Abstract
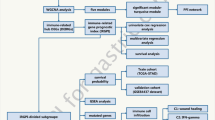
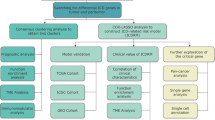
Immunogenic cell death (ICD) induces anti-tumor immunity and aids in dismantling the immunosuppressive immune microenvironment (TME), which belongs to a type of regulated cell death. The differentiation of gastric cancer (GC) subtypes and the discovery of prognostic biomarkers are crucial for its treatment because GC is a disease that is both highly heterogeneous and aggressive. However, although the induction of ICD in tumor cells is associated with a favorable prognosis, the exact mechanism of its role in GC remains unclear. Transcriptome profiling data and clinical data of GC patients were retrieved from The Cancer Genome Atlas (TCGA) database. Herein, patients were classified with the consensus clustering algorithm, and the associated biological functions and immune microenvironment infiltration were explored based on the expression of ICD-associated genes. A risk score signature consisting of 11 ICD-related genes was established via the least absolute shrinkage and selection operator regression (LASSO) method. We have retrieved similar studies in recent years and compared them with our study using the time-dependent receiver operating characteristic (ROC) curves. Gene set variation analysis (GSVA) and single sample gene set enrichment analysis (ssGSEA) were performed to explore the association between the signature and tumor microenvironment (TME). Two distinct subtypes associated with ICD in GC were identified, each with a different prognosis. The ICD-high expression subtype was associated with higher immune cell infiltration and a better prognosis. The ICD-related gene signature containing 11 genes (CGB5, Z84468.1, APOA5, EPHA8, CLEC18C, TLR7, MUC7, MUC15, CTLA4, CALB2, and UGT2B28), could independently and accurately predict the prognosis of GC. In this study, an ICD-based classification was conducted to assist in the diagnosis and personalized therapy for GC. The ICD-related genes risk score model was established to predict prognosis.








Similar content being viewed by others
Data Availability
All data generated and described in this paper are available from the corresponding web servers. Further inquiries can be directed to the corresponding authors.
Abbreviations
- GC:
-
Gastric cancer
- ICD:
-
Immunogenic cell death
- IRGs:
-
ICD-related genes
- TCGA:
-
The Cancer Genome Atlas
- GEO:
-
The Gene Expression Omnibus
- LASSO:
-
Least absolute shrinkage and selection operator regression
- TME:
-
Tumor microenvironment
- CDF:
-
Cumulative distribution function
- PCA:
-
Principal component analysis
- OS:
-
Overall survival
- ssGSEA:
-
Single sample gene set enrichment analysis
- ICGs:
-
Immune checkpoint genes
- ICB:
-
Immune checkpoint blockade
- TIDE:
-
The Tumor Immune Dysfunction and Exclusion
- IPS:
-
The immunophenoscore
- ICIs:
-
Immune checkpoint inhibitors
- DEGs:
-
Differentially expressed genes
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- ROC:
-
Receiver operating characteristic
- AUC:
-
Areas under the curve
- C-index:
-
The concordance index
- TMB:
-
Tumor mutation burden
- IC50:
-
The semi-inhibitory concentration
- GSVA:
-
Gene Set Enrichment Analysis
- TLRs:
-
Toll-like receptors
References
Adkins I, Fucikova J, Garg AD, Agostinis P, Špíšek R (2014) Physical modalities inducing immunogenic tumor cell death for cancer immunotherapy. Oncoimmunology 3(12):e968434. https://doi.org/10.4161/21624011.2014.968434
Ajani JA, Lee J, Sano T, Janjigian YY, Fan D, Song S (2017) Gastric adenocarcinoma. Nat Rev Dis Primers 3:17036. https://doi.org/10.1038/nrdp.2017.36
Alcantara M, Du Rusquec P, Romano E (2020) Current clinical evidence and potential solutions to increase benefit of CAR T-cell therapy for patients with solid tumors. Oncoimmunology 9(1):1777064. https://doi.org/10.1080/2162402x.2020.1777064
