Abstract
Objective
To gain insights on the degree of heterogeneity and kinetic differences of streptokinase (SK) from group G (SKG) Streptococci compared with standard SK from group C (SKC) and identification of potentially contributing critical residues (hotspots).
Results
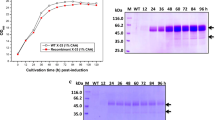
DNA and sequencing analyses confirmed the proper construction of all SK encoding vectors (two SKGs and one standard SKC). SDS-PAGE and western blot analyses confirmed the expression and proper purification of the recombinant SKs from E.coli with the expected size of 47 kDa. Kinetic analyses of two SKGs, compared with SKC, showed higher levels of specific [(×103 IU/mg) of 725 and 715 vs. 536] and fibrin-dependent proteolytic activities [Kcat/KM (min−1/µM) of 37 and 30 vs. 23], accompanied by declined fibrin-independent amidolytic activities [Kcat/KM (min−1/mM) of 109 and 84 vs. 113], respectively. Sequence alignments identified 10 novel residual substitutions scattered in SKα (I33F, R45Q, SKG132, A47D, and G55 N), SKβ (N228 K, F287I), and SKγ domains (L335 V, V396A, T403S) of SKGs, as potential hotspots.
Conclusion
The residue substitutions identified might critically contribute as hot spots to different kinetic parameters of SKGs and might assist in further elucidation of structure/function relations and rational design of SKs with improved (fibrin-dependent) therapeutic properties.

Similar content being viewed by others
References
Aneja R, Datt M, Singh B, Kumar S, Sahni G (2009) Identification of a new exosite involved in catalytic turnover by the streptokinase-plasmin activator complex during human plasminogen activation. J Biol Chem 284:32642–32650
Aneja R, Datt M, Yadav S, Sahni G (2013) Multiple exosites distributed across the three domains of streptokinase co-operate to generate high catalytic rates in the streptokinase-plasmin activator complex. Biochemistry 52(49):8957–8968
Arabi R, Arabi R, Roohvand F, Norouzian D, Sardari S et al (2011) A comparative study on the activity and antigenicity of truncated and full-length forms of streptokinase. Pol J Microbiol 60:243–251
Boxrud PD, Fay WP, Bock PE (2000) Streptokinase binds to human plasmin with high affinity, perturbs the plasmin active site, and induces expression of a substrate recognition exosite for plasminogen. J Biol Chem 275:14579–14589
Chaudhary A, Vasudha S, Rajagopal K et al (1999) Function of the central domain of streptokinase in substrate plasminogen docking and processing revealed by site directed mutagenesis. Protein Sci 8:2791–2805
Cook SM, Skora A, Gillen CM, Walker MJ, McArthur JD (2012) Streptokinase variants from Streptococcus pyogenes isolates display altered plasminogen activation characteristics–implications for pathogenesis. Mol Microbiol 86:1052–1062
Huang TT, Malke H, Ferretti J (1989) The streptokinase gene of group A streptococci: cloning, expression in Escherichia coli, and sequence analysis. Mol Microbiol 3:197–205
Kalia A, Bessen DE (2004) Natural selection and evolution of streptococcal virulence genes involved in tissue-specific adaptations. J Bacteriol 186:110
Keramati M, Roohvand F, Eslaminejad Z, Mirzaie A, Nikbin VS, Aslani MM (2012) PCR/RFLP-based allelic variants of streptokinase and their plasminogen activation potencies. FEMS Microbiol Lett 335:79–85
Keramati M, Mianroodi RA, Memarnejadian A, Mirzaie A, Sazvari S, Aslani MM, Roohvand F (2013a) Towards a superior streptokinase for fibrinolytic therapy of vascular thrombosis. Cardiovasc Hematol Agents Med Chem 11:218–229
Keramati M, Roohvand F, Aslani MM, Motevalli F, Memarnejadian A (2013b) Pitfalls in screening streptococci for retrieving superior streptokinase (SK) genes: no activity correlation for streptococcal culture supernatant and recombinant SK. J Ind Microbiol Biotechnol 40:151–158
