Abstract
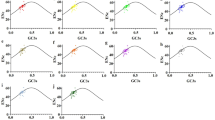
Codon usage bias of coding sequences has been usually used for exploring the evolutionary factors that affect the variation of genes. We took 20 chloroplast genomes of Malus species into account to explore the codon usage patterns, including the composition, relationship between GC3s and effective number of codons, the parity rule two analyses, the relative synonymous codon usage, the codon adaptation index, the frequency of optimal codons, the codon bias index, etc., of their coding genes. The relationship between GC3 and the ENC values showed that when the separate genes are concerned, the distribution of their GC3 contents is relatively concentrated and the distribution of the ENC values are from 35 to 61 or so. The neutrality plot showed that the correlation coefficient between GC12 and GC3 is 0.095, revealing the mutation factor played a weak role in codon pattern formation. Correspondence analysis results revealed that the codon usage patterns in the chloroplast genomes of Malus species are similar. All these results showed that all Malus chloroplast genomes are AT rich ones, the third bases of the codons are affected by the natural selection pressure, the first two nucleotide base of the codon are affected by mutation pressure. Some genes, such as the rsp7, psbA and ycf2 are of lower codon usage divergences, while the rps12, rps16 and ndhD are of higher codon usage divergences. Codon usage bias exists in the Malus genomes could be used for exploring the evolutionary characteristics in chloroplast genomes and for further study on evolutionary phenomenon in other species.







Similar content being viewed by others
References
Amandine C, Tatiana G, Marinus JMS, Isabel R, Pierre G (2014) The domestication and evolutionary ecology of apples. Trends Genet 30(2):57–65. https://doi.org/10.1016/j.tig.2013.10.002
Bao L, Li K, Liu Z, Han M, Zhang D (2016) Characterization of the complete chloroplast genome of the Chinese crabapple Malus prunifolia (Rosales: Rosaceae: Maloideae). Conserv Genet Resour 8(3):227–229. https://doi.org/10.1007/s12686-016-0540-0
Bastolla U, Dehouck Y, Echave J (2017) What evolution tells us about protein physics, and protein physics tells us about evolution. Curr Opin Struct Biol 42:59–66. https://doi.org/10.1016/j.sbi.2016.10.020
Challabathula D, Zhang Q, Bartels D (2018) Protection of photosynthesis in desiccation-tolerant resurrection plants. J Plant Physiol 227:84–92. https://doi.org/10.1016/j.jplph.2018.05.002
Deb B, Uddin A, Mazumder GA, Chakraborty S (2018) Analysis of codon usage pattern of mitochondrial protein-coding genes in different hookworms. Mol Biochem Parasitol 219:24–32. https://doi.org/10.1016/j.molbiopara.2017.11.005
Debadin B, Subhasis M (2019) Comparative genomics of a few members of the family Aquificaceae on the basis of their codon usage profile. Gene Reports 14(10):54–64. https://doi.org/10.1016/j.genrep.2018.11.003
Dong W, Xu C, Li C, Sun J, Zuo Y, Shi S, Cheng T, Guo J, Zhou S (2015) ycf1, the most promising plastid DNA barcode of land plants. Sci Rep 5:8348. https://doi.org/10.1038/srep08348
Gichira AW, Avoga S, Li Z, Hu G, Wang Q, Chen J (2019) Comparative genomics of 11 complete chloroplast genomes of Senecioneae (Asteraceae) species: DNA barcodes and phylogenetics. Bot Stud 60(1):17. https://doi.org/10.1186/s40529-019-0265-y
Haruo S, Brian RM (2016) Codon adaptation of plastid genes. PLoS One 11(5):e0154306. https://doi.org/10.1371/journaLpone.0154306
He Y, Xiao H, Deng C, Xiong L, Yang J, Peng C (2016) The complete chloroplast genome sequences of the medicinal plant Pogostemon cablin. Int J Mol Sci 17:820. https://doi.org/10.3390/ijms17060820