Auslander N, Zhang G, Lee JS, Frederick DT, Miao B, Moll T, Tian T, Wei Z, Madan S, Sullivan RJ, Boland G, Flaherty K, Herlyn M, Ruppin E (2018) Robust prediction of response to immune checkpoint blockade therapy in metastatic melanoma. Nat Med 24(10):1545–1549. https://doi.org/10.1038/s41591-018-0157-9
Bagchi S, Yuan R, Engleman EG (2021) Immune checkpoint inhibitors for the treatment of cancer: clinical impact and mechanisms of response and resistance. Annu Rev Pathol 16:223–249. https://doi.org/10.1146/annurev-pathol-042020-042741
Barbie DA, Tamayo P, Boehm JS, Kim SY, Moody SE, Dunn IF, Schinzel AC, Sandy P, Meylan E, Scholl C, Fröhling S, Chan EM, Sos ML, Michel K, Mermel C, Silver SJ, Weir BA, Reiling JH, Sheng Q, Gupta PB et al (2009) Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462(7269):108–112. https://doi.org/10.1038/nature08460
Bezu L, Gomes-de-Silva LC, Dewitte H, Breckpot K, Fucikova J, Spisek R, Galluzzi L, Kepp O, Kroemer G (2015) Combinatorial strategies for the induction of immunogenic cell death. Front Immunol 6:187. https://doi.org/10.3389/fimmu.2015.00187
Boku N, Satoh T, Ryu MH, Chao Y, Kato K, Chung HC, Chen JS, Muro K, Kang WK, Yeh KH, Yoshikawa T, Oh SC, Bai LY, Tamura T, Lee KW, Hamamoto Y, Kim JG, Chin K, Oh DY, Minashi K et al (2021) Nivolumab in previously treated advanced gastric cancer (ATTRACTION-2): 3-year update and outcome of treatment beyond progression with nivolumab. Gastric Cancer 24(4):946–958. https://doi.org/10.1007/s10120-021-01173-w
Casares N, Pequignot MO, Tesniere A, Ghiringhelli F, Roux S, Chaput N, Schmitt E, Hamai A, Hervas-Stubbs S, Obeid M, Coutant F, Métivier D, Pichard E, Aucouturier P, Pierron G, Garrido C, Zitvogel L, Kroemer G (2005) Caspase-dependent immunogenicity of doxorubicin-induced tumor cell death. J Exp Med 202(12):1691–1701. https://doi.org/10.1084/jem.20050915
Charoentong P, Finotello F, Angelova M, Mayer C, Efremova M, Rieder D, Hackl H, Trajanoski Z (2017) Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep 18(1):248–262. https://doi.org/10.1016/j.celrep.2016.12.019
Colaprico A, Silva TC, Olsen C, Garofano L, Cava C, Garolini D, Sabedot TS, Malta TM, Pagnotta SM, Castiglioni I, Ceccarelli M, Bontempi G, Noushmehr H (2016) TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res 44(8):e71. https://doi.org/10.1093/nar/gkv1507
Conte A, Paladino S, Bianco G, Fasano D, Gerlini R, Tornincasa M, Renna M, Fusco A, Tramontano D, Pierantoni GM (2017) High mobility group A1 protein modulates autophagy in cancer cells. Cell Death Differ 24(11):1948–1962. https://doi.org/10.1038/cdd.2017.117
Cristescu R, Lee J, Nebozhyn M, Kim KM, Ting JC, Wong SS, Liu J, Yue YG, Wang J, Yu K, Ye XS, Do IG, Liu S, Gong L, Fu J, Jin JG, Choi MG, Sohn TS, Lee JH, Bae JM et al (2015) Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nat Med 21(5):449–456. https://doi.org/10.1038/nm.3850
Davis S, Meltzer PS (2007) GEOquery: a bridge between the gene expression omnibus (GEO) and bioconductor. Bioinformatics 23(14):1846–1847. https://doi.org/10.1093/bioinformatics/btm254
DeNardo DG, Brennan DJ, Rexhepaj E, Ruffell B, Shiao SL, Madden SF, Gallagher WM, Wadhwani N, Keil SD, Junaid SA, Rugo HS, Hwang ES, Jirström K, West BL, Coussens LM (2011) Leukocyte complexity predicts breast cancer survival and functionally regulates response to chemotherapy. Cancer Discov 1(1):54–67. https://doi.org/10.1158/2159-8274.Cd-10-0028
Doig KD, Fellowes A, Scott P, Fox SB (2022) Tumour mutational burden: an overview for pathologists. Pathology 54(3):249–253. https://doi.org/10.1016/j.pathol.2021.11.008
Elliott MR, Chekeni FB, Trampont PC, Lazarowski ER, Kadl A, Walk SF, Park D, Woodson RI, Ostankovich M, Sharma P, Lysiak JJ, Harden TK, Leitinger N, Ravichandran KS (2009) Nucleotides released by apoptotic cells act as a find-me signal to promote phagocytic clearance. Nature 461(7261):282–286. https://doi.org/10.1038/nature08296
