Abstract
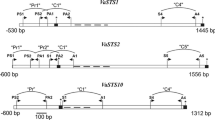
Resveratrol is a plant-derived phenol but the mechanism that regulates its biosynthesis remains unidentified. Stilbene synthase (STS) catalyzes resveratrol formation in vivo and we have proposed that inducers of resveratrol production affect STS expression through an unidentified epigenetic mechanism. To investigate the role of DNA methylation in resveratrol biosynthesis, we treated both rolB transgenic and empty vector control Vitis amurensis cell cultures with the DNA demethylation agent, 5-azacytidine. Treated cells had increased resveratrol production through activation of VaSTS10 expression. The lowest levels of cytosine methylation were at the 5′- and 3′-ends of the VaSTS1 protein-coding sequence. Cytosine methylation decreased mostly at the 5′- and 3′-ends of VaSTS10 after azaC treatment with an intriguing regularity in the number of cytosine nucleotides within the 5′- and 3′- ends of the protein-coding sequences. Thus, cytosine methylation is crucial for the regulation of the resveratrol biosynthetic pathway.



Similar content being viewed by others
References
Agius F, Kapoor A, Zhu JK (2006) Role of the Arabidopsis DNA glycosylase/lyase ROS1 in active DNA demethylation. Proc Natl Acad Sci USA 103:11796–11801
Alvarez ME, Nota F, Cambiagno DA (2010) Epigenetic control of plant immunity. Mol Plant Pathol 11:563–576
Aziz MH, Kumar R, Ahmad N (2003) Cancer chemoprevention by resveratrol: in vitro and in vivo studies and the underlying mechanisms. Int J Oncol 23:17–28
Bird A (2002) DNA methylation patterns and epigenetic memory. Gene Dev 16:6–21
Chandra S (2012) Natural plant genetic engineer Agrobacterium rhizogenes: role of T-DNA in plant secondary metabolism. Biotechnol Lett 3:407–415
Dao TTH, Linthorst HJM, Verpoorte R (2011) Chalcone synthase and its functions in plant resistance. Phytochem Rev 10:397–412
Finnegan EJ, Genger RK, Peacock WJ, Dennis ES (1998) DNA methylation in plants. Annu Rev Plant Physiol Plant Mol Biol 49:223–247
Frankel EN, Waterhouse AL (1993) Inhibition of human LDL oxidation by resveratrol. Lancet 341:1103–1104
Hudson K, Luo S, Hagemann N, Preuss D (2011) Changes in global gene expression in response to chemical and genetic perturbation of chromatin structure. PLoS ONE 6:e20587
Kawiak A, Krolicka A, Lojkowska E (2011) In vitro cultures of Drosera aliciae as a source of a cytotoxic naphthoquinone: ramentaceone. Biotechnol Lett 33:2309–2316
Kiselev KV (2011) Perspectives for production and application of resveratrol. Appl Microbiol Biotechnol 90:417–425
Kiselev KV, Dubrovina AS (2010) A new method for analysing gene expression based on frequency analysis of RT-PCR products obtained with degenerate primers. Acta Physiol Plant 32:495–502
Kiselev KV, Dubrovina AS, Veselova MV, Bulgakov VP, Fedoreyev SA, Zhuravlev YN (2007) The rolB gene-induced overproduction of resveratrol in Vitis amurensis transformed cells. J Biotechnol 128:681–692
Kiselev KV, Dubrovina AS, Bulgakov VP (2009) Phenylalanine ammonia–lyase and stilbene synthase gene expression in rolB transgenic cell cultures of Vitis amurensis. Appl Microbiol Biotechnol 82:647–655
Kiselev KV, Tyunin AP, Manyakhin AY, Zhuravlev YN (2011) Resveratrol content and expression patterns of stilbene synthase genes in Vitis amurensis cells treated with 5-azacytidine. Plant Cell Tissue Organ Cult 105:65–72
Kiselev KV, Tyunin AP, Zhuravlev YN (2012) Involvement of DNA methylation in the regulation of STS10 gene expression in Vitis amurensis. Planta. doi:10.1007/s00425-012-1806-8
Ku KL, Chang PS, Cheng YC, Lien CY (2005) Production of stilbenoids from the callus of Arachis hypogaea: a novel source of the anticancer compound piceatannol. J Agric Food Chem 53:3877–3881
