Abstract
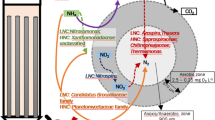
A glass bead biofilm reactor was operated using H2 as an electron donor to remove nitrate at 150 mg NO3–N l−1 to below detection level. The microbial community in the glass beads biofilm reactor was investigated by using denaturing gradient gel electrophoresis (DGGE) and phylogenetic analysis. In DGGE analysis of the biofilm, five bands were dominant and indicated the presence of eight β-proteobacteria, one γ-proteobacteria and twelve clostridia. An unculturable Hydrogenophaga sp., which is a new genus of hydrogen-oxidizing bacterium was dominant in microbial community of the biofilm reactor.
Similar content being viewed by others
References
RI Amann W Ludwig KK Schleifer (1995) ArticleTitlePhylogenetic identification and in situ detection of individual microbial cells without cultivation Microbiol. Rev 59 143–169 Occurrence Handle7535888
InstitutionalAuthorNameAPHA, AWWA, WPCF (1995) Standard Methods for the Examination of Water and Wastewater EditionNumber19 American Public Health Association, American Water Works Association and Water Pollution Control Federation New York, NY
F Azam DC Smith CF Steward A Hagstrom (1993) ArticleTitleBactriea-organic matter coupling and its significance for ocean carbon cycling Microbiol. Ecol 28 167–179 Occurrence Handle10.1007/BF00166806
AT Brain JW Lenly JJ Alvarez (1998) ArticleTitleFe(0)-supported autotrophic denitrification Environ. Sci. Technol 32 634–639 Occurrence Handle10.1021/es9707769
TC Driscoll JJ Bisogni (1978) ArticleTitleThe use of sulfur and sulfide in packed bed reactors for autotrophic denitrification J. Water Poll. Control Fedn 3 569–577
DC Kross GR Hallberg DR Bruner K Cherryholmes JK Johnson (1993) ArticleTitleThe nitrate contamination of private well water in Iowa Am. J. Public Health 83 270–272 Occurrence Handle8427340
LA Lawrence PL McCarty (1970) ArticleTitleUnified basis for biological treatment design and operation J. San. Eng. (American Society of Civil Engineers) 6 IssueIDSA3 757
G Muyzer EC Waal Particlede AG Uitterlinden (1993) ArticleTitleProfiling of complex microbial population by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA Appl. Environ. Microbiol 59 659–700
HI Park JS Kim DK Kim DW Pak (2004) ArticleTitleAutohydrogenotrophic denitrification of high nitrate concentration in a glass beads biofilm reactor J. Kor. Soc. Water Quality 20 236–240
BE Rittmann V Snoeyink (1984) ArticleTitleAchieving biologically stable drinking water J. Am. Water Works Assoc 76 106–114
V Urbain R Benoid J Manem (1996) ArticleTitleMembrane bioreactor: a new treatment tool J. Am. Water Works Assoc 88 75–86
A Willems J Busse M Goor B Pot (1989) ArticleTitleHydrogenophaga, a new genus of hydrogen-oxidizing bacteria that includes Hydrogenophaga flava comb. nov. (formerly Pseudomonas flava), Hydrogenophaga palleronii (formerly Pseudomonas palleronii), Hydrogenophaga pseudoflava (formerly Pseudomonas pseudoflava and Pseudomonas carboxydoflava) and Hydrogenophaga taeniospiralis (formerly Pseudomonas taeniospiralis) Int. J. Syst. Bacteriol 39 319–333
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Park, H.I., Choi, YJ. & Pak, D. Autohydrogenotrophic Denitrifying Microbial Community in a Glass Beads Biofilm Reactor. Biotechnol Lett 27, 949–953 (2005). https://doi.org/10.1007/s10529-005-7654-x
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s10529-005-7654-x