Abstract
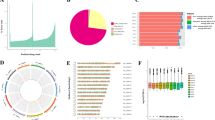
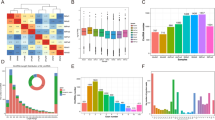
The Long non-coding RNA (lncRNA) expression profile data of ten samples including human Mesenchymal Stem Cell (MSC) adipogenic differentiation 0, 3, and 6 days from the GEO database, and then perform gene ID conversion, BLAST comparison, and annotation marking. Finally, group A (treatment group on day 3 of differentiation and control group on day 0 of differentiation) obtained a total of 1180 mRNA and 185 lncRNA; group B (treatment group on day 6 of differentiation and control group on day 0 of differentiation). A total of 1376 mRNA and 206 lncRNA were obtained. Finally, we processed the differential lncRNAs and mRNAs obtained in the two groups, and obtained 113 shared differential lncRNAs to further predict the targeted miRNA, a total of 815 lncRNA-miRNA pairs. The targeted mRNA was further predicted, and the grouped differential mRNAs were combined to obtain 64 differential mRNAs. In the end, we obtained 216 ceRNAs containing 26 lncRNAs, 27 miRNAs and 64 mRNAs. We found that the mRNAs in the ceRNA network were mainly enriched with 45 Gene Ontology (GO) terms, mainly including glucose homeostasis mechanism and insulin stimulation response. 69 Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were mainly enriched. It mainly includes many pathways related to lipid metabolism such as Adenosine 5′-monophosphate (AMP)-activated protein kinase (AMPK), Rap1, cAMP, mitogen-activated protein kinase (MAPK), Ras, hypoxia inducible factor-1 (HIF-1), PI3K-Akt, insulin signaling and so on. In the end, we identified 216 ceRNA regulatory relationships related to obesity research. Our research provides a clearer direction for understanding the molecular mechanism of obesity, the screening and determination of drug targets biomarkers in the future.





Similar content being viewed by others
Data Availability
All the data are available within the manuscript.
References
Agapito G (2019) Computer tools to analyze microarray data. Methods Mol Biol 1986:267–282
Alonso-Betanzos A, Bolón-Canedo V, Morán-Fernández L, Sánchez-Maroño N (2019) A Review of microarray datasets: where to find them and specific characteristics. Methods Mol Biol 1986:65–85
Behzadi P, Ranjbar R (2019) DNA microarray technology and bioinformatic web services. Acta Microbiol Immunol Hung 66(1):19–30
Brill MJ, Diepstraten J, van Rongen A, van Kralingen S, van den Anker JN, Knibbe CA (2012) Impact of obesity on drug metabolism and elimination in adults and children. Clin Pharmacokinet 51(5):277–304
Cabiati M, Randazzo E, Salvadori C, Peroni D, Federico G, Del Ry S (2020) Circulating microRNAs associated with C-type natriuretic peptide in childhood obesity. Peptides 133:170387
Cai C, Min S, Yan B, Liu W, Yang X, Li L, Wang T, Jin A (2019) MiR-27a promotes the autophagy and apoptosis of IL-1β treated-articular chondrocytes in osteoarthritis through PI3K/AKT/mTOR signaling. Aging 11(16):6371–6384
Chen T, Zhang Y, Liu Y, Zhu D, Yu J, Li G, Sun Z, Wang W, Jiang H, Hong Z (2019) MiR-27a promotes insulin resistance and mediates glucose metabolism by targeting PPAR-γ-mediated PI3K/AKT signaling. Aging 11(18):7510–7524
Cox AJ, West NP, Cripps AW (2015) Obesity, inflammation, and the gut microbiota. Lancet Diabetes Endocrinol 3(3):207–215
