Abstract
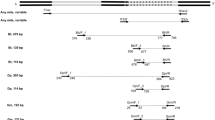
Mites are known causes of allergic diseases. Currently, identification of mites based on morphology is difficult if only one mite is isolated from a (dust) sample, or when only one gender is found, or when the specimen is not intact especially with the loss of the legs. The purpose of this study was to use polymerase chain reaction–restriction fragment length polymorphism (PCR–RFLP) of the ITS2 gene, to complement the morphological data for the identification of mites to the species level. For this, six species were cultured: Dermatophagoides pteronyssinus, D. farinae, Blomia tropicalis, Tyrophagus putrescentiae, Aleuroglyphus ovatus and Glycycometus malaysiensis. Genomic DNA of the mites was extracted, quantified, amplified and digested individually with restriction enzymes. Hinf I and Ple I differentiated the restriction patterns of D. pteronyssinus and D. farinae. Bfa I and Alu I enzymes differentiated B. tropicalis and G. malaysiensis. Ple I enzyme was useful for the differentiation between T. putrescentiae and A. ovatus. Bfa I was useful for the differentiation of G. malaysiensis from the rest of the species. In conclusion, different species of mites can be differentiated using PCR–RFLP of ITS2 region. With the established PCR–RFLP method in this study, identification of these mites to the species level is possible even if complete and intact adult specimens of both sexes are not available. As no study to date has reported PCR–RFLP method for the identification of domestic mites, the established method should be validated for the identification of other species of mites that were not included in this study.







Similar content being viewed by others
References
Anderson DL, Morgan MJ (2007) Genetic and morphological variation of bee parasitic Tropilaelaps mites (Acari: Laelapidae): new and re-defined species. Exp Appl Acarol 43:1–24
Arlian LG, Morgan MS (2003) Biology, ecology and prevalence of dust mites. Immunol Allergy Clin North Am 23:443–468
Ben-David T, Melamed S, Gerson U (2007) ITS2 sequences as barcodes for identifying and analyzing spider mites (Acari: Tetranychidae). Exp Appl Acarol 41:169–181
Bronswijk JE, de Cock W, Oshima S (1974) The genus Blomia oudemans (Acari: Glycyphagidae). Description of Blomia tropicalis from house dust in tropical and sub-tropical regions. Acarología 15:477–489
Chew FT, Zhang L, Ho TM, Lee BW (1999) House dust mite fauna of tropical Singapore. Clin Exp Allergy 29:201–206
Colloff MJ (2009) Identification and taxonomy, classification and phylogeny. In: Findlay A (ed) Dust mites, 1st edn. CSIRO Publishing, Australia, pp 1–44
Colloff MJ, Spieksma FTM (1992) Pictorial keys for the identification of domestic mites. Clin Exp Allergy 22:823–830
Cruickshank RH (2002) Molecular markers for the phylogenetics of mites and ticks. System Appl Acarol 7:3–14
Fenton B, Birch ANE, Malloch G, Lanham PG, Brennan RM (2000) Gall mite molecular phylogeny and its relationship to the evolution of plant host specificity. Exp Appl Acarol 24:831–861
Hage-Hamsten M, Johansson SGO (1992) Storage mites. Exp Appl Acarol 16:117–128
Jeyaprakash A, Hoy MA (2010) A DNA extraction procedure that allows mite specimens to be slide mounted: phytoseiid species evaluated as a model. Exp Appl Acarol 40:241–247
Konakandla B, Park Y, Margolies D (2006) Whole genome amplification of Chelex-extracted DNA from a single mite: a method for studying genetics of the predatory mite Phytoseiulus persimilis. Exp Appl Acarol 40:241–247
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Li-Mattila TY, Paavanen-Huhtala S, Fenton B, Tuovinen T (2000) Species and strain identification by RAPD-PCR and ITS sequences. Exp Appl Acarol 24:863–880
