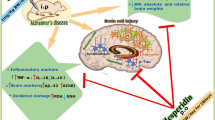
In the cerebellum, hippocampus, and prefrontal cortex of mature male Wistar rats with trained spatial navigational skill in the Morris water maze, the transcriptional activity the NAPA gene that regulates the transport and secretion of synaptic vesicles, release of neurotransmitters, and protein degradation was determined by real-time PCR. Animals subjected to forced swimming in a time-matched regime (active control) and naïve rats were used as the comparison groups. Suppression of NAPA gene activity was found in the hippocampus and cerebellum of the active control group, while navigation skill training led to a significant increase in gene expression in all brain structures under study. The findings suggest the existence of specific mechanisms regulating NAPA gene activity during the formation of spatial memory and adaptive behavior under stress conditions.
Similar content being viewed by others
References
Arleo A, Rondi-Reig L. Multimodal sensory integration and concurrent navigation strategies for spatial cognition in real and artificial organisms. J. Integr. Neurosci. 2007;6(3):327-366. doi: https://doi.org/10.1142/s0219635207001593
Sardoo AM, Zhang S, Ferraro TN, Keck TM, Chen Y. Decoding brain memory formation by single-cell RNA sequencing. Brief Bioinform. 2022;23(6):bbac412. doi: https://doi.org/10.1093/bib/bbac412
Reshetnikov VV, Kisaretova PE, Ershov NI, Shulyupova AS, Oshchepkov DY, Klimova NV, Ivanchihina AV, Merkulova TI, Bondar NP. Genes associated with cognitive performance in the Morris water maze: an RNA-seq study. Sci. Rep. 2020;10(1):22078. doi: https://doi.org/10.1038/s41598-020-78997-6
Rizo J. Molecular mechanisms underlying neurotransmitter release. Annu. Rev. Biophys. 2022;51:377-408. doi: https://doi.org/10.1146/annurev-biophys-111821-104732
Wang L, Brautigan DL. α-SNAP inhibits AMPK signaling to reduce mitochondrial biogenesis and dephosphorylates Thr172 in AMPKα in vitro. Nat. Commun. 2013;4:1559. doi: https://doi.org/10.1038/ncomms2565
Federighi G, Traina G, Macchi M, Ciampini C, Bernardi R, Baldi E, Bucherelli C, Brunelli M, Scuri R. Modulation of gene expression in contextual fear conditioning in the rat. PLoS One. 2013;8(11):e80037. doi: https://doi.org/10.1371/journal.pone.0080037
Sauvola CW, Akbergenova Y, Cunningham KL, Aponte-Santiago NA, Littleton JT. The decoy SNARE Tomosyn sets tonic versus phasic release properties and is required for homeostatic synaptic plasticity. Elife. 2021;10:e72841. doi: https://doi.org/10.7554/eLife.72841
Gruden MA, Storozheva ZI, Sewell RD, Kolobov VV, Sherstnev VV. Distinct functional brain regional integration of Casp3, Ascl1 and S100a6 gene expression in spatial memory. Behav. Brain Res. 2013;252:230-238. doi: https://doi.org/10.1016/j.bbr.2013.06.024
Gruden’ MA, Storozheva ZI, Ratmirov AM, Sherstnev VV. Pattern of Notch2, Numb, and Cas8 gene expression in relevant structures of the rat brain during formation of spatial memory. Bull. Exp. Biol. Med. 2017;163(6):785-788. doi: https://doi.org/10.1007/s10517-017-3903-y
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25(4):402-408. doi: https://doi.org/10.1006/meth.2001.1262
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B. 1995;57(1):289-300. doi: https://doi.org/10.2307/2346101
Poulter S, Austen JM, Kosaki Y, Dachtler J, Lever C, McGregor A. En route to delineating hippocampal roles in spatial learning. Behav. Brain Res. 2019;369:111936. doi: https://doi.org/10.1016/j.bbr.2019.111936
Andre P, Zaccaroni M, Fiorenzani P, Della Seta D, Menzocchi M, Farabollini F. Offline consolidation of spatial memory: Do the cerebellar output circuits play a role? A study utilizing a Morris water maze protocol in male Wistar rats. Brain Res. 2019;1718:148-158. doi: https://doi.org/10.1016/j.brainres.2019.05.010
Tong L, Shen H, Perreau VM, Balazs R, Cotman CW. Effects of exercise on gene-expression profile in the rat hippocampus. Neurobiol. Dis. 2001;8(6):1046-1056. doi: https://doi.org/10.1006/nbdi.2001.0427
Ghosh Dastidar S, Das Sharma S, Chakraborty S, Chattarji S, Bhattacharya A, Muddashetty RS. Distinct regulation of bioenergetics and translation by group I mGluR and NMDAR. EMBO Rep. 2022;23(2):e54501. doi: https://doi.org/10.15252/embr.202154501
Author information
Authors and Affiliations
Corresponding author
Additional information
Translated from Byulleten’ Eksperimental’noi Biologii i Meditsiny, Vol. 175, No. 6, pp. 773-777, June, 2023
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gruden, M.A., Ratmirov, A.M., Storozheva, Z.I. et al. Analysis of NAPA Gene Expression in Brain Structures of Wistar Rats during the Formation of Long-Term Spatial Memory and Physical Activity under Stress Situation. Bull Exp Biol Med 175, 810–813 (2023). https://doi.org/10.1007/s10517-023-05952-6
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10517-023-05952-6