Abstract
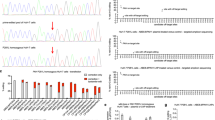
Treatment with tetrahydrobiopterin (BH4) is the latest therapeutic option approved for patients with phenylketonuria (PKU)—one of the most frequent inborn metabolic diseases. PKU or phenylalanine hydroxylase (PAH) deficiency is caused by mutations in the PAH gene. Given that some PAH mutations are responsive to BH4 treatment while others are non-responsive, for every novel mutation that is discovered it is essential to confirm its pathogenic effect and to assess its responsiveness to a BH4 treatment in vitro, before the drug is administered to patients. We found a c.676C>A (p.Gln226Lys) mutation in the PAH gene in two unrelated patients with PKU. The corresponding aberrant protein has never been functionally characterized in vitro and its response to BH4 treatment is unknown. Computational analyses proposed that glutamine at position 226 is an important, evolutionary conserved amino acid while the substitution with lysine probably disturbs tertiary protein structure and impacts posttranslational PAH modifications. Using hepatoma cellular model, we demonstrated that the amount of mutant p.Gln226Lys PAH detected by Western blot was only 1.2% in comparison to wild-type PAH. The addition of sepiapterin, intracellular precursor of BH4, did not increase PAH protein yield thus marking p.Gln226Lys as BH4-non-responsive mutation. Therefore, computational, experimental, and clinical data were all in accordance showing that p.Gln226Lys is a severe pathogenic PAH mutation. Its non-responsiveness to BH4 treatment in hepatoma cellular model should be considered when deciding treatment options for PKU patients carrying this mutation. Consequently, our study will facilitate clinical genetic practice, particularly genotype-based stratification of PKU treatment.


Similar content being viewed by others

References
Aguado C, Perez B, Ugarte M, Desviat LR (2006) Analysis of the effect of tetrahydrobiopterin on PAH gene expression in hepatoma cells. FEBS Lett 580:1697–1701. https://doi.org/10.1016/j.febslet.2006.02.005
Anjema K, van Rijn M, Hofstede FC, Bosch AM, Hollak CE, Rubio-Gozalbo E et al (2013) Tetrahydrobiopterin responsiveness in phenylketonuria: prediction with the 48-hour loading test and genotype. Orphanet J Rare Dis 8:103. https://doi.org/10.1186/1750-1172-8-103
Blau N (2016) Genetics of phenylketonuria: then and now. Hum Mutat 37:508–515. https://doi.org/10.1002/humu.22980
Djordjevic M, Klaassen K, Sarajlija A, Tosic N, Zukic B, Kecman B et al (2013) Molecular genetics and genotype-based estimation of BH4-responsiveness in Serbian PKU patients: spotlight on phenotypic implications of p. L48S. JIMD Rep 9:49–58. https://doi.org/10.1007/8904_2012_178
Gemperle-Britschgi C, Iorgulescu D, Mager MA, Anton-Paduraru D, Vulturar R, Thony B (2016) A novel common large genomic deletion and two new missense mutations identified in the Romanian phenylketonuria population. Gene 576:182–188. https://doi.org/10.1016/j.gene.2015.10.020
Groselj U, Tansek MZ, Smon A, Angelkova N, Anton D, Baric I et al (2014) Newborn screening in southeastern Europe. Mol Genet Metab 113:42–45. https://doi.org/10.1016/j.ymgme.2014.07.020
Guldberg P, Rey F, Zschocke J, Romano V, Francois B, Michiels L et al (1998) A European multicenter study of phenylalanine hydroxylase deficiency: classification of 105 mutations and a general system for genotype-based prediction of metabolic phenotype. Am J Hum Genet 63:71–79. https://doi.org/10.1086/301920
Keil S, Anjema K, van Spronsen FJ, Lambruschini N, Burlina A, Belanger-Quintana A et al (2013) Long-term follow-up and outcome of phenylketonuria patients on sapropterin: a retrospective study. Pediatrics 131:e1881–1888. https://doi.org/10.1542/peds.2012-3291
