Abstract
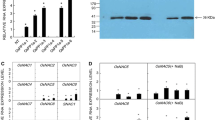
A phosphate starvation-induced, purple, acid phosphatase cDNA was cloned from rice, Oryza sativa. The cDNA encoding the phosphatase (OsPAP2) has 1,893 bp with an open reading frame of 630 amino acid residues. The deduced amino acid sequence of OsPAP2 shows identities of 60–63% with other plant purple acid phosphatases and appears to have five conserved motifs containing the residues involved in metal binding. OsPAP2 expression is up-regulated in the rice plant and in cell cultures in the absence of phosphate (P i ). The induced expression of OsPAP2 is a specific response to P i starvation, and is not affected by the deprivation of other nutrients. OsPAP2 expression was responsive to the level of P i -supply, and transcripts of OsPAP2 were abundant in P i -deprived roots. The OsPAP2 cDNA was expressed as a 69 kDa polypeptide in baculovirus-infected insect Sf9 cells. In addition, the OsPAP2 gene was introduced into Arabidopsis via an Agrobacterium-mediated transformation. Functional expression of the OsPAP2 gene in the transgenic Arabidopsis line was confirmed by northern and western blot analyses, as well as by phosphatase activity assays. These results suggest that the OsPAP2 gene can be used to develop new transgenic dicotyledonous plants that are able to adapt to P i -deficient conditions.




Similar content being viewed by others
References
Baldwin JC, Karthikeyan AS, Raghothama KG (2001) LEPS2, a phosphorus starvation-induced novel acid phosphatase from tomato. Plant Physiol 125:728–737
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
del Pozo JC, Allona I, Rubio V, Leyva A, de la Pena A, Aragoncillo C, Paz-Ares J (1999) A type 5 acid phosphatase gene from Arabidopsis thaliana is induced by phosphate starvation and by some other types of phosphate mobilising/oxidative stress conditions. Plant J 19:579–589
Goldstein AH, Baertlein DA, McDaniel RG (1988) Phosphate starvation inducible metabolism in lycopersicon esculentum I. Excretion of acid phosphatase by tomato plants and suspension-cultured cells. Plant Physiol 87:711–715
Hegeman CE, Grabau EA (2001) A novel phytase with similarity to purple acid phosphatases is expressed in cotyledons of germinating soybean seedlings. Plant Physiol 126:1598–1608
Hur YJ, Lee HG, Jeon EJ, Lee YY, Nam MH, Yi G, Eun MY, Nam J, Lee JH, Kim DH (2007) A phosphate starvation-induced acid phosphatase from Oryza sativa: phosphate regulation and transgenic expression. Biotechnol Lett 29:829–835
Je YH, Chang JH, Choi JY, Roh JY, Jin BR, O’Reilly DR, Kang SK (2001) A defective viral genome maintained in Escherichia coli for the generation of baculovirus expression vectors. Biotechnol Lett 23:575–582
Klabunde T, Strater N, Frohlich R, Witzel H, Krebs B (1996) Mechanism of FeIII-ZnII purple acid phosphatase based on crystal structures. J Mol Biol 259:737–748
Li D, Zhu H, Liu K, Liu X, Leggewie G, Udvardi M, Wang D (2002) Purple acid phosphatases of Arabidopsis thaliana. J Biol Chem 277:27772–27781
Miller SS, Liu J, Allan DL, Menzhuber CJ, Fedorova M, Vance CP (2001) Molecular control of acid phosphatase secretion into the rhizosphere of proteoid roots from phosphorus-stressed white lupin. Plant Physiol 127:594–606
O’Reilly DR, Miller LK, Luckow VA (1992) Baculovirus expression vectors: a laboratory manual. W H. Freeman & Co., New York
Schenk G, Guddat LW, Carrington LE, Hume DA, Hamilton S, de Jersey J (2000) Identification of mammalian-like purple acid phosphatases in a wide range of plants. Gene 250:117–125
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Yu SM, Kuo YH, Sheu G, Sheu YJ, Liu LF (1991) Metabolic derepression of alpha-amylase gene expression in suspension-cultured cells of rice. J Biol Chem 266:21131–21137
Zimmermann P, Regrierer B, Kossmann J, Frossard E, Amrhein N, Bucher M (2004) Differential expression of three purple acid phosphatases from potato. Plant Biol 6:519–528
Acknowledgments
This work was supported by a grant from the Crop Functional Genomics Center of the 21st Century Frontier Research Program (No. CG 3123) and the Biogreen 21 programs, Rural Development Administration (RDA), Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hur, Y.J., Jin, B.R., Nam, J. et al. Molecular characterization of OsPAP2: transgenic expression of a purple acid phosphatase up-regulated in phosphate-deprived rice suspension cells. Biotechnol Lett 32, 163–170 (2010). https://doi.org/10.1007/s10529-009-0131-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-009-0131-1