Fachel AA, Tahira AC, Vilella-Arias SA, Maracaja-Coutinho V, Gimba ER, Vignal GM, Campos FS, Reis EM, Verjovski-Almeida S (2013) Expression analysis and in silico characterization of intronic long noncoding RNAs in renal cell carcinoma: emerging functional associations. Mol Cancer 12(1):140
Font-Burgada J, Sun B, Karin M (2016) Obesity and cancer: the oil that feeds the flame. Cell Metab 23(1):48–62
Friedman JM (2000) Obesity in the new millennium. Nature 404(6778):632–634
Guo W, Jiang H, Li H, Li F, Yu Q, Liu Y, Jiang W, Zhang M (2019) LncRNA-SRA1 suppresses osteosarcoma cell proliferation while promoting cell apoptosis. Technol Cancer Res Treat 18:1533033819841438
Guo Y, Zhang X, Huang W, Miao X (2017) Identification and characterization of differentially expressed miRNAs in subcutaneous adipose between Wagyu and Holstein cattle. Sci Rep 7:44026
Hsu SD, Tseng YT, Shrestha S, Lin YL, Khaleel A, Chou CH, Chu CF, Huang HY, Lin CM, Ho SY, Jian TY, Lin FM, Chang TH, Weng SL, Liao KW, Liao IE, Liu CC, Huang HD (2014) miRTarBase update 2014: an information resource for experimentally validated miRNA-target interactions. Nucleic Acids Res 42:D78-85
Hu Y, Chen HY, Yu CY, Xu J, Wang JL, Qian J, Zhang X, Fang JY (2014) A long non-coding RNA signature to improve prognosis prediction of colorectal cancer. Oncotarget 5(8):2230–2242
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57
Huang S, Wang S, Bian C, Yang Z, Zhou H, Zeng Y, Li H, Han Q, Zhao RC (2012) Upregulation of miR-22 promotes osteogenic differentiation and inhibits adipogenic differentiation of human adipose tissue-derived mesenchymal stem cells by repressing HDAC6 protein expression. Stem Cells Dev 21(13):2531–2540
Janderová L, McNeil M, Murrell AN, Mynatt RL, Smith SR (2003) Human mesenchymal stem cells as an in vitro model for human adipogenesis. Obes Res 11(1):65–74
Jeggari A, Marks DS, Larsson E (2012) miRcode: a map of putative microRNA target sites in the long non-coding transcriptome. Bioinformatics 28(15):2062–2063
Ji L, Li X (2019) Long noncoding RNA MEG3 is a tumor suppressor in choriocarcinoma by upregulation of microRNA-211. J Cell Physiol 234(12):22911–22920
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120(1):15–20
Li J, Chen Z, Tian L, Zhou C, He MY, Gao Y, Wang S, Zhou F, Shi S, Feng X, Sun N, Liu Z, Skogerboe G, Dong J, Yao R, Zhao Y, Sun J, Zhang B, Yu Y, Shi X, Luo M, Shao K, Li N, Qiu B, Tan F, Chen R, He J (2014) LncRNA profile study reveals a three-lncRNA signature associated with the survival of patients with oesophageal squamous cell carcinoma. Gut 63(11):1700–1710
Li M, Sun X, Cai H, Sun Y, Plath M, Li C, Lan X, Lei C, Lin F, Bai Y, Chen H (2016) Long non-coding RNA ADNCR suppresses adipogenic differentiation by targeting miR-204. Biochem Biophys Acta 1859(7):871–882
Li S, Mi L, Yu L, Yu Q, Liu T, Wang GX, Zhao XY, Wu J, Lin JD (2017) Zbtb7b engages the long noncoding RNA Blnc1 to drive brown and beige fat development and thermogenesis. Proc Natl Acad Sci USA 114(34):E7111-e7120
Li X, Wu Z, Fu X, Han W (2013) Long Noncoding RNAs: Insights from Biological Features and Functions to Diseases. Med Res Rev 33(3):517–553
Lin Y, Wen-Jie Z, Chang-Qing L, Sheng-Xiang A, Yue Z (2020) mir-22-3p/KLF6/MMP14 axis in fibro-adipogenic progenitors regulates fatty infiltration in muscle degeneration. FASEB J 34(9):12691–12701
Liu M, Li B, Peng W, Ma Y, Huang Y, Lan X, Lei C, Qi X, Liu GE, Chen H (2019) LncRNA-MEG3 promotes bovine myoblast differentiation by sponging miR-135. J Cell Physiol 234(10):18361–18370
