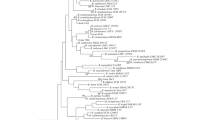
Opportunistic microorganisms in the gut biocenosis were studied in adolescents with normal body weight and obesity (patients consulted at the Clinical Department of Research Center of Family Health and Human Reproduction Problems). The biological material was studied by standard bacteriological methods, representatives of Enterobacteriaceae family were also characterized using metagenomic sequencing of V3-V4 variable regions of 16S gene rRNA. Gut microbiota of obese adolescents was unbalanced and was characterized by low levels of bifido- and lactoflora representatives, a spectrum of E. coli associations, and high prevalence of opportunistic microorganisms and their associations. Representatives of Enterobacteriaceae family were most often found in the gut microbiota of obese adolescents.
Similar content being viewed by others
References
Belkova NL, Nemchenko UM, Pogodina AV, Feranchuk SI, Romanitsa AI, Novikova EA, Rychkova LV. Composition and Structure of Gut Microbiome in Adolescents with Obesity and Different Breastfeeding Duration. Bull. Exp. Biol. Med. 2019;167(6):717-721. 2019;167(6):759-762. doi: https://doi.org/10.1007/s10517-019-04617-7
Volovnikova VA, Kotrova AD, Ivanova KA, Ermolenko EI, Shishkin AN. The role of gut microbiota in the development of obesity. Juvenis Scientia. 2019;(6):4-10. Russian.
Methods of Clinical Laboratory Tests: in 3 vol. Vol. 3. Clinical Microbiology / Menshchikov VV, ed. Moscow, 2009. Russian.
Rychkova LV, Ajurova ZG, Pogodina AV. Obesity and associated risk factors in adolescents in rural areas of Buryatia, Russia. Ozhirenie Metabolizm. 2018;15(3):42-48. Russian.
Abenavoli L, Scarpellini E, Colica C, Boccuto L, Salehi B, Sharifi-Rad J, Aiello V, Romano B, De Lorenzo A, Izzo AA, Capasso R. Gut microbiota and obesity: a role for probiotics. Nutrients. 2019;11(11). pii: E2690. doi: https://doi.org/10.3390/nu11112690
Castaner O, Goday A, Park YM, Lee SH, Magkos F, Shiow STE, Schröder H. The gut microbiome profile in obesity: a systematic review. Int. J. Endocrinol. 2018;2018:4095789. doi: https://doi.org/10.1155/2018/4095789
Chassard C, Dapoigny M, Scott KP, Crouzet L, Del’homme C, Marquet P, Martin JC, Pickering G, Ardid D, Eschalier A, Dubray C, Flint HJ, Bernalier-Donadille A. Functional dysbiosis within the gut microbiota of patients with constipated-irritable bowel syndrome. Aliment. Pharmacol. Ther. 2012;35(7):828-838.
Gagliardi A, Totino V, Cacciotti F, Iebba V, Neroni B, Bonfiglio G, Trancassini M, Passariello C, Pantanella F, Schippa S. Rebuilding the gut microbiota ecosystem. Int. J. Environ. Res. Public Health. 2018;15(8). pii: E1679. doi: https://doi.org/10.3390/ijerph15081679
Ho JT, Chan GC, Li JC. Systemic effects of gut microbiota and its relationship with disease and modulation. BMC Immunol. 2015;16:21. doi: https://doi.org/10.1186/s12865-015-0083-2
Hollister EB, Riehle K, Luna RA, Weidler EM, Rubio-Gonzales M, Mistretta TA, Raza S, Doddapaneni HV, Metcalf GA, Muzny DM, Gibbs RA, Petrosino JF, Shulman RJ, Versalovic J. Structure and function of the healthy pre-adolescent pediatric gut microbiome. Microbiome. 2015;3:36. doi: https://doi.org/10.1186/s40168-015-0101-x
Ley RE, Lozupone CA, Hamady M, Knight R, Gordon JI. Worlds within worlds: evolution of the vertebrate gut microbiota. Nat. Rev. Microbiol. 2008;6(10):776-788.
McBurney MI, Davis C, Fraser CM, Schneeman BO, Huttenhower C, Verbeke K, Walter J, Latulippe ME. Establishing what constitutes a healthy human gut microbiome: state of the science, regulatory considerations, and future directions. J. Nutr. 2019;149(11):1882-1895.
Nouvenne A, Ticinesi A, Tana C, Prati B, Catania P, Miraglia C, De’ Angelis GL, Di Mario F, Meschi T. Digestive disorders and Intestinal microbiota. Acta Biomed. 2018;89(9-S):47-51.
O’Toole PW, Paoli M. The contribution of microbial biotechnology to sustainable development goals: microbiome therapies. Microb. Biotechnol. 2017;10(5):1066-1069.
Parte AC. LPSN — List of Prokaryotic names with standing in nomenclature (bacterio.net), 20 years on. Int. J. Syst. Evol. Microbiol. 2018;68(6):1825-1829.
Author information
Authors and Affiliations
Corresponding author
Additional information
Translated from Byulleten’ Eksperimental’noi Biologii i Meditsiny, Vol. 170, No. 9, pp. 312-317, September, 2020
Rights and permissions
About this article
Cite this article
Grigorova, E.V., Belkova, N.L., Nemchenko, U.M. et al. Metasequencing of V3-V4 Variable Regions of 16S rRNA Gene in Opportunistic Microbiota and Gut Biocenosis in Obese Adolescents. Bull Exp Biol Med 170, 321–325 (2021). https://doi.org/10.1007/s10517-021-05060-3
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10517-021-05060-3