Abstract
Objectives
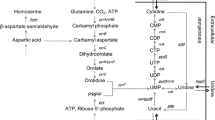
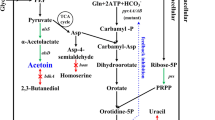
The metabolic pathway related to uridine production was modified in Bacillus subtilis in order to increase the production of uridine.
Results
Decreasing the relative transcriptional level of pur operon in Bacillus subtilis TD300 to 80%, and the production of the derived strain TD312 was increased to 11.81 g uridine/l and the yield was increased to 270 mg uridine/g glucose. The expression of pucR gene in situ by PccpA resulting in a 194.01-fold increase in the relative transcriptional level of pucR gene and 349.71-fold increase in the relative transcriptional level of ure operon, respectively. Furthermore, the production of TD314 reached 13.06 g uridine/l, while the yield reached 250 mg uridine/g glucose.
Conclusion
This is the first report that more than 13 g uridine/l with a yield of 250 mg uridine/g glucose is produced in shake flask fermentation of genetically engineered Bacillus subtilis.




Similar content being viewed by others
References
Beier L, Nygaard P, Jarmer H, Saxild HH (2002) Transcription analysis of the Bacillus subtilis PucR regulon and identification of a cis-acting sequence required for PucR-regulated expression of genes involved in purine catabolism. J Bacteriol 184:3232–3241. https://doi.org/10.1128/JB.184.12.3232-3241.2002
Belitsky BR, Sonenshein AL (2013) Genome-wide identification of Bacillus subtilis CodY-binding sites at single-nucleotide resolution. Proc Natl Acad Sci USA 110:7026–7031. https://doi.org/10.1073/pnas.1300428110
Bera AK, Zhu J, Zalkin H, Smith JL (2003) Functional dissection of the Bacillus subtilis pur operator site. J Bacteriol 185:4099–4109. https://doi.org/10.1128/JB.185.14.4099-4109.2003
Doi M, Asahi S, Tsunemi Y, Akiyama SI (1989) Mechanism of uridine production by Bacillus subtilis mutants. Appl Microbiol Biotechnol 30:234–238. https://doi.org/10.1007/BF00256210
Fan X, Wu H, Li G, Yuan H, Zhang H, Li Y, Xie X, Chen N (2017) Improvement of uridine production of Bacillus subtilis by atmospheric and room temperature plasmamutagenesis and high-throughput screening. PLoS ONE 12:e0176545. https://doi.org/10.1371/journal.pone.0176545
Fan X, Wu H, Li G, Jia Z, Li G, Li Q, Chen N, Xie X (2018) Metabolic engineering of Bacillus subtilis for the co-production of uridine and acetoin. PLoS ONE 12:e0176545. https://doi.org/10.1007/s00253-018-9316-7
Han AR, Kang HR, Son J, Kwon DH, Kim S, Lee WC et al (2016) The structure of the pleiotropic transcription regulator CodY provides insight into its GTP-sensing mechanism. Nucleic Acids Res 44:9483–9493. https://doi.org/10.1093/nar/gkw775
Hobl B, Mack M (2007) The regulator protein PyrR of Bacillus subtilis specifically interacts in vivo with three untranslated regions within pyr mRNA of pyrimidine biosynthesis. Microbiology 153:693–700. https://doi.org/10.1099/mic.0.2006/003772-0
Jordheim LP, Durantel D, Zoulim F, Dumontet C (2013) Advances in the development of nucleoside and nucleotide analogues for cancer and viral diseases. Nat Rev Drug Dis 12:447–464. https://doi.org/10.1038/nrd4010
Liu S, Endo K, Ara K, Ozaki K, Ogasawara N (2008) Introduction of marker-free deletions in Bacillus subtilis using the AraR repressor and the ara promoter. Microbiology 154:2562–2570. https://doi.org/10.1099/mic.0.2008/016881-0
Wang Y, Ma R, Liu L, He L, Ban R (2018) Improvement of uridine production in Bacillus subtilis by metabolic engineering. Biotech Lett 40:151–155. https://doi.org/10.1007/s10529-017-2453-8
Zhu H, Yang SM, Yuan ZM, Ban R (2015) Metabolic and genetic factors affecting the productivity of pyrimidine nucleoside in Bacillus subtilis. Microb Cell Fact 14:54. https://doi.org/10.1186/s12934-015-0237-1
Acknowledgements
This work was supported by the National High-tech R&D Program of China (Grant No. 2012AA02A701).
Supporting information
Supplementary Table 1—Strains used.
Supplementary Table 2—Primers used.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they have no conflicts of interest to this work.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, X., Wang, C., Liu, L. et al. Improve uridine production by modifying related metabolic pathways in Bacillus subtilis. Biotechnol Lett 42, 551–555 (2020). https://doi.org/10.1007/s10529-020-02820-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-02820-5