Abstract
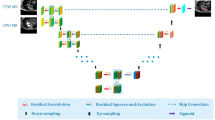
Multi-modal learning of unpaired anatomical and functional modalities is challenging because the large discrepancies between feature representations, and the absence of registration, hinder deeper cross-modality information extraction. To address this issue, we propose a novel unpaired multi-modal learning framework for lung tumor segmentation in anatomical and functional images (i.e., T2w-MRI and DWI-MRI). The framework consists of two models: a multi-modal segmentation network (Segmentor) and an adversarial learning-based structure adaptation (SA) model. The Segmentor contains a modality-specific encoder for intensity offset reduction and a shared decoder for cross-modality information fusion. The SA model helps the Segmentor obtain high-level features with similar distributions from different modalities through adversarial training. To address the modality imbalance problem caused by the large gap in the number of training samples and image characteristics, we introduce a harmonic constraint term derived from two weighted dice losses to maintain the balance of multi-modal training. Experimental results based on a clinical dataset of 326 T2w-MRI and 112 DWI-MRI scans show that the proposed framework improves segmentation performance compared with several state-of-the-art unpaired multi-modal segmentation methods.







Similar content being viewed by others
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global cancer statistics 2020: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries CA: a cancer journal for clinicians
Sim A. J., Kaza E, Singer L, Rosenberg SA (2020) A review of the role of mri in diagnosis and treatment of early stage lung cancer. Clinical and Translational Radiation Oncology 24:16–22
Yin Y, Sedlaczek O, Müller B, Warth