Fuchs CS, Doi T, Jang RW, Muro K, Satoh T, Machado M, Sun W, Jalal SI, Shah MA, Metges JP, Garrido M, Golan T, Mandala M, Wainberg ZA, Catenacci DV, Ohtsu A, Shitara K, Geva R, Bleeker J, Ko AH et al (2018) Safety and efficacy of pembrolizumab monotherapy in patients with previously treated advanced gastric and gastroesophageal junction cancer: phase 2 clinical KEYNOTE-059 trial. JAMA Oncol 4(5):e180013. https://doi.org/10.1001/jamaoncol.2018.0013
Galluzzi L, Bravo-San Pedro JM, Demaria S, Formenti SC, Kroemer G (2017) Activating autophagy to potentiate immunogenic chemotherapy and radiation therapy. Nat Rev Clin Oncol 14(4):247–258. https://doi.org/10.1038/nrclinonc.2016.183
Galluzzi L, Vitale I, Aaronson SA, Abrams JM, Adam D, Agostinis P, Alnemri ES, Altucci L, Amelio I, Andrews DW, Annicchiarico-Petruzzelli M, Antonov AV, Arama E, Baehrecke EH, Barlev NA, Bazan NG, Bernassola F, Bertrand MJM, Bianchi K, Blagosklonny MV et al (2018) Molecular mechanisms of cell death: recommendations of the nomenclature committee on cell death 2018. Cell Death Differ 25(3):486–541. https://doi.org/10.1038/s41418-017-0012-4
Galluzzi L, Vitale I, Warren S, Adjemian S, Agostinis P, Martinez AB, Chan TA, Coukos G, Demaria S, Deutsch E, Draganov D, Edelson RL, Formenti SC, Fucikova J, Gabriele L, Gaipl US, Gameiro SR, Garg AD, Golden E, Han J et al (2020) Consensus guidelines for the definition, detection and interpretation of immunogenic cell death. J Immunother Cancer. https://doi.org/10.1136/jitc-2019-000337
Galon J, Costes A, Sanchez-Cabo F, Kirilovsky A, Mlecnik B, Lagorce-Pagès C, Tosolini M, Camus M, Berger A, Wind P, Zinzindohoué F, Bruneval P, Cugnenc PH, Trajanoski Z, Fridman WH, Pagès F (2006) Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313(5795):1960–1964. https://doi.org/10.1126/science.1129139
Garg AD, De Ruysscher D, Agostinis P (2016) Immunological metagene signatures derived from immunogenic cancer cell death associate with improved survival of patients with lung, breast or ovarian malignancies: a large-scale meta-analysis. Oncoimmunology 5(2):e1069938. https://doi.org/10.1080/2162402x.2015.1069938
Halama N, Michel S, Kloor M, Zoernig I, Benner A, Spille A, Pommerencke T, von Knebel DM, Folprecht G, Luber B, Feyen N, Martens UM, Beckhove P, Gnjatic S, Schirmacher P, Herpel E, Weitz J, Grabe N, Jaeger D (2011) Localization and density of immune cells in the invasive margin of human colorectal cancer liver metastases are prognostic for response to chemotherapy. Cancer Res 71(17):5670–5677. https://doi.org/10.1158/0008-5472.Can-11-0268
Hänzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14:7. https://doi.org/10.1186/1471-2105-14-7
Hinshaw DC, Shevde LA (2019) the tumor microenvironment innately modulates cancer progression. Cancer Res 79(18):4557–4566. https://doi.org/10.1158/0008-5472.Can-18-3962
Hunger A, Medrano RF, Zanatta DB, Del Valle PR, Merkel CA, Salles TA, Ferrari DG, Furuya TK, Bustos SO, de Freitas SR, Costanzi-Strauss E, Strauss BE (2017) Reestablishment of p53/Arf and interferon-β pathways mediated by a novel adenoviral vector potentiates antiviral response and immunogenic cell death. Cell Death Discov 3:17017. https://doi.org/10.1038/cddiscovery.2017.17
Jiang P, Gu S, Pan D, Fu J, Sahu A, Hu X, Li Z, Traugh N, Bu X, Li B, Liu J, Freeman GJ, Brown MA, Wucherpfennig KW, Liu XS (2018) Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med 24(10):1550–1558. https://doi.org/10.1038/s41591-018-0136-1
Kepp O, Zitvogel L, Kroemer G (2019) Clinical evidence that immunogenic cell death sensitizes to PD-1/PD-L1 blockade. Oncoimmunology 8(10):e1637188. https://doi.org/10.1080/2162402x.2019.1637188
Kroemer G, Galluzzi L, Kepp O, Zitvogel L (2013) Immunogenic cell death in cancer therapy. Annu Rev Immunol 31:51–72. https://doi.org/10.1146/annurev-immunol-032712-100008