Klaassen K, Stojiljkovic M, Brasil S, Djordjevic M, Desviat LR, Pavlovic S et al (2015) Novel p.Gln226Lys mutation in phenylalanine hydroxylase gene resulting in classical PKU. J Inherit Metab Dis 38:S110
Reblova K, Hruba Z, Prochazkova D, Pazdirkova R, Pouchla S, Zeman J et al (2013) Hyperphenylalaninemia in the Czech Republic: genotype-phenotype correlations and in silico analysis of novel missense mutations. Clin Chim Acta 419:1–10. https://doi.org/10.1016/j.cca.2013.01.006
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 17:405–424. https://doi.org/10.1038/gim.2015.30
Scriver CR, Hurtubise M, Konecki D, Phommarinh M, Prevost L, Erlandsen H et al (2003) PAHdb 2003: what a locus-specific knowledgebase can do. Hum Mutat 21:333–344. https://doi.org/10.1002/humu.10200
Scriver CR, Levy H, Donlon J (2008) Hyperphenylalaninemia: phenylalanine hydroxylase deficiency. In: Valle D, Beaudet AL, Vogelstein B, Kinzler KW, Antonarakis S, Ballabio A (editors), Scriver CR, Childs B, Sly WS (editors emeritus) (eds) The online metabolic and molecular basis of inherited disease. McGraw-Hill, New York, p 77
Stojiljkovic M, Jovanovic J, Djordjevic M, Grkovic S, Cvorkov Drazic M, Petrucev B et al (2006) Molecular and phenotypic characteristics of patients with phenylketonuria in Serbia and Montenegro. Clin Genet 70:151–155. https://doi.org/10.1111/j.1399-0004.2006.00650.x
Stojiljkovic M, Perez B, Desviat LR, Aguado C, Ugarte M, Pavlovic S (2009) The Missense p. S231F phenylalanine hydroxylase gene mutation causes complete loss of enzymatic activity in vitro. Protein J 28:294–299. https://doi.org/10.1007/s10930-009-9194-z
Wettstein S, Underhaug J, Perez B, Marsden BD, Yue WW, Martinez A et al (2015) Linking genotypes database with locus-specific database and genotype-phenotype correlation in phenylketonuria. Eur J Hum Genet 23:302–309. https://doi.org/10.1038/ejhg.2014.114
Zekanowski C, Perez B, Desviat LR, Wiszniewski W, Ugarte M (2000) In vitro expression analysis of R68G and R68S mutations in phenylalanine hydroxylase gene. Acta Biochim Pol 47:365–369
Zurfluh MR, Zschocke J, Lindner M, Feillet F, Chery C, Burlina A et al (2008) Molecular genetics of tetrahydrobiopterin-responsive phenylalanine hydroxylase deficiency. Hum Mutat 29:167–175. https://doi.org/10.1002/humu.20637
Acknowledgements
This work has been funded by grants from the Ministry of Education, Science and Technological Development, Serbia (III41004) and European Commission (EU-FP7-REGPOT-316088) given to SP; the Spanish–Serbian cooperation project funded by the Ministry of Economy and Competitiveness, Spain (PRIAIBSE-2011-1126); and the Ministry of Education, Science and Technological Development, Serbia (451-03-02635/2011-14/14) given to BP and MS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
This study has been approved by the Ethics Committee of the Mother and Child Health Care Institute of Serbia “Dr Vukan Cupic” in Belgrade, Serbia, and has therefore been performed in accordance with the ethical standards laid down in the 1975 Declaration of Helsinki and its later amendments. Informed consent was obtained from all individual participants included in the study and/or their parents or legally authorized representatives.
Rights and permissions
About this article
Cite this article
Klaassen, K., Djordjevic, M., Skakic, A. et al. Functional Characterization of Novel Phenylalanine Hydroxylase p.Gln226Lys Mutation Revealed Its Non-responsiveness to Tetrahydrobiopterin Treatment in Hepatoma Cellular Model. Biochem Genet 56, 533–541 (2018). https://doi.org/10.1007/s10528-018-9858-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-018-9858-5