Abstract
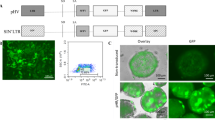
293T and Sk-Hep-1 cells were transduced with a replication-defective self-inactivating HIV-1 derived vector carrying FVIII cDNA. The genomic DNA was sequenced to reveal LTR/human genome junctions and integration sites. One hundred and thirty-two sequences matched human sequences, with an identity of at least 98%. The integration sites in 293T-FVIIIDB and in Sk-Hep-FVIIIDB cells were preferentially located in gene regions. The integrations in both cell lines were distant from the CpG islands and from the transcription start sites. A comparison between the two cell lines showed that the lentiviral-transduced DNA had the same preferred regions in the two different cell lines.



Similar content being viewed by others
References
Antequera F, Bird A (1999) CpG islands as genomic footprints of promoters that are associated with replication origins. Curr Biol 9:R661–R667
Beard BC, Dickerson D, Beebe K, Gooch C, Fletcher J, Okbinoglu T, Miller DG, Jacobs MA, Kaul R, Kiem HP, Trobridge GD (2007) Comparison of HIV-derived lentiviral and MLV-based gammaretroviral vector integration sites in primate repopulating cells. Mol Ther 15:1356–1365
Cavazzana-Calvo M, Hacein-Bey S, de Saint Basile G, Gross F, Yvon E, Nusbaum P, Selz F, Hue C, Certain S, Casanova JL, Bousso P, Deist FL, Fischer A (2000) Gene therapy of human severe combined immunodeficiency (SCID)-X1 disease. Science 288:669–672
Hacein-Bey-Abina S, von Kalle C, Schmidt M, Le Deist F, Wulffraat N, McIntyre E, Radford I, Villeval JL, Fraser CC, Cavazzana-Calvo M, Fischer A (2003a) A serious adverse event after successful gene therapy for X-linked severe combined immunodeficiency. N Engl J Med 348:255–256
Hacein-Bey-Abina S, Von Kalle C, Schmidt M, McCormack MP, Wulffraat N et al (2003b) LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1. Science 302:415–419
Hacker CV, Vink CA, Wardell TW, Lee S, Treasure P, Kingsman SM, Mitrophanous KA, Miskin JE (2006) The integration profile of EIAV-based vectors. Mol Ther 14:536–545
Kim S, Kim N, Dong B, Boren D, Lee SA, Das Gupta J, Gaughan C, Klein EA, Lee C, Silverman RH, Chow SA (2008) Integration site preference of xenotropic murine leukemia virus-related virus, a new human retrovirus associated with prostate cancer. J Virol 82:9964–9977
Kustikova O, Fehse B, Modlich U, Yang M, Dullmann J, Kamino K, von Neuhoff N, Schlegelberger B, Li Z, Baum C (2005) Clonal dominance of hematopoietic stem cells triggered by retroviral gene marking. Science 308:1171–1174
Laufs S, Guenechea G, Gonzalez-Murillo A, Zsuzsanna Nagy K, Luz Lozano M, del Val C, Jonnakuty S, Hotz-Wagenblatt A, Jens Zeller W, Bueren JA, Fruehauf S (2006) Lentiviral vector integration sites in human NOD/SCID repopulating cells. J Gene Med 8:1197–1207
Lavie L, Maldener E, Brouha B, Meese EU, Mayer J (2004) The human L1 promoter: variable transcription initiation sites and a major impact of upstream flanking sequence on promoter activity. Genome Res 14:2253–2260
Lewinski MK, Yamashita M, Emerman M, Ciuffi A, Marshall H, Crawford G, Collins F, Shinn P, Leipzig J, Hannenhalli S, Berry CC, Ecker JR, Bushman FD (2006) Retroviral DNA integration: viral and cellular determinants of target-site selection. PLoS Pathog 2:e60
Li Z, Dullmann J, Schiedlmeier B, Schmidt M, von Kalle C, Meyer J, Forster M, Stocking C, Wahlers A, Frank O, Ostertag W, Kuhlcke K, Eckert HG, Fehse B, Baum C (2002) Murine leukemia induced by retroviral gene marking. Science 296:497
Mitchell RS, Beitzel BF, Schroder AR, Shinn P, Chen H, Berry CC, Ecker JR, Bushman FD (2004) Retroviral DNA integration: ASLV, HIV, and MLV show distinct target site preferences. PLoS Biol 2:E234
Montini E, Cesana D, Schmidt M, Sanvito F, Ponzoni M, Bartholomae C, Sergi Sergi L, Benedicenti F, Ambrosi A, Di Serio C, Doglioni C, von Kalle C, Naldini L (2006) Hematopoietic stem cell gene transfer in a tumor-prone mouse model uncovers low genotoxicity of lentiviral vector integration. Nat Biotechnol 24:687–696
Picanco V, Heinz S, Bott D, Behrmann M, Covas DT, Seifried E, Tonn T (2007) Recombinant expression of coagulation factor VIII in hepatic and non-hepatic cell lines stably transduced with third generation lentiviral vectors comprising the minimal factor VIII promoter. Cytotherapy 9:785–794
Picanço-Castro V, Fontes AM, Russo-Carbolante EMS, Covas DT (2008) Lentiviral-mediated gene transfer—a patent review. Exp Opin Therap Patents 18(5):525–539
Rohdewohld H, Weiher H, Reik W, Jaenisch R, Breindl M (1987) Retrovirus integration and chromatin structure: Moloney murine leukemia proviral integration sites map near DNase I-hypersensitive sites. J Virol 61:336–343
Tonn T, Herder C, Becker S, Seifried E, Grez M (2002) Generation and characterization of human hematopoietic cell lines expressing factor VIII. J Hematother Stem Cell Res 11:695–704
Venter JC, Adams MD, Meyers E, Li PW, Mural RJ et al. (2001) The Sequence of the Human Genome. Science 1304–1351
Wu X, Li Y, Crise B, Burgess SM (2003) Transcription start regions in the human genome are favored targets for MLV integration. Science 300:1749–1751
Acknowledgments
We thank Adriana Aparecida Marques who helped with the sequencing. This work was supported by CNPq (Conselho Nacional de Desenvolvimento Tecnológico) and FAPESP (Fundação de Amparo a Pesquisa do Estado de São Paulo).
Author information
Authors and Affiliations
Corresponding author
Additional information
Purpose of work We have examined lentiviral integration site selection in hepatocyte and kidney cell lines to see whether there are determinants of integration tendency other than the structure and biological characteristics of lentiviruses.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
de Sousa Russo-Carbolante, E.M., Picanço-Castro, V., Alves, D.C.C. et al. Integration pattern of HIV-1 based lentiviral vector carrying recombinant coagulation factor VIII in Sk-Hep and 293T cells. Biotechnol Lett 33, 23–31 (2011). https://doi.org/10.1007/s10529-010-0387-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-010-0387-5