Ladoire S, Mignot G, Dabakuyo S, Arnould L, Apetoh L, Rébé C, Coudert B, Martin F, Bizollon MH, Vanoli A, Coutant C, Fumoleau P, Bonnetain F, Ghiringhelli F (2011) In situ immune response after neoadjuvant chemotherapy for breast cancer predicts survival. J Pathol 224(3):389–400. https://doi.org/10.1002/path.2866
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Maeser D, Gruener RF, Huang RS (2021) OncoPredict: an R package for predicting in vivo or cancer patient drug response and biomarkers from cell line screening data. Brief Bioinform. https://doi.org/10.1093/bib/bbab260
Mayakonda A, Lin DC, Assenov Y, Plass C, Koeffler HP (2018) Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res 28(11):1747–1756. https://doi.org/10.1101/gr.239244.118
Mlecnik B, Tosolini M, Kirilovsky A, Berger A, Bindea G, Meatchi T, Bruneval P, Trajanoski Z, Fridman WH, Pagès F, Galon J (2011) Histopathologic-based prognostic factors of colorectal cancers are associated with the state of the local immune reaction. J Clin Oncol 29(6):610–618. https://doi.org/10.1200/jco.2010.30.5425
Network TCGAR (2014) Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513(7517):202–209. https://doi.org/10.1038/nature13480
Niccolai E, Taddei A, Prisco D, Amedei A (2015) Gastric cancer and the epoch of immunotherapy approaches. World J Gastroenterol 21(19):5778–5793. https://doi.org/10.3748/wjg.v21.i19.5778
Obeid M, Tesniere A, Ghiringhelli F, Fimia GM, Apetoh L, Perfettini JL, Castedo M, Mignot G, Panaretakis T, Casares N, Métivier D, Larochette N, van Endert P, Ciccosanti F, Piacentini M, Zitvogel L, Kroemer G (2007) Calreticulin exposure dictates the immunogenicity of cancer cell death. Nat Med 13(1):54–61. https://doi.org/10.1038/nm1523
Pardoll DM (2012) The blockade of immune checkpoints in cancer immunotherapy. Nat Rev Cancer 12(4):252–264. https://doi.org/10.1038/nrc3239
Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, Lee W, Yuan J, Wong P, Ho TS, Miller ML, Rekhtman N, Moreira AL, Ibrahim F, Bruggeman C, Gasmi B, Zappasodi R, Maeda Y, Sander C, Garon EB et al (2015) Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348(6230):124–128. https://doi.org/10.1126/science.aaa1348
Rosenberg JE, Hoffman-Censits J, Powles T, van der Heijden MS, Balar AV, Necchi A, Dawson N, O’Donnell PH, Balmanoukian A, Loriot Y, Srinivas S, Retz MM, Grivas P, Joseph RW, Galsky MD, Fleming MT, Petrylak DP, Perez-Gracia JL, Burris HA, Castellano D et al (2016) Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387(10031):1909–1920. https://doi.org/10.1016/s0140-6736(16)00561-4
Ruan H, Leibowitz BJ, Zhang L, Yu J (2020) Immunogenic cell death in colon cancer prevention and therapy. Mol Carcinog 59(7):783–793. https://doi.org/10.1002/mc.23183
Rusakiewicz S, Semeraro M, Sarabi M, Desbois M, Locher C, Mendez R, Vimond N, Concha A, Garrido F, Isambert N, Chaigneau L, Le Brun-Ly V, Dubreuil P, Cremer I, Caignard A, Poirier-Colame V, Chaba K, Flament C, Halama N, Jäger D et al (2013) Immune infiltrates are prognostic factors in localized gastrointestinal stromal tumors. Cancer Res 73(12):3499–3510. https://doi.org/10.1158/0008-5472.Can-13-0371
Sharma P, Allison JP (2015) The future of immune checkpoint therapy. Science 348(6230):56–61. https://doi.org/10.1126/science.aaa8172
Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A, Walsh LA, Postow MA, Wong P, Ho TS, Hollmann TJ, Bruggeman C, Kannan K, Li Y, Elipenahli C, Liu C, Harbison CT, Wang L, Ribas A, Wolchok JD et al (2014) Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med 371(23):2189–2199. https://doi.org/10.1056/NEJMoa1406498
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249. https://doi.org/10.3322/caac.21660
Tesniere A, Schlemmer F, Boige V, Kepp O, Martins I, Ghiringhelli F, Aymeric L, Michaud M, Apetoh L, Barault L, Mendiboure J, Pignon JP, Jooste V, van Endert P, Ducreux M, Zitvogel L, Piard F, Kroemer G (2010) Immunogenic death of colon cancer cells treated with oxaliplatin. Oncogene 29(4):482–491. https://doi.org/10.1038/onc.2009.356
