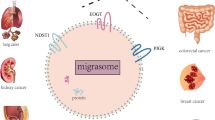
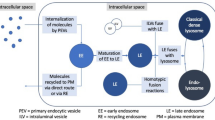
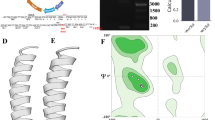
One of the potential causes of cancer recurrence is disruption of the cell—cell communication in the primary tumors that is realized, among other things, through secretion and uptake of exosomes by cells. Low expression of the IGFBP6 gene (insulin-like growth factor binding protein 6) is associated with a high recurrence rate and can serve as a prognostic marker of luminal breast cancer. The knockdown of the IGFBP6 gene leads to significant changes in lipid metabolism. We performed a quantitative analysis of both exosomes and proteins involved in the mechanism of their biogenesis. Changes in the expression profile of mRNAs and their proteins responsible for the synthesis and secretion of exosomes were revealed. We showed a decrease in the expression of the of the VPS28 gene mRNA (vacuolar protein sorting-associated protein 28) and the corresponding protein by 2.3 and 5.6 times, respectively. The secretion of exosomes by MDA-MB-231 cells with IGFBP6 knockdown decreased by 2 times. We discussed a mechanism of disruption of cell—cell communication.
Similar content being viewed by others
References
Nikulin S, Zakharova G, Poloznikov A, Raigorodskaya M, Wicklein D, Schumacher U, Nersisyan S, Bergquist J, Bakalkin G, Astakhova L, Tonevitsky A. Effect of the expression of ELOVL5 and IGFBP6 genes on the metastatic potential of breast cancer cells. Front. Genet. 2021;12:662843. https://doi.org/10.3389/fgene.2021.662843
Song F, Zhou X.X, Hu Y, Li G, Wang Y. The roles of Insulin-Like Growth Factor binding protein family in development and diseases. Adv. Ther. 2021;38(2):885-903. https://doi.org/10.1007/s12325-020-01581-x
Bach LA, Fu P, Yang Z. Insulin-like growth factor-binding protein-6 and cancer. Clin. Sci. (Lond). 2013;124(4):215-229. https://doi.org/10.1042/CS20120343
Turchinovich A, Tonevitsky AG, Cho WC, Burwinkel B. Check and mate to exosomal extracellular miRNA: new lesson from a new approach. Front. Mol. Biosci. 2015;2:11. https://doi.org/10.3389/fmolb.2015.00011
Palazzolo G, Albanese NN, DI Cara G, Gygax D, Vittorelli ML, Pucci-Minafra I. Proteomic analysis of exosome-like vesicles derived from breast cancer cells. Anticancer Res. 2012;32(3):847-860.
Melo SA, Sugimoto H, O’Connell JT, Kato N, Villanueva A, Vidal A, Qiu L, Vitkin E, Perelman LT, Melo CA, Lucci A, Ivan C, Calin GA, Kalluri R. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell. 2014;26(5):707-721. https://doi.org/10.1016/j.ccell.2014.09.005
Meldolesi J. Exosomes and ectosomes in intercellular communication. Curr. Biol. 2018;28(8):R435-R444. https://doi.org/10.1016/j.cub.2018.01.059
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D. The human genome browser at UCSC. Genome Res. 2002;12(6):996-1006. https://doi.org/10.1101/gr.229102
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics. 2012;13:134. https://doi.org/10.1186/1471-2105-13-134
Owczarzy R, Tataurov AV, Wu Y, Manthey JA, McQuisten KA, Almabrazi HG, Pedersen KF, Lin Y, Garretson J, McEntaggart NO, Sailor CA, Dawson RB, Peek AS. IDT SciTools: a suite for analysis and design of nucleic acid oligomers. Nucleic Acids Res. 2008;36(Web Server issue):W163-9. https://doi.org/10.1093/nar/gkn198
Maltseva DV, Khaustova NA, Fedotov NN, Matveeva EO, Lebedev AE, Shkurnikov MU, Galatenko VV, Schumacher U, Tonevitsky AG. High-throughput identification of reference genes for research and clinical RT-qPCR analysis of breast cancer samples. J. Clin. Bioinforma. 2013;3(1):13. https://doi.org/10.1186/2043-9113-3-13
Miyanishi M, Tada K, Koike M, Uchiyama Y, Kitamura T, Nagata S. Identification of Tim4 as a phosphatidylserine receptor. Nature. 2007;450:435-439. https://doi.org/10.1038/nature06307
Muller L, Hong CS, Stolz DB, Watkins SC, Whiteside TL. Isolation of biologically-active exosomes from human plasma. J. Immunol. Methods. 2014;411:55-65. https://doi.org/10.1016/j.jim.2014.06.007
Wu Y, Deng W, Klinke DJ 2nd. Exosomes: improved methods to characterize their morphology, RNA content, and surface protein biomarkers. Analyst. 2015;140(19):6631-6642. https://doi.org/10.1039/c5an00688k
Williams RL, Urbé S. The emerging shape of the ESCRT machinery. Nat. Rev. Mol. Cell Biol. 2007;8(5):355-368. https://doi.org/10.1038/nrm2162
Hessvik NP, Llorente A. Current knowledge on exosome biogenesis and release. Cell. Mol. Life Sci. 2018;75(2):193-208. https://doi.org/10.1007/s00018-017-2595-9
Malhi H. Emerging role of extracellular vesicles in liver diseases. Am. J. Physiol. Gastrointest. Liver Physiol. 2019;317(5):G739-G749. https://doi.org/10.1152/ajpgi.00183.2019
Spencer N, Yeruva L. Role of bacterial infections in extracellular vesicles release and impact on immune response. Biomed. J. 2021;44(2):157-164. https://doi.org/10.1016/j.bj.2020.05.006
Phuyal S, Hessvik NP, Skotland T, Sandvig K, Llorente A. Regulation of exosome release by glycosphingolipids and flotillins. FEBS J. 2014;281(9):2214-2227. https://doi.org/10.1111/febs.12775
Villarroya-Beltri C, Baixauli F, Gutiérrez-Vázquez C, Sánchez-Madrid F, Mittelbrunn M. Sorting it out: regulation of exosome loading. Semin. Cancer Biol. 2014;28:3-13. https://doi.org/10.1016/j.semcancer.2014.04.009
Bobrie A, Colombo M, Raposo G, Théry C. Exosome secretion: molecular mechanisms and roles in immune responses. Traffic. 2011;12(12):1659-1668. https://doi.org/10.1111/j.1600-0854.2011.01225.x
Huang J, Xiong J, Yang L, Zhang J, Sun S, Liang Y. Cell-free exosome-laden scaffolds for tissue repair. Nanoscale. 2021;13(19): 8740-8750. https://doi.org/10.1039/d1nr01314a
Author information
Authors and Affiliations
Corresponding author
Additional information
Translated from Kletochnye Tekhnologii v Biologii i Meditsine, No. 1, pp. 47-51, March, 2023
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Efimova, A.S., Antipenko, I.D., Evtushenko, E.A. et al. Effect of IGFBP6 Knockdown on Proteins Regulating Exosome Synthesis and Secretion in MDA-MB-231 Cell Line. Bull Exp Biol Med 175, 157–161 (2023). https://doi.org/10.1007/s10517-023-05828-9
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10517-023-05828-9