Abstract
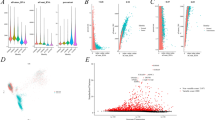
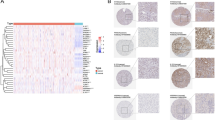
Kidney renal clear cell carcinoma (KIRC) is the most common histopathologic type of renal cell carcinoma. PANoptosis, a cell death pathway that involves an interplay between pyroptosis, apoptosis and necroptosis, is associated with cancer immunity and development. However, the prognostic significance of PANoptosis in KIRC remains unclear. RNA-sequencing expression and mutational profiles from 532 KIRC samples and 72 normal samples with sufficient clinical data were retrieved from the Cancer Genome Atlas (TCGA) database. A prognostic model was constructed using differentially expressed genes (DEGs) related to PANoptosis in the TCGA cohort and was validated in a Gene Expression Omnibus (GEO) cohorts. Incorporating various clinical features, the risk model remained an independent prognostic factor in multivariate analysis, and it demonstrated superior performance compared to unsupervised clustering of the 21 PANoptosis-related genes alone. Further mutational analysis showed fewer VHL and more BAP1 alterations in the high-risk group, with alterations in both genes also associated with patient prognosis. The high-risk group was characterized by an unfavorable immune microenvironment, marked by reduced levels of CD4 + T cells and natural killer cells, but increased M2 macrophages and regulatory T cells. Finally, the risk model was predictive of response to immune checkpoint blockade, as well as sensitivity to sunitinib and paclitaxel. The PANoptosis-related risk model developed in this study enables accurate prognostic prediction in KIRC patients. Its associations with the tumor immune microenvironment and drug efficacy may offer potential therapeutic targets and inform clinical decisions.






Similar content being viewed by others
Data availability
The data that support the findings of this study are openly available in TCGA database at https://portal.gdc.cancer.gov/, GEO database at https://www.ncbi.nlm.nih.gov/geo/, or are available from the corresponding author upon reasonable request.
References
Vuong L, Kotecha RR, Voss MH, Hakimi AA (2019) Tumor microenvironment dynamics in clear-cell renal cell carcinoma. Cancer Discov 9(10):1349–1357. https://doi.org/10.1158/2159-8290.CD-19-0499
Choueiri TK, Motzer RJ (2017) Systemic therapy for metastatic renal-cell carcinoma. N Engl J Med 376(4):354–366. https://doi.org/10.1056/NEJMra1601333
Leibovich BC, Lohse CM, Crispen PL, Boorjian SA, Thompson RH, Blute ML, Cheville JC (2010) Histological subtype is an independent predictor of outcome for patients with renal cell carcinoma. J Urol 183(4):1309–1315. https://doi.org/10.1016/j.juro.2009.12.035
Shuch B, Amin A, Armstrong AJ, Eble JN, Ficarra V, Lopez-Beltran A, Martignoni G, Rini BI, Kutikov A (2015) Understanding pathologic variants of renal cell carcinoma: distilling therapeutic opportunities from biologic complexity. Eur Urol 67(1):85–97. https://doi.org/10.1016/j.eururo.2014.04.029
Shao N, Tang H, Mi Y, Zhu Y, Wan F, Ye D (2020) A novel gene signature to predict immune infiltration and outcome in patients with prostate cancer. Oncoimmunology 9(1):1762473. https://doi.org/10.1080/2162402X.2020.1762473
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12(5):453–457. https://doi.org/10.1038/nmeth.3337
Janiszewska AD, Poletajew S, Wasiutyński A (2013) Spontaneous regression of renal cell carcinoma. Contemp Oncol (Pozn) 17(2):123–127. https://doi.org/10.5114/wo.2013.34613
Inman BA, Harrison MR, George DJ (2013) Novel immunotherapeutic strategies in development for renal cell carcinoma. Eur Urol 63(5):881–889. https://doi.org/10.1016/j.eururo.2012.10.006
