Abstract
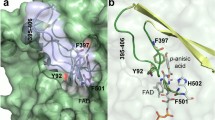
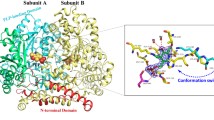
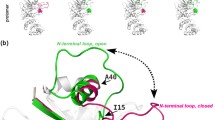
A three-dimensional structural model of fructosyl amine oxidase from the marine yeast Pichia N1-1 was generated using the crystal structure of monomeric sarcosine oxidase from Bacillus sp. B-0618 as template. The putative active site region was investigated by site-directed mutagenesis, identifying several amino acid residues likely playing important roles in the enzyme reaction. Asn354 was identified as a residue that plays an important role in substrate recognition and that can be substituted in order to change substrate specificity while maintaining high catalytic activity. While the Asn354Ala substitution had no effect on the V max K −1m value for fructosyl valine, the V max K −1m value for fructosyl-ε N-lysine was decreased 3-fold, thus resulting in a 3-fold improvement in specificity for fructosyl valine over fructosyl-ε N-lysine.


Similar content being viewed by others
References
Ferri S, Miura S, Sakaguchi A, Ishimura F, Tsugawa W, Sode K (2004) Cloning and expression of fructosyl-amine oxidase from marine yeast Pichia sp. N1-1. Mar Biotechnol 6:625–632
Ferri S, Sakaguchi A, Goto H, Tsugawa W, Sode K (2005) Isolation and characterization of fructosyl-amine oxidase from an Arthrobacter sp. Biotechnol Lett 27:27–32
Horiuchi T, Kurokawa T, Saito N (1989) Purification and properties of fructosyl amino acid oxidase from Corynebacterium sp. 2-4-1. Agric Biol Chem 53:103–110
Keil P, Mortensen HB, Christophersen C (1985) Fructosylvaline. A simple model of the N-terminal residue of human haemoglobin A1c. Acta Chem Scand B 39:191–193
Ogawa K, Stollner D, Scheller F, Warsinke A, Ishimura F, Tsugawa W, Ferri S, Sode K (2002) Development of a flow-injection analysis (FIA) enzyme sensor for fructosyl amine monitoring. Anal Bioanal Chem 373:211–214
Sode K, Ishimura F, Tsugawa W (2001) Screening and characterization of fructosyl-valine-utilizing marine microorganisms. Mar Biotechnol 3:126–132
Trickey P, Wagner MA, Jorns MS, Mathews FS (1999) Monomeric sarcosine oxidase: structure of covalently flavinylated amine oxidizing enzyme. Structure 7: 331–345
Tsugawa W, Ishimura F, Ogawa K, Sode K (2000) Development of an enzyme sensor utilizing a novel fructosyl amine oxidase from a marine yeast. Electrochemistry 68:869–871
Tsugawa W, Ogawa K, Ishimura F, Sode K (2001) Fructosyl amine sensing based on prussian blue modified enzyme electrode. Electrochemistry 69:973–975
Wierenga RK, Terpstra P, Hol WG (1986) Prediction of the occurrence of the ADP-binding beta alpha beta-fold in proteins, using an amino acid sequence fingerprint. J Mol Biol 187:101–107
Wu X, Takahashi M, Chen SG, Monnier VM (2000) Cloning of Amadoriase I isoenzyme from Aspergillus sp.: evidence of FAD Covalently Linked to Cys342. Biochemistry 39:1515–1521
Yoshida N, Sakai Y, Serata M, Tani Y, Kato N (1995) Distribution and properties of fructosyl amino acid oxidase in fungi. Appl Environ Microbiol 61:4487–4489
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Miura, S., Ferri, S., Tsugawa, W. et al. Active site analysis of fructosyl amine oxidase using homology modeling and site-directed mutagenesis. Biotechnol Lett 28, 1895–1900 (2006). https://doi.org/10.1007/s10529-006-9173-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-006-9173-9