Hosokawa K, Shibata T, Nakamura I, Hishida A (2004) Discrimination among species of papaver based on the plastid rpl16 gene and the rpl16–rpl14 spacer sequence. Forensic Sci Int 139(2–3):195–199. https://doi.org/10.1016/j.forsciint.2003.11.001
Jenkins GM, Holmes EC (2003) The extent of codon usage bias in human RNA viruses and its evolutionary origin. Virus Res 92(1):1–7. https://doi.org/10.1016/s0168-1702(02)00309-x
Koh EJ, Song WY, Lee Y, Kim HK, Kim K, Chung N, Lee KW, Hong SW, Lee H (2006) Expression of yeast cadmium factor 1 (YCF1) confers salt tolerance to Arabidopsis thaliana. Plant Sci 170(3):534–541. https://doi.org/10.1016/j.plantsci.2005.10.007
Kong WQ, Yang JH (2017) The complete chloroplast genome sequence of Morus cathayana and Morus multicaulis, and comparative analysis within genus Morus L. PeerJ 8(5):e3037. https://doi.org/10.7717/peerj.3037
Li S, Yang J (2014) System analysis of synonymous codon usage biases in archaeal virus genomes. J Theor Biol 355:128–139. https://doi.org/10.1016/j.jtbi.2014.03.022
Li N, Sun M, Jiang Z, Shu H, Zhang S (2016) Genome-wide analysis of the synonymous codon usage patterns in apple. J Integr Agr 15(5):983–991. https://doi.org/10.1016/S2095-3119(16)61333-3
Li G, Ren Y, Pan H, Zhang L (2018) Comprehensive analysis and comparison on the codon usage pattern of whole Mycobacterium Tuberculosis genome from different area. Biomed Res Int 2018:3574976. https://doi.org/10.1155/2018/3574976
Li Y, Liu Y, Wu P, Zhou S, Wang L, Zhou S (2020) The complete chloroplast genome sequence of Malus toringoides (Rosaceae). Mitochondrial DNA B 5(3):2790–2792. https://doi.org/10.1080/23802359.2020.1780977
Li C, Cai C, Tao Y, Sun Z, Jiang M, Chen L, Li J (2021a) Variation and evolution of the whole chloroplast genomes of Fragaria spp (Rosaceae). Front Plant Sci 12:754209
Li G, Zhang L, Xue P (2021b) Codon usage pattern and genetic diversity in chloroplast genomes of Panicum species. Gene 802:145866. https://doi.org/10.1016/j.gene.2021.145866
Li G, Zhang L, Xue P (2022) Codon usage divergence of important functional genes in Mycobacterium tuberculosis. Int J Biol Macromol 209:1197–1204. https://doi.org/10.1016/j.ijbiomac.2022.04.112
Liang C, Wang L, Lei J, Duan B, Ma W, Xiao S, Qi H, Wang Z, Liu Y, Shen X, Guo S, Hu H, Xu J, Chen S (2019) A comparative analysis of the chloroplast genomes of four salvia medicinal plants. Engineering 5(5):907–915. https://doi.org/10.1016/j.eng.2019.01.017
Liu M, Dong H, Wang M, Liu Q (2020a) Evolutionary divergence of function and expression of laccase genes in plants. J Genet 99:23. https://doi.org/10.1007/s12041-020-1184-0
Liu XY, Li Y, Ji KK, Zhu J, Ling P, Zhou T, Fan LY, Xie SQ (2020b) Genome-wide codon usage pattern analysis reveals the correlation between codon usage bias and gene expression in Cuscuta australis. Genomics 112(4):2695–2702. https://doi.org/10.1016/j.ygeno.2020.03.002
Maldonado LL, Stegmayer G, Milone DH, Oliveira G, Rosenzvit M, Kamenetzky L (2018) Whole genome analysis of codon usage in Echinococcus. Mol Biochem Parasitol 225:54–66. https://doi.org/10.1016/j.molbiopara.2018.08.001
Mazumdar P, Binti Othman R, Mebus K, Ramakrishnan N, Ann Harikrishna J (2017) Codon usage and codon pair patterns in non-grass monocot genomes. Ann Bot 120(6):893–909. https://doi.org/10.1093/aob/mcx112
Mazumder GA, Uddin A, Chakraborty S (2020) Analysis of codon usage pattern of mitochondrial ND genes in Platyhelminthes. Mol Biochem Parasitol 238:111294. https://doi.org/10.1016/j.molbiopara.2020.111294