Desbois M, Udyavar AR, Ryner L, Kozlowski C, Guan Y, Dürrbaum M, Lu S, Fortin JP, Koeppen H, Ziai J, Chang CW, Keerthivasan S, Plante M, Bourgon R, Bais C, Hegde P, Daemen A, Turley S, Wang Y (2020) Integrated digital pathology and transcriptome analysis identifies molecular mediators of T-cell exclusion in ovarian cancer. Nat Commun 11(1):5583. https://doi.org/10.1038/s41467-020-19408-2
Bi K, He MX, Bakouny Z, Kanodia A, Napolitano S, Wu J, Grimaldi G, Braun DA, Cuoco MS, Mayorga A, DelloStritto L, Bouchard G, Steinharter J, Tewari AK, Vokes NI, Shannon E, Sun M, Park J, Chang SL, McGregor BA, Haq R, Denize T, Signoretti S, Guerriero JL, Vigneau S, Rozenblatt-Rosen O, Rotem A, Regev A, Choueiri TK, Van Allen EM (2021) Tumor and immune reprogramming during immunotherapy in advanced renal cell carcinoma. Cancer Cell 39(5):649-661.e5. https://doi.org/10.1016/j.ccell.2021.02.015
Motzer RJ, Rini BI, McDermott DF, Redman BG, Kuzel TM, Harrison MR, Vaishampayan UN, Drabkin HA, George S, Logan TF, Margolin KA, Plimack ER, Lambert AM, Waxman IM, Hammers HJ (2015) Nivolumab for metastatic renal cell carcinoma: results of a randomized phase II trial. J Clin Oncol 33(13):1430–1437. https://doi.org/10.1200/JCO.2014.59.0703
Bertheloot D, Latz E, Franklin BS (2021) Necroptosis, pyroptosis and apoptosis: an intricate game of cell death. Cell Mol Immunol 18(5):1106–1121. https://doi.org/10.1038/s41423-020-00630-3
Hanahan D (2022) Hallmarks of cancer: new dimensions. Cancer Discov 12(1):31–46. https://doi.org/10.1158/2159-8290.CD-21-1059
Galluzzi L, Vitale I, Aaronson SA, Abrams JM, Adam D, Agostinis P, Alnemri ES, Altucci L, Amelio I, Andrews DW, Annicchiarico-Petruzzelli M, Antonov AV, Arama E, Baehrecke EH, Barlev NA, Bazan NG, Bernassola F, Bertrand MJM, Bianchi K, Blagosklonny MV, Blomgren K, Borner C, Boya P, Brenner C, Campanella M, Candi E, Carmona-Gutierrez D, Cecconi F, Chan FK, Chandel NS, Cheng EH, Chipuk JE, Cidlowski JA, Ciechanover A, Cohen GM, Conrad M, Cubillos-Ruiz JR, Czabotar PE, D'Angiolella V, Dawson TM, Dawson VL, De Laurenzi V, De Maria R, Debatin KM, DeBerardinis RJ, Deshmukh M, Di Daniele N, Di Virgilio F, Dixit VM, Dixon SJ, Duckett CS, Dynlacht BD, El-Deiry WS, Elrod JW, Fimia GM, Fulda S, García-Sáez AJ, Garg AD, Garrido C, Gavathiotis E, Golstein P, Gottlieb E, Green DR, Greene LA, Gronemeyer H, Gross A, Hajnoczky G, Hardwick JM, Harris IS, Hengartner MO, Hetz C, Ichijo H, Jäättelä M, Joseph B, Jost PJ, Juin PP, Kaiser WJ, Karin M, Kaufmann T, Kepp O, Kimchi A, Kitsis RN, Klionsky DJ, Knight RA, Kumar S, Lee SW, Lemasters JJ, Levine B, Linkermann A, Lipton SA, Lockshin RA, López-Otín C, Lowe SW, Luedde T, Lugli E, MacFarlane M, Madeo F, Malewicz M, Malorni W, Manic G, Marine JC, Martin SJ, Martinou JC, Medema JP, Mehlen P, Meier P, Melino S, Miao EA, Molkentin JD, Moll UM, Muñoz-Pinedo C, Nagata S, Nuñez G, Oberst A, Oren M, Overholtzer M, Pagano M, Panaretakis T, Pasparakis M, Penninger JM, Pereira DM, Pervaiz S, Peter ME, Piacentini M, Pinton P, Prehn JHM, Puthalakath H, Rabinovich GA, Rehm M, Rizzuto R, Rodrigues CMP, Rubinsztein DC, Rudel T, Ryan KM, Sayan E, Scorrano L, Shao F, Shi Y, Silke J, Simon HU, Sistigu A, Stockwell BR, Strasser A, Szabadkai G, Tait SWG, Tang D, Tavernarakis N, Thorburn A, Tsujimoto Y, Turk B, Vanden Berghe T, Vandenabeele P, Vander Heiden MG, Villunger A, Virgin HW, Vousden KH, Vucic D, Wagner EF, Walczak H, Wallach D, Wang Y, Wells JA, Wood W, Yuan J, Zakeri Z, Zhivotovsky B, Zitvogel L, Melino G, Kroemer G (2018) Molecular mechanisms of cell death: recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ 25(3):486–541. https://doi.org/10.1038/s41418-017-0012-4
Kesavardhana S, Malireddi RKS, Kanneganti TD (2020) Caspases in cell death, inflammation, and pyroptosis. Annu Rev Immunol 38:567–595. https://doi.org/10.1146/annurev-immunol-073119-095439