Langcake P, Pryce RJ (1977) A new class of phytoalexins from grapevines. Experientia 33:151–152
Law JA, Jacobsen SE (2010) Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet 11:204–220
Lister R, O’Malley RC, Tonti-Filippini J, Gregory BD, Berry CC, Millar AH, Ecker JR (2008) Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell 3:523–536
Messeguer R, Ganal MW, Steffens JC, Tanksley SD (1991) Characterization of the level, target sites and inheritance of cytosine methylation in tomato nuclear DNA. Plant Mol Biol 16:753–770
Parage C, Tavares R, Rety S, Baltenweck-Guyot R, Poutaraud A, Renault L, Heintz D, Lugan R, Marais GAB, Aubourg S, Hugueney P (2012) Structural, functional, and evolutionary analysis of the unusually large stilbene synthase gene family in grapevine. Plant Physiol 160:1407–1419
Pavet V, Quintero C, Cecchini NM, Rosa AL, Alvarez ME (2006) Arabidopsis displays centromeric DNA hypomethylation and cytological alterations of heterochromatin upon attack by Pseudomonas syringae. Mol Plant-Microbe Interact 19:577–587
Pecinka A, Dinh HQ, Baubec T, Rosa M, Lettner N, Mittelsten-Scheid O (2010) Epigenetic regulation of repetitive elements is attenuated by prolonged heat stress in Arabidopsis. Plant Cell 9:3118–3129
Penterman J, Zilberman D, Huh JH, Ballinger T, Henikoff S, Fischer RL (2007) DNA demethylation in the Arabidopsis genome. Proc Natl Acad Sci USA 104:6752–6757
Shankar S, Singh G, Srivastava RK (2007) Chemoprevention by resveratrol: molecular mechanisms and therapeutic potential. Front Biosci 12:4839–4854
Shumakova OA, Manyakhin AY, Kiselev KV (2011) Resveratrol content and expression of phenylalanine ammonia-lyase and stilbene synthase genes in cell cultures of Vitis amurensis treated with coumaric acid. Appl Biochem Biotechnol 165:1427–1436
Spena A, Schmulling T, Koncz C, Schell JS (1987) Independent and synergistic activity of rol A, B and C loci in stimulating abnormal growth in plants. EMBO J 6:3891–3899
Tassoni A, Fornale S, Franceschetti M, Musiani F, Michael AJ, Perry B, Bagni N (2005) Jasmonates and Na-orthovanadate promote resveratrol production in Vitis vinifera cv. Barbera cell cultures. New Phytol 166:895–905
Tyunin AP, Kiselev KV, Zhuravlev YN (2012) Effects of total DNA demethylation on methyltransferase gene expression and resveratrol production in cell cultures of Vitis amurensis. Plant Cell Tissue Organ Cult 111:91–100
Vanyushin BF, Ashapkin VV (2011) DNA methylation in higher plants: past, present and future. Biochim Biophys Acta 8:360–368
Velasco R, Zharkikh A, Troggio M, Cartwright DA, Cestaro A, Pruss D, Pindo M, Fitzgerald LM, Vezzulli S (2007) A high quality draft consensus sequence of the genome of a heterozygous grapevine variety. PLoS ONE 2:e1326
Ya-Ut P, Chareonsap P, Sukrong S (2011) Micropropagation and hairy root culture of Ophiorrhiza alata Craib for camptothecin production. Biotechnol Lett 33:2519–2526
Zhu JK (2009) Active DNA demethylation mediated by DNA glycosylases. Annu Rev Genet 43:143–166
Acknowledgments
The authors express their thanks to Korben Dallas for inspiring and helpful comments on the manuscript. This work was supported by Grant of the Russian Foundation for Basic Research (12-04-33069-mol_ved), by grants of the Far East Division of the Russian Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tyunin, A.P., Kiselev, K.V. & Karetin, Y.A. Differences in the methylation patterns of the VaSTS1 and VaSTS10 genes of Vitis amurensis Rupr.. Biotechnol Lett 35, 1525–1532 (2013). https://doi.org/10.1007/s10529-013-1235-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-013-1235-1