Abstract
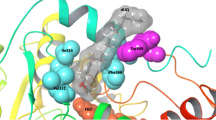
Comparison of three-dimensional structures of flavin-dependent azoreductases revealed two conserved loops around the flavin mononucleotide (FMN) cofactor. Tyr74, His75 and Lys109 in the two loops of azoreductase AZR from Rhodobacter sphaeroides were replaced with Trp, Asn and Ala/His by site-directed mutagenesis, respectively. The optimal pH values of K109H and H75N were pH 6, and those of K109A and Y74W were pH 9. The optimal temperature (30°C) was not affected by mutation. Positively charged residues at position 109 is critical for the binding of methyl red. K109 might only be involved in the binding of the 2′-phosphate group of NADPH and have no effect on the binding of NADH. Y74W and H75N mutations decreased the binding of methyl red/nitrofurazone and had no affect on the binding of NADPH.




Similar content being viewed by others
References
Bellamacina CR (1996) The nicotinamide dinucleotide binding motif: a comparison of nucleotide binding proteins. FASEB J 10:1257–1269
Bottoms CA, Smith PE, Tanner JJ (2002) A structurally conserved water molecule in Rossmann fold dinucleotide-binding domains. Protein Sci 11:2125–2137
Chen H (2006) Recent advances in azo dye degrading enzyme research. Curr Protein Pept Sci 7:101–111
Chen H, Hopper SL, Cerniglia CE (2005) Biochemical and molecular characterization of an azoreductase from Staphylococcus aureus, a tetrameric NADPH-dependent flavoprotein. Microbiology 151:1433–1441
Chung KT (1983) The significance of azo-reducion in the mutagenesis and carcinogenesis of azo dyes. Mutat Res 114:269–281
DeLano WL (2003) PyMOL reference manual. DeLano Scientific LLC, San Carlos
Dos Santos AB, Cervantes FJ, Van Lier JB (2007) Review paper on current technologies for decolourisation of textile wastewaters: perspectives for anaerobic biotechnology. Biores Technol 98:2369–2385
Freigang J, Diederichs K, Schäfer KP, Welte W, Paul R (2002) Crystal structure of oxidized flavodoxin, an essential protein in Helicobacter pylori. Protein Sci 11:253–261
Hanauer SB (1996) Inflammatory bowel disease. N Engl J Med 334:841–848
Ito K, Nakanishi M, Lee WC, Sasaki H, Zenno S, Saigo K, Kitade Y, Tanokura M (2006) Three-dimensional structure of AzoR from Escherichia coli. An oxidereductase conserved in microorganisms. J Biol Chem 281:20567–20576
Jornvall H, Persson B, Krook M, Atrian S, Gonzalez-Duarte R, Jeffery J, Ghosh D (1995) Short-chain dehydrogenases/reductases (SDR). Biochemistry 34:6003–6013
Kobori T, Sasaki H, Lee WC, Zenno S, Saigo K, Murphy ME, Tanokura M (2001) Structure and site-directed mutagenesis of a flavoprotein from Escherichia coli that reduces nitrocompounds: alteration of pyridine nucleotide binding by a single amino acid substitution. J Biol Chem 276:2816–2823
Liger D, Graille M, Zhou CZ, Leulliot N, Quevillon-Cheruel S, Blondeau K, Janin J, van Tilbeurgh H (2004) Crystal structure and functional characterization of yeast YLR011wp, an enzyme with NAD(P)H-FMN and ferric iron reductase activities. J Biol Chem 279:34890–34897
Liu G, Zhou J, Lv H, Xiang X, Wang J, Zhou M, Qv Y (2007a) Azoreductase from Rhodobacter sphaeroides AS1.1737 is a flavodoxin that also functions as nitroreductase and flavin mononucleotide reductase. Appl Microbiol Biotechnol 76:1271–1279
Liu ZJ, Chen H, Shaw N, Hopper SL, Chen L, Chen S, Cerniglia CE, Wang BC (2007b) Crystal structure of an aerobic FMN-dependent azoreductase (AzoA) from Enterococcus faecalis. Arch Biochem Biophys 463:68–77
Nakanishi M, Yatome C, Ishida N, Kitade Y (2001) Putative ACP phosphodiesterase gene (acpD) encodes an azoreductase. J Biol Chem 276:46394–46399
Nicholas KB, Nicholas HB Jr, Deerfield DW II (1997) GeneDoc: analysis and visualization of genetic variation. EMBnet News 4:1–4
Raffi F, Hall JD, Cerniglia CE (1997) Mutagenicity of azo dyes used in foods, drugs and cosmetics before and after reduction by Clostridium species from the human intestinal tract. Food Chem Toxicol 35:897–901
Reyes VM, Sawaya MR, Brown KA, Kraut J (1995) Isomorphous crystal structures of Escherichia coli dihydrofolate reductase complexed with folate, 5-deazafolate, and 5,10-dideazatetrahydrofolate: mechanistic implications. Biochemistry 34:2710–2723
Shore J (1996) Advances in direct dyes. Ind J Fibre Text Res 21:1–29
Stolz A (2001) Basic and applied aspects in the microbial degradation of azo dyes. Appl Microbiol Biotechnol 56:69–80
Suzuki Y, Yoda T, Ruhul A, Sugiura W (2001) Molecular cloning and characterization of the gene coding for azoreductase from Bacillus sp. OY1-2 isolated from soil. J Biol Chem 276:9059–9065
Wang CJ, Hagemeier C, Rahman N, Lowe E, Noble M, Coughtrie M, Sim E, Westwood I (2007) Molecular cloning, characterization and ligand-bound structure of an azoreductase from Pseudomonas aeruginosa. J Mol Biol 373:1213–1228
Wilson DK, Bohren KM, Gabbay KH, Quiocho FA (1992) An unlikely sugar substrate site in the 1.65 Å structure of the human aldose reductase holoenzyme implicated in diabetic complications. Science 257:81–84
Yan B, Zhou J, Wang J, Du C, Hou H, Song Z, Bao Y (2004) Expression and characteristics of the gene encoding azoreductase from Rhodobacter sphaeroides AS1.1737. FEMS Microbiol Lett 236:129–136
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, G., Zhou, J., Wang, J. et al. Site-directed mutagenesis of substrate binding sites of azoreductase from Rhodobacter sphaeroides . Biotechnol Lett 30, 869–875 (2008). https://doi.org/10.1007/s10529-007-9627-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-007-9627-8