Malireddi RKS, Kesavardhana S, Kanneganti TD (2019) ZBP1 and TAK1: master regulators of NLRP3 inflammasome/pyroptosis, apoptosis, and necroptosis (PAN-optosis). Front Cell Infect Microbiol 9:406. https://doi.org/10.3389/fcimb.2019.00406
Samir P, Malireddi RKS, Kanneganti TD (2020) The PANoptosome: a deadly protein complex driving pyroptosis, apoptosis, and necroptosis (PANoptosis). Front Cell Infect Microbiol 10:238. https://doi.org/10.3389/fcimb.2020.00238
Fritsch M, Günther SD, Schwarzer R, Albert MC, Schorn F, Werthenbach JP, Schiffmann LM, Stair N, Stocks H, Seeger JM, Lamkanfi M, Krönke M, Pasparakis M, Kashkar H (2019) Caspase-8 is the molecular switch for apoptosis, necroptosis and pyroptosis. Nature 575(7784):683–687. https://doi.org/10.1038/s41586-019-1770-6
Jiang M, Qi L, Li L, Wu Y, Song D, Li Y (2021) Caspase-8: a key protein of cross-talk signal way in “PANoptosis” in cancer. Int J Cancer 149(7):1408–1420. https://doi.org/10.1002/ijc.33698
Karki R, Sundaram B, Sharma BR, Lee S, Malireddi RKS, Nguyen LN, Christgen S, Zheng M, Wang Y, Samir P, Neale G, Vogel P, Kanneganti TD (2021) ADAR1 restricts ZBP1-mediated immune response and PANoptosis to promote tumorigenesis. Cell Rep 37(3):109858. https://doi.org/10.1016/j.celrep.2021.109858
Malireddi RKS, Karki R, Sundaram B, Kancharana B, Lee S, Samir P, Kanneganti TD (2021) Inflammatory cell death, PANoptosis, mediated by cytokines in diverse cancer lineages inhibits tumor growth. Immunohorizons 5(7):568–580. https://doi.org/10.4049/immunohorizons.2100059
Karki R, Sharma BR, Lee E, Banoth B, Malireddi RKS, Samir P, Tuladhar S, Mummareddy H, Burton AR, Vogel P, Kanneganti TD (2020) Interferon regulatory factor 1 regulates PANoptosis to prevent colorectal cancer. JCI Insight 5(12):e136720. https://doi.org/10.1172/jci.insight.136720
Wang X, Sun R, Chan S, Meng L, Xu Y, Zuo X, Wang Z, Hu X, Han Q, Dai L, Bai T, Yu Z, Wang M, Yang W, Zhang H, Chen W (2022) PANoptosis-based molecular clustering and prognostic signature predicts patient survival and immune landscape in colon cancer. Front Genet 13:955355. https://doi.org/10.3389/fgene.2022.955355
Chen G, He Z, Jiang W, Li L, Luo B, Wang X, Zheng X (2022) Construction of a machine learning-based artificial neural network for discriminating PANoptosis related subgroups to predict prognosis in low-grade gliomas. Sci Rep 12(1):22119. https://doi.org/10.1038/s41598-022-26389-3
Pan H, Pan J, Li P, Gao J (2022) Characterization of PANoptosis patterns predicts survival and immunotherapy response in gastric cancer. Clin Immunol 238:109019. https://doi.org/10.1016/j.clim.2022.109019
Geeleher P, Cox N, Huang RS (2014) pRRophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS ONE 9(9):e107468. https://doi.org/10.1371/journal.pone.0107468
Zheng M, Kanneganti TD (2020) The regulation of the ZBP1-NLRP3 inflammasome and its implications in pyroptosis, apoptosis, and necroptosis (PANoptosis). Immunol Rev 297(1):26–38. https://doi.org/10.1111/imr.12909
Karki R, Sharma BR, Tuladhar S, Williams EP, Zalduondo L, Samir P, Zheng M, Sundaram B, Banoth B, Malireddi RKS, Schreiner P, Neale G, Vogel P, Webby R, Jonsson CB, Kanneganti TD (2021) Synergism of TNF-α and IFN-γ triggers inflammatory cell death, tissue damage, and mortality in SARS-CoV-2 infection and cytokine shock syndromes. Cell 184(1):149-168.e17. https://doi.org/10.1016/j.cell.2020.11.025
Chen W, Lin W, Wu L, Xu A, Liu C, Huang P (2022) A novel prognostic predictor of immune microenvironment and therapeutic response in kidney renal clear cell carcinoma based on necroptosis-related gene signature. Int J Med Sci 19(2):377–392. https://doi.org/10.7150/ijms.69060
