Abstract
Objective
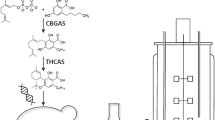
The Δ9-tetrahydrocannabinolic acid synthase (THCAS) from Cannabis sativa was expressed intracellularly in different organisms to investigate the potential of a biotechnological production of Δ9-tetrahydrocannabinolic acid (THCA) using whole cells.
Results
Functional expression of THCAS was obtained in Saccharomyces cerevisiae and Pichia (Komagataella) pastoris using a signal peptide from the vacuolar protease, proteinase A. No functional expression was achieved in Escherichia coli. The highest volumetric activities obtained were 98 pkat ml−1 (intracellular) and 44 pkat ml−1 (extracellular) after 192 h of cultivation at 15 °C using P. pastoris cells. Low solubility of CBGA prevents the THCAS application in aqueous cell-free systems, thus whole cells were used for a bioconversion of cannabigerolic acid (CBGA) to THCA. Finally, 1 mM (0.36 g THCA l−1) THCA could be produced by 10.5 gCDW l−1 before enzyme activity was lost.
Conclusion
Whole cells of P. pastoris offer the capability of synthesizing pharmaceutical THCA production




Similar content being viewed by others
References
Carlini EA (2004) The good and the bad effects of (-) trans-delta-9-tetrahydrocannabinol (Delta 9-THC) on humans. Toxicon 44:461–467
Delic M, Graf A, Koellensperger G et al (2014) Overexpression of the transcription factor Yap1 modifies intracellular redox conditions and enhances recombinant protein secretion. Microb Cell 1:376–386
Ernst O, Zor T (2010) Linearization of the bradford protein assay. J Vis Exp 38:1–6
Mechoulam R (1970) Marihuana Chemistry. Science 168:1159–1165
Pertwee RG (2006) Cannabinoid pharmacology: the first 66 years. Br J Pharmacol 147:S163–S171
Rothman J, Stevens TH (1986) Protein sorting in yeast: mutants defective in vacuole biogenesis mislocalize vacuolar proteins into the late secretory pathway 47:1041–1051
Shoyama Y, Tamada T, Kurihara K et al (2012) Structure and function of ∆1-tetrahydrocannabinolic acid (THCA) synthase, the enzyme controlling the psychoactivity of Cannabis sativa. J Mol Biol 423:96–105
Sirikantaramas S, Morimoto S, Shoyama Y et al (2004) The gene controlling marijuana psychoactivity: molecular cloning and heterologous expression of Delta1-tetrahydrocannabinolic acid synthase from Cannabis sativa L. J Biol Chem 279:39767–39774
Stehle F, Stubbs MT, Strack D, Milkowski C (2008) Heterologous expression of a serine carboxypeptidase-like acyltransferase and characterization of the kinetic mechanism. FEBS J 275:775–787
Taura F, Dono E, Sirikantaramas S et al (2007) Production of Delta(1)-tetrahydrocannabinolic acid by the biosynthetic enzyme secreted from transgenic Pichia pastoris. Biochem Biophys Res Commun 361:675–680
Tolner B, Smith L, Begent RHJ, Chester KA (2006) Production of recombinant protein in Pichia pastoris by fermentation. Nat Protoc 1:1006–1021
Trost BM, Dogra K (2007) Synthesis of (-)-Delta9-trans-tetrahydrocannabinol: stereocontrol via Mo-catalyzed asymmetric allylic alkylation reaction. Org Lett 9:861–863
Acknowledgments
This study was financially supported by the Graduate Cluster Industrial Biotechnology (CLIB). The authors are thankful to the thesis students for their excellent help during the laboratory work: Dirk Münker, David Dannheisig and Madeleine Dorsch. We are also grateful to Parijat Kusari for critically reading this manuscript. Studies were conducted with the permission of No. 4584989 issued by the Federal Institute for Drugs and Medical Devices (BfArM), Germany.
Supporting information
Supplementary Table 1: List of microorganisms used for expression of THCAS.
Supplementary Table 2: List of plasmids.
Supplementary Fig. 1: Screening of P. pastoris clones—volumetric THCAS activity; cultures were inoculated at 0.105 gCDW l−1. Cultures were grown at 200 rpm and 20 °C. Methanol was added every 24 h at 0.5 % (v/v). Values are calculated from biological duplicates.
Supplementary Fig. 2: Screening of P. pastoris clones—specific THCAS activity; cultures were inoculated at 0.105 gCDW l−1. Cultures were grown at 200 rpm and 20 °C. Methanol was added every 24 h at 0.5 % (v/v). Values are calculated from biological duplicates.
Supplementary Fig. 3: Expression of THCAS using PP2_HC; Cultures were grown in 3-baffled shake-flasks at 200 rpm and 10 °C. Methanol was added every 24 h at 0.5 % (v/v). Data points represent the means of three biological replicates with two technical replicates and error bars represent the standard deviation.
Supplementary Fig. 4: Expression of THCAS using PP2_HC; Cultures were grown in 3-baffled shake-flasks at 200 rpm and 20 °C. Methanol was added every 24 h at 0.5 % (v/v). Data points represent the means of three biological replicates with two technical replicates and error bars represent the standard deviation.
Supplementary Fig. 5: Expression of THCAS using PP2_HC; Cultures were grown in 3-baffled shake-flasks at 200 rpm and 25 °C. Methanol was added every 24 h at 0.5 % (v/v). Data points represent the means of three biological replicates with two technical replicates and error bars represent the standard deviation.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zirpel, B., Stehle, F. & Kayser, O. Production of Δ9-tetrahydrocannabinolic acid from cannabigerolic acid by whole cells of Pichia (Komagataella) pastoris expressing Δ9-tetrahydrocannabinolic acid synthase from Cannabis sativa l.. Biotechnol Lett 37, 1869–1875 (2015). https://doi.org/10.1007/s10529-015-1853-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-015-1853-x