Mahani MK, Inomata N, Saboori A, Tabatabaei BES, Ishiyama H, Ariana A (2009) Genetic variation in populations of Allothrombium pulvinum (Acari: Trombidiidae) from Northern Iran revealed by mitochondrial COX I and nuclear rDNA ITS2 sequences. Exp Appl Acarol 48:273–289
Mariana A, Ho TM, Sofian-Azirun M, Wong AL (2000) House dust mite fauna in the Klang Valley, Malaysia. Southeast Asian J Trop Med Publ Health 31(4):712–721
Mtambo J, Bortel WV, Madder M, Roelants P, Backeljau T (2006) Comparison of preservation methods of Rhipicephalus appendiculatus (Acari: Ixodidae) for reliable DNA amplification by PCR. Exp Appl Acarol 38:189–199
Mtambo J, Madder M, Bortel WV, Berkvens D, Backeljau T (2007a) Rhipicephalus appendiculatus and R. zambeziensis (Acari: Ixodidae) from Zambia: a molecular reassessment of their species status and identification. Exp Appl Acarol 41:115–128
Mtambo J, Madder M, Bortel WV, Chaka G, Berkvens D, Backeljau T (2007b) Further evidence for geographic differentiation in R. appendiculatus (Acari: Ixodidae) from Eastern and Southern provinces of Zambia. Exp Appl Acarol 41:129–138
Nadchatram M (2005) House dust mites, our intimate associates. Trop Biomed 22(1):23–37
Navajas M (1998) Host plant associations in the spider mite Tetranychus urticae (Acari: Tetranychidae): insights from molecular phylogeography. Exp Appl Acarol 22:201–214
Navajas M, Lagnel J, Fauvel G, de Moraes G (1999) Sequence variation of ribosomal internal transcribed spacers (ITS) in commercially important Phytoseiidae mites. Exp Appl Acarol 23:851–859
Navajas M, Tsagkarakov A, Lagnel J, Perrot-Minnot MJ (2000) Genetic differentiation in Tetranychus urticae (Acari: Tetranychidae): polymorphism, host races or sibling species? Exp Appl Acarol 24:365–376
Noge K, Mori N, Tanaka C, Nishida R, Tsuda M, Kuwahara Y (2005) Identification of astigmatid mites using the second internal transcribed spacer (ITS2) region and its application for phylogenetic study. Exp Appl Acarol 35:29–46
Palyvos NE, Emmanouel NG, Saitanis CJ (2008) Mites associated with stored products in Greece. Exp Appl Acarol 44:213–226
Potenza L, Cafiero MA, Camarda A, Salandra GL, Cucchiarini L, Dachà M (2009) Characterization of Dermanyssus gallinae (Acarina: Dermanissydae) by sequence analysis of the ribosomal internal transcribed spacer regions. Vet Res Commun 33:611–618
Sambrook J, Russel DW (2001) Gel electrophoresis of DNA and pulsed-field agarose gel electrophoresis. In: William D (ed) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press. Cold Spring Harbor, pp 5.4–5.17
Suarez-Martinez EB, Montealegre F, Sierra-Montes JM, Herrera RJ (2005) Molecular identification of pathogenic house dust mites using 12S rRNA sequences. Electrophoresis 26:2927–2934
Tsai JJ, Wu HH, Shen HD, Hsu EL, Wang SR (1998) Sensitization to Blomia tropicalis among asthmatic patients in Taiwan. Int Arch Allergy Immunol 115:144–149
Vargas M, García-Varela M, Laclette JP, Pérez TM (2005) Application of ITS 2 sequences as markers for identification and phylogenetic inference within the genus Geomylichus (Acari: Listrophoridae). Exp Appl Acarol 35:223–238
Zahler M, Gothe R (1997) Evidence for the reproductive isolation of Dermacentor marginatus and Dermacentor reticulatus (Acari: Ixodidae) ticks based on cross-breeding, morphology and molecular studies. Exp Appl Acarol 21:685–696
Acknowledgments
We are grateful to the Institute for Medical Research (IMR), Kuala Lumpur, Malaysia for providing the dust mites. This research was supported by grants from Ministry of Science, Technology and Innovation Malaysia (02-02-09-SF0007) and International Medical University [Grant number: BMS I02-2009(05)].
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wong, S.F., Chong, A.L., Mak, J.W. et al. Molecular identification of house dust mites and storage mites. Exp Appl Acarol 55, 123–133 (2011). https://doi.org/10.1007/s10493-011-9460-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10493-011-9460-6