Li D, Liu S, Xu J, Chen L, Xu C, Chen F, Xu Z, Zhang Y, Xia S, Shao Y, Wang Y. Ferroptosis-related gene CHAC1 is a valid indicator for the poor prognosis of kidney renal clear cell carcinoma. J Cell Mol Med. 2021;25(7):3610–3621. https://doi.org/10.1111/jcmm.16458. Epub 2021 Mar 16. Erratum in: J Cell Mol Med. 2022;26(18):4874–4875
Xie L, Li H, Zhang L, Ma X, Dang Y, Guo J, Liu J, Ge L, Nan F, Dong H, Yan Z, Guo X (2020) Autophagy-related gene P4HB: a novel diagnosis and prognosis marker for kidney renal clear cell carcinoma. Aging (Albany NY) 12(2):1828–1842. https://doi.org/10.18632/aging.102715
Sun Z, Tao W, Guo X, Jing C, Zhang M, Wang Z, Kong F, Suo N, Jiang S, Wang H (2022) Construction of a lactate-related prognostic signature for predicting prognosis, tumor microenvironment, and immune response in kidney renal clear cell carcinoma. Front Immunol 13:818984. https://doi.org/10.3389/fimmu.2022.818984
Zamyatina A, Heine H (2020) Lipopolysaccharide recognition in the crossroads of TLR4 and caspase-4/11 mediated inflammatory pathways. Front Immunol 11:585146. https://doi.org/10.3389/fimmu.2020.585146. (Erratum in: Front Immunol 2021;12:649442)
Li T, Liu N, Zhang G, Chen M (2022) CASP4 and CASP8 as newly defined autophagy-pyroptosis-related genes associated with clinical and prognostic features of renal cell carcinoma. J Cancer Res Ther 18(7):1952–1960. https://doi.org/10.4103/jcrt.jcrt_126_22
Liao G, Lv J, Ji A, Meng S, Chen C (2021) TLR3 serves as a prognostic biomarker and associates with immune infiltration in the renal clear cell carcinoma microenvironment. J Oncol 2021:3336770. https://doi.org/10.1155/2021/3336770
Wang L, Zhu Y, Ren Z, Sun W, Wang Z, Zi T, Li H, Zhao Y, Qin X, Gao D, Zhang L, He Z, Le W, Wu Q, Wu G (2023) An immunogenic cell death-related classification predicts prognosis and response to immunotherapy in kidney renal clear cell carcinoma. Front Oncol 13:1147805. https://doi.org/10.3389/fonc.2023.1147805
Xu J, Guo Y (2021) LY96 Upregulation Promotes Kidney Renal Clear Cell Carcinoma (KIRC). Research Square 2021. https://doi.org/10.21203/rs.3.rs-154342/v1
Lee N, Zakka LR, Mihm MC Jr, Schatton T (2016) Tumour-infiltrating lymphocytes in melanoma prognosis and cancer immunotherapy. Pathology 48(2):177–187. https://doi.org/10.1016/j.pathol.2015.12.006
Jochems C, Schlom J (2011) Tumor-infiltrating immune cells and prognosis: the potential link between conventional cancer therapy and immunity. Exp Biol Med (Maywood) 236(5):567–579. https://doi.org/10.1258/ebm.2011.011007
Acknowledgements
We are grateful to Sangerbox (http://vip.sangerbox.com) and Nanjing Geneseeq Technology Inc. for providing technical support of this manuscript.
Funding
This research is supported by the National Natural Science Foundation of China (82274607), the Tianjin Health and Family Planning Commission program (No. 2017166) and Beijing Xisike Clinical Oncology Research Foundation (Y-HS202102-0120, Y-HS202202-0005).
Author information
Authors and Affiliations
Contributions
ZJ and JW made substantial contributions to the conception or design of the work; JW and CD made substantial contributions to the acquisition of the work; MZ,Y L,FL,YZ and JL made substantial contributions to the creation of new software used in the work; ZJ, ZP and YY drafted the work or revised it critically for important intellectual content; ZP and YY approved the version to be published; ZP and YY agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Corresponding authors
Ethics declarations
Competing interest
The authors declare no competing interest.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jiang, Z., Wang, J., Dao, C. et al. Utilizing a novel model of PANoptosis-related genes for enhanced prognosis and immune status prediction in kidney renal clear cell carcinoma. Apoptosis 29, 681–692 (2024). https://doi.org/10.1007/s10495-023-01932-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10495-023-01932-3