McLean MJ, Wolfe KH, Devine KM (1998) Base composition skews, replication orientation, and gene orientation in 12 prokaryote genomes. J Mol Evol 47(6):691–496. https://doi.org/10.1007/pl00006428
Naizaier R, Qu Z, Wu S, Tian X (2019) The complete chloroplast genome of Malus sieversii (Rosaceae), a wild apple tree in Xinjiang. China Mitochondrial DNA B 4(1):983–984. https://doi.org/10.1080/23802359.2019.1581108
Nakamura M, Sugiura M (2007) Translation efficiencies of synonymous codons are not always correlated with codon usage in tobacco chloroplasts. Plant J 49(1):128–134. https://doi.org/10.1111/j.1365-313X.2006.02945.x
Naver H, Boudreau E, Rochaix JD (2001) Functional studies of Ycf3: its role in assembly of photosystem I and interactions with some of its subunits. Plant Cell 13(12):2731–2745. https://doi.org/10.1105/tpc.010253
Pandey S, Prasad A, Sharma N, Prasad M (2020) Linking the plant stress responses with RNA helicases. Plant Sci 299:110607. https://doi.org/10.1016/j.plantsci.2020.110607
Saurabh P, Mehanathan M, Namisha S, Vaishali C, Priya D, Shweta S, Sarita J, Saloni M, Manoj P (2019) Characterization of DEAD-box family of RNA helicases in tomato provides insights into their roles in biotic and abiotic stresses. Environ Exp Bot 158:107–116. https://doi.org/10.1016/j.envexpbot.2018.11.018
Shackelton LA, Parrish CR, Holmes EC (2006) Evolutionary basis of codon usage and nucleotide composition bias in vertebrate DNA viruses. J Mol Evol 62(5):551–563. https://doi.org/10.1007/s00239-005-0221-1
Sharp PM, Cowe E, Higgins DG, Shields DC, Wolfe KH, Wright F (1988) Codon usage patterns in Escherichia coli, Bacillus subtilis, Saccharomyces cerevisiae, Schizosaccharomyces pombe, Drosophila melanogaster and Homo sapiens; a review of the considerable within-species diversity. Nucleic Acids Res 16(17):8207–8211. https://doi.org/10.1093/nar/16.17.8207
Sophiarani Y, Arif U, Supriyo C (2019) Deciphering codon usage patterns and evolutionary forces in chloroplast genes of Camellia sinensis var. assamica and Camellia sinensis var. sinensis in comparison to Camellia pubicosta. J Integr Agr 18(12):2771–2785. https://doi.org/10.1016/S2095-3119(19)62716-4
Sueoka N (1988) Directional mutation pressure and neutral molecular evolution. P Natl Acad Sci Usa 85(8):2653–2657. https://doi.org/10.1073/pnas.85.8.2653
Sueoka N (1999) Translation-coupled violation of parity rule 2 in human genes is not the cause of heterogeneity of the DNA G+C content of third codon position. Gene 238(1):53–58. https://doi.org/10.1016/s0378-1119(99)00320-0
Supriyo C, Debojyoti N, Tarikul HM, Arif U (2017) Codon usage pattern and prediction of gene expression level in Bungarus species. Gene 604:48–60. https://doi.org/10.1016/j.gene.2016.11.023
Supriyo C, Sophiarani Y, Arif U (2020) Analysis of codon usage bias of chloroplast genes in Oryza species. Planta 252(4):67. https://doi.org/10.1007/s00425-020-03470-7
Svetlana VN, Duccio C, Vadim RVG (2013) Phylogenetic analysis of 47 chloroplast genomes clarifies the contribution of wild species to the domesticated apple maternal line. Mol Biol Evol 30(8):1751–1760. https://doi.org/10.1093/molbev/mst092
Tan W, Gao H, Zhang H, Yu X, Tian X, Jiang W, Zhou K (2020) The complete chloroplast genome of Chinese medicine (Psoralea corylifolia): molecular structures, barcoding and phylogenetic analysis. Plant Gene 21:100216. https://doi.org/10.1016/j.plgene.2019.100216
Tao P, Dai L, Luo M, Tang F, Tien P, Pan Z (2009) Analysis of synonymous codon usage in classical swine fever virus. Virus Genes 38(1):104–112. https://doi.org/10.1007/s11262-008-0296-z