Liu S, Chen Q, Wang Y (2020) MiR-125b-5p suppresses the bladder cancer progression via targeting HK2 and suppressing PI3K/AKT pathway. Hum Cell 33(1):185–194
Liu S, Sheng L, Miao H, Saunders TL, MacDougald OA, Koenig RJ, Xu B (2014a) SRA gene knockout protects against diet-induced obesity and improves glucose tolerance. J Biol Chem 289(19):13000–13009
Liu S, Xu R, Gerin I, Cawthorn WP, Macdougald OA, Chen XW, Saltiel AR, Koenig RJ, Xu B (2014) SRA regulates adipogenesis by modulating p38/JNK phosphorylation and stimulating insulin receptor gene expression and downstream signaling. PloS one 9(4):e95416
Matsuzaki S (2011) DNA microarray analysis in endometriosis for development of more effective targeted therapies. Front Biosci (elite Ed) 3:1139–1153
Mercer TR, Dinger ME, Mattick JS (2009) Long non-coding RNAs: insights into functions. Nat Rev Genet 10(3):155–159
Mi L, Zhao XY, Li S, Yang G, Lin JD (2017) Conserved function of the long noncoding RNA Blnc1 in brown adipocyte differentiation. Mol Metab 6(1):101–110
Ren S, Zhang Y, Li B, Bu K, Wu L, Lu Y, Lu Y, Qiu Y (2019) Downregulation of lncRNA-SRA participates in the development of cardiovascular disease in type II diabetic patients. Exp Ther Med 17(5):3367–3372
Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, Chang HY (2007) Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129(7):1311–1323
Seravalle G, Grassi G (2017) Obesity and hypertension. Pharmacol Res 122:1–7
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504
Shi C, Huang F, Gu X, Zhang M, Wen J, Wang X, You L, Cui X, Ji C, Guo X (2016) Adipogenic miRNA and meta-signature miRNAs involved in human adipocyte differentiation and obesity. Oncotarget 7(26):40830–40845
Shi X, Sun M, Liu H, Yao Y, Song Y (2013) Long non-coding RNAs: a new frontier in the study of human diseases. Cancer Lett 339(2):159–166
Stamatopoulos A, Stamatopoulos T, Gamie Z, Kenanidis E, Ribeiro RDC, Rankin KS, Gerrand C, Dalgarno K, Tsiridis E (2019) Mesenchymal stromal cells for bone sarcoma treatment: Roadmap to clinical practice. J Bone Onco 16:100231
Tang S, Zhu W, Zheng F, Gui W, Zhang W, Lin X, Li H (2020) The long noncoding RNA Blnc1 protects against diet-induced obesity by promoting mitochondrial function in white fat. Diabet Metab Syndr Obes 13:1189–1201
Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F, Shi Y, Segal E, Chang HY (2010) Long noncoding RNA as modular scaffold of histone modification complexes. Science 329(5992):689–693
Vecchié A, Dallegri F, Carbone F, Bonaventura A, Liberale L, Portincasa P, Frühbeck G, Montecucco F (2018) Obesity phenotypes and their paradoxical association with cardiovascular diseases. Eur J Intern Med 48:6–17
Wang F, Yuan JH, Wang SB, Yang F, Yuan SX, Ye C, Yang N, Zhou WP, Li WL, Li W, Sun SH (2014) Oncofetal long noncoding RNA PVT1 promotes proliferation and stem cell-like property of hepatocellular carcinoma cells by stabilizing NOP2. Hepatology 60(4):1278–1290
Wang Y, Liu W, Liu Y, Cui J, Zhao Z, Cao H, Fu Z, Liu B (2018) Long noncoding RNA H19 mediates LCoR to impact the osteogenic and adipogenic differentiation of mBMSCs in mice through sponging miR-188. J Cell Physiol 233(9):7435–7446
Wang Z, Lachmann A, Ma’ayan A (2019) Mining data and metadata from the gene expression omnibus. Biophys Rev 11(1):103–110