Keramati M, Roohvand F, Aslani MM, Khatami S, Aghasadeghi M, Sadat M, Memarnejadian A, Motevalli F (2013c) Screening, cloning and expression of active streptokinase from an Iranian isolate of S. equisimilis Group C in E. coli. Iran J Basic Med Sci 16:620–627
Kim DM, Lee SJ, Kim IC, Kim ST, Byun SM (2000) Asp41-His48 region of streptokinase is important in binding to a substrate plasminogen. Thromb Res 99:93–98
McArthur JD, McKay FC, Ramachandran V, Shyam P et al (2008) Allelic variants of streptokinase from Streptococcus pyogenes display functional differences in plasminogen activation. FASEB J 22:3146–3153
Mundada LV, Prorok M, DeFord ME, Figuera M, Castellino FJ, Fay WP (2003) Structure-function analysis of the streptokinase amino terminus (residues 1–59). J Biol Chem 278:24421–24427
Tharp AC, Laha M, Panizzi P, Thompson MW, Fuentes-Prior P, Bock PE (2009) Plasminogen substrate recognition by the streptokinase-plasminogen catalytic complex is facilitated by Arg253, Lys256, and Lys257 in the streptokinase gamma-domain and Kringle 5 of the substrate. J Biol Chem 284:19511–19521
Wang X, Lin X, Loy JA, Tang J, Zhang XC (1998) Crystal structure of the catalytic domain of human plasmin complexed with streptokinase. Science 281:1662–1665
Wohl RC, Summaria L, Robbins K (1980) Kinetics of activation of human plasminogen by different activator species at pH 7.4 and 37 degrees C. J Biol Chem 255:2005–2013
Yadav S, Aneja R, Kumar P, Datt M, Sinha S, Sahni G (2011) Identification through combinatorial random and rational mutagenesis of a substrate-interacting exosite in the domain of streptokinase. J Biol Chem 286:6458–6469
Young KC, Shi GY, Wu DH, Chang LC, Chang BI, Ou CP, Wu HL (1998) Plasminogen activation by streptokinase via a unique mechanism. J Biol Chem 273:3110–3116
Zhang Y, Liang Z, Glinton K, Ploplis VA, Castellino FJ (2013) Functional differences between Streptococcus pyogenes cluster 1 and cluster 2b streptokinases are determined by their β-domains. FEBS Lett 587:1304–1309
Zhang Y, Mayfield JA, Ploplis VA, Castellino FJ (2014) The β-domain of cluster 2b streptokinase is a major determinant for the regulation of its plasminogen activation activity by cellular plasminogen receptors. Biochem Biophys Res Commun 444:595–598
Acknowledgements
This work was supported by Pasteur Institute of Iran in partial fulfilment of Ph.D thesis of M.K. in Pharmaceutical Biotechnology program.
Supporting information
Supplementary Table 1—Extracted kinetic data from literature for random/directed mutations and deletions studies on SKC compared to SKGs of the present study.
Supplementary Fig. 1—Schematic presentation of HPG activation mechanism by SK in amidolytic (fibrin-independent) and proteolytic (fibrin-dependent) pathways.
Supplementary Fig. 2—Construction of streptokinase expression plasmids.
Supplementary Figs. 3, 4 and 5—Time course of single stage HPG activation, amidolytic and proteolytic activities (by SKC, SKG88 and SKG132), respectively.
Supplementary Fig. 6—Complete amino acid sequence alignment for SKG88, SKG132 and SKC9542.
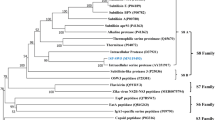
Supplementary Fig. 7—Construction of the phylogenetic tree, based on 339 bp hyper variable region of SKβ domain (sk-V1; residues 147–218).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Keramati, M., Aslani, M.M., Khatami, S. et al. Sequence and kinetic analyses of streptokinase from two group G streptococci with high fibrin-dependent plasminogen activities and the identification of novel altered amino acids as potential hot spots. Biotechnol Lett 39, 889–895 (2017). https://doi.org/10.1007/s10529-017-2310-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-017-2310-9