Torre AR, Lin YC, Van de Peer Y, Ingvarsson PK (2015) Genome-wide analysis reveals diverged patterns of codon bias, gene expression, and rates of sequence evolution in picea gene families. Genome Biol Evol 7(4):1002–1015. https://doi.org/10.1093/gbe/evv044
Wei L, He J, Jia X, Qi Q, Liang Z, Zheng H, Ping Y, Liu S, Sun J (2014) Analysis of codon usage bias of mitochondrial genome in Bombyx mori and its relation to evolution. BMC Evol Biol 14:262. https://doi.org/10.1186/s12862-014-0262-4
Xiong AS, Peng RH, Zhuang J, Gao F, Zhu B, Fu XY, Xue Y, Jin XF, Tian YS, Zhao W, Yao QH (2009) Gene duplication, transfer, and evolution in the chloroplast genome. Biotechnol Adv 27(4):340–347. https://doi.org/10.1016/j.biotechadv.2009.01.012
Xu C, Cai X, Chen Q, Zhou H, Cai Y, Ben A (2011) Factors affecting synonymous codon usage bias in chloroplast genome of oncidium gower ramsey. Evol Bioinform 7:271–278. https://doi.org/10.4137/EBO.S8092
Xu X, Fei D, Han H, Liu H, Zhang J, Zhou Y, Xu C, Wang H, Cao H, Zhang H (2017) Comparative characterization analysis of synonymous codon usage bias in classical swine fever virus. Microb Pathog 107:368–371. https://doi.org/10.1016/j.micpath.2017.04.019
Xun M, Song J, Shi J, Li J, Shi Y, Yan J, Zhang W, Yang H (2021) Genome-wide identification of sultr genes in Malus domestica and low sulfur-induced MhSultr3;1a to increase cysteine-improving growth. Front Plant Sci 12:748242. https://doi.org/10.3389/fpls.2021.748242
Yan M, Zhao X, Zhou J, Huo Y, Ding Y, Yuan Z (2019) The complete chloroplast genome of cultivated apple (Malus domestica Cv. ‘Yantai Fuji 8’). Mitochondrial DNA B 4(1):1213–1216. https://doi.org/10.1080/23802359.2019.1591182
Yan L, Wang H, Huang X, Li Y, Yue Y, Wang Z, Tang S (2022) Chloroplast genomes of genus Tilia: comparative genomics and molecular evolution. Front Genet 13:925726. https://doi.org/10.3389/fgene.2022.925726
Yang Y, Zhu J, Feng L, Zhou T, Bai G, Yang J, Zhao G (2018) Plastid genome comparative and phylogenetic analyses of the key genera in Fagaceae: highlighting the effect of codon composition bias in phylogenetic inference. Front Plant Sci 9:82. https://doi.org/10.3389/fpls.2018.00082
Zhang W (2000) Phylogeny of the grass family (Poaceae) from rpl16 intron sequence data. Mol Phylogenet Evol 15(1):135–146. https://doi.org/10.1006/mpev.1999.0729
Zhang SD, Jin JJ, Chen SY, Chase MW, Soltis DE, Li HT, Yang JB, Li DZ, Yi TS (2017) Diversification of rosaceae since the late cretaceous based on plastid phylogenomics. New Phytol 214(3):1355–1367. https://doi.org/10.1111/nph.14461
Zhang X, Rong C, Qin L, Mo C, Fan L, Yan J, Zhang M (2018) Complete chloroplast genome sequence of Malus hupehensis: genome structure, comparative analysis, and phylogenetic relationships. Molecules 23:2917. https://doi.org/10.3390/molecules23112917
Acknowledgements
The authors of this manuscript sincerely appreciate the great efforts of all researchers who have contributed the data to the public databases of GenBank.
Funding
The authors have not any funding support.
Author information
Authors and Affiliations
Contributions
GL conceived, designed, and supervised the overall study. LZ, PX, and MZ conducted data processing and computational analyses. GL plotted the figures and drafted the manuscript, MZ revised the manuscript. All authors contributed to the article and agreed to the submitted version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, G., Zhang, L., Xue, P. et al. Comparative Analysis on the Codon Usage Pattern of the Chloroplast Genomes in Malus Species. Biochem Genet 61, 1050–1064 (2023). https://doi.org/10.1007/s10528-022-10302-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-022-10302-8