growth curve of E. coli BL21 (DE3) harboring recombinant pET47b-Pa-SOD. The results are expressed as means ± SE of the means, n=3.
Fig. S2 SDS-PAGE analysis of differentially expressed proteins upon induction of Pa-SOD. M represents protein molecular weight marker (kDa). pQE-Pa-SOD/M15 indicates total protein obtained from E. coli M15 strain harboring pQE-Pa-SOD recombinant plasmid (genetically engineered mutant of Pa-SOD WT) before induction at 0 h and after induction at 3 h (lanes 2-3). The arrow indicates the differentially expressed proteins.
Fig. S3 Relative expressions of rne, ispF, ftsZ, ftsA, and Pa-SOD at 0, 1, 3 and 5 h of induction as analyzed by qRT-PCR. Endogenous hcaT of E. coli was used to normalize the data. The results are expressed as means ± SE of the means, n=3. Different alphabet above bars represent significant difference at p ≤ 0.05 according to Duncan’s multiple range test.
Table S1 List of primers used for qRT-PCR.
Table S2 Differentially expressed proteins identified by UHPLC-Q-TOF-IMS in E. coli upon Pa-SOD induction.
Table S3 Differentially expressed proteins identified by MALDI-TOF-MS/MS/MS in E. coli upon Pa-SOD induction.
Table S4 Functional classification and cluster analysis of differentially expressed protein obtained from PANTHER and DAVID, respectively.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with animals or human participants performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Guleria, S., Joshi, R., Singh, D. et al. Identification of host factors limiting the overexpression of recombinant Cu, Zn superoxide dismutase in Escherichia coli. Biotechnol Lett 42, 2389–2401 (2020). https://doi.org/10.1007/s10529-020-02962-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-02962-6