Vacchelli E, Ma Y, Baracco EE, Sistigu A, Enot DP, Pietrocola F, Yang H, Adjemian S, Chaba K, Semeraro M, Signore M, De Ninno A, Lucarini V, Peschiaroli F, Businaro L, Gerardino A, Manic G, Ulas T, Günther P, Schultze JL et al (2015) Chemotherapy-induced antitumor immunity requires formyl peptide receptor 1. Science 350(6263):972–978. https://doi.org/10.1126/science.aad0779
Vanpouille-Box C, Alard A, Aryankalayil MJ, Sarfraz Y, Diamond JM, Schneider RJ, Inghirami G, Coleman CN, Formenti SC, Demaria S (2017) DNA exonuclease Trex1 regulates radiotherapy-induced tumour immunogenicity. Nat Commun 8:15618. https://doi.org/10.1038/ncomms15618
Wang B, Zou D, Wang N, Wang H, Zhang T, Gao L, Ma C, Zheng P, Gu B, Li X, Wang Y, He P, Ma Y, Wang X, Chen H (2022) Construction and validation of a novel coagulation-related 7-gene prognostic signature for gastric cancer. Front Genet 13:957655. https://doi.org/10.3389/fgene.2022.957655
Wilkerson MD, Hayes DN (2010) ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26(12):1572–1573. https://doi.org/10.1093/bioinformatics/btq170
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, Feng T, Zhou L, Tang W, Zhan L, Fu X, Liu S, Bo X, Yu G (2021) ClusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation (Camb) 2(3):100141. https://doi.org/10.1016/j.xinn.2021.100141
Xin D, Man Y, Yang Y, Wang F (2022) A novel prognostic and therapeutic target biomarker based on necroptosis-related gene signature and immune microenvironment infiltration in gastric cancer. Front Genet 13:953997. https://doi.org/10.3389/fgene.2022.953997
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW, Levine DA, Carter SL, Getz G, Stemke-Hale K, Mills GB, Verhaak RG (2013) Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun 4:2612. https://doi.org/10.1038/ncomms3612
Zhan X, Wang B, Li Z, Li J, Wang H, Chen L, Jiang H, Wu M, Xiao J, Peng X, Ma H, Feng D, Wang D, Fu Q, Wang M, Shen F, Hao Q, Zhang L, Xu J, Zhang Y (2019) Phase I trial of Claudin 18.2-specific chimeric antigen receptor T cells for advanced gastric and pancreatic adenocarcinoma. J Clin Oncol 37:2509–2509. https://doi.org/10.1200/JCO.2019.37.15_suppl.2509
Zhang C, Liu T, Wang J, Zhang J (2022) Development and verification of an immune-related gene prognostic index for gastric cancer. Sci Rep 12(1):15693. https://doi.org/10.1038/s41598-022-20007-y
Zhang K, Peng Z, Huang X, Qiao Z, Wang X, Wang N, Xi H, Cui J, Gao Y, Huang X, Gao H, Wei B, Chen L (2017) Phase II trial of adjuvant immunotherapy with autologous tumor-derived Gp96 vaccination in patients with gastric cancer. J Cancer 8(10):1826–1832. https://doi.org/10.7150/jca.18946
Zhou X, Zhang B, Zheng G, Zhang Z, Wu J, Du K, Zhang J (2022) Novel necroptosis-related gene signature for predicting early diagnosis and prognosis and immunotherapy of gastric cancer. Cancers (Basel) 14(16):3891. https://doi.org/10.3390/cancers14163891
Funding
This study was supported by the National Key Research and Development Program of China (2018YFC2002000).
Author information
Authors and Affiliations
Contributions
The authors confirm contribution to the paper as follows: study conception and design: YM; data collection: ZJ; analysis and interpretation of results: DC; validation of results: MW; visualization of results: MZ and YH; draft manuscript preparation: YM and SX; manuscript review & editing: ZX; All authors reviewed the results and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Ethical Approval
This study does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Meng, Y., Jin, Z., Wang, M. et al. Definition of a Novel Immunogenic Cell Death-Relevant Gene Signature Associated with Immune Landscape in Gastric Cancer. Biochem Genet 61, 2092–2115 (2023). https://doi.org/10.1007/s10528-023-10361-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-023-10361-5