Wei S, Du M, Jiang Z, Hausman GJ, Zhang L, Dodson MV (2016) Long noncoding RNAs in regulating adipogenesis: new RNAs shed lights on obesity. Cell Mol Life Sci 73(10):2079–2087
Wong N, Wang X (2015) miRDB: an online resource for microRNA target prediction and functional annotations. Nucleic Acids Res 43:D146-152
Xiao T, Liu L, Li H, Sun Y, Luo H, Li T, Wang S, Dalton S, Zhao RC, Chen R (2015) Long noncoding RNA ADINR regulates adipogenesis by transcriptionally activating C/EBPα. Stem Cell Rep 5(5):856–865
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li CY, Wei L (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39:W316-322
Xu B, Gerin I, Miao H, Vu-Phan D, Johnson CN, Xu R, Chen XW, Cawthorn WP, MacDougald OA, Koenig RJ (2010) Multiple roles for the non-coding RNA SRA in regulation of adipogenesis and insulin sensitivity. PloS One 5(12):e14199
Yu Y, Du H, Wei S, Feng L, Li J, Yao F, Zhang M, Hatch GM, Chen L (2018) Adipocyte-derived exosomal MiR-27a induces insulin resistance in skeletal muscle through repression of PPARγ. Theranostics 8(8):2171–2188
Zhang CJ, Liu C, Wang YX, Zhu N, Hu ZY, Liao DF, Qin L (2019) Long non-coding RNA-SRA promotes neointimal hyperplasia and vascular smooth muscle cells proliferation via MEK-ERK-CREB pathway. Vascul Pharmacol 116:16–23
Zhao J, Sun BK, Erwin JA, Song JJ, Lee JT (2008) Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science (new York, NY) 322(5902):750–756
Zheng Y, Cai B, Li X, Li D, Yin G (2019) MiR-125b-5p and miR-181b-5p inhibit keratinocyte proliferation in skin by targeting Akt3. Eur J Pharmacol 862:172659
Acknowledgements
All the authors acknowledge and thank their respective institutes & universities.
Funding
This work was supported by grants from the National Key Research and Development Program of China (2018YFD0501700), the National Key Technology Support Program (2015BAD03B04), the National Beef and Yak Industrial Technology System (CARS-37), the Agricultural Science and Technology Innovation and Transformation Project of Shaanxi Province (NYKJ-2018-LY09) and the Technical Innovation Engineering Project of Shaanxi Province (2016 KTCL02-15).
Author information
Authors and Affiliations
Contributions
CL and SHAR conceptualization, data curation, formal analysis, investigation, methodology, software, validation, writing—original draft, writing—review and editing. MARN and RK; YF conceptualization, data curation, formal analysis, methodology, software, validation, writing—review and editing. ZMM and AFS, Data curation, formal analysis, writing—review and editing. BMA and FMS Writing—review and editing. MAB software, methodology, visualization software. LZ: conceptualization, funding acquisition, project administration, resources, supervision, visualization, writing—review and editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical Approval
The data used in this study met the following criteria: these relevant data were obtained on basis of the GEO ethics committee, which is open access and public; therefore, there is no need to worry about approval.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liang, C., Raza, S.H.A., Naqvi, M.A.R. et al. Construction of Adipogenic ceRNA Network Based on lncRNA Expression Profile of Adipogenic Differentiation of Human MSC Cells. Biochem Genet 60, 543–557 (2022). https://doi.org/10.1007/s10528-021-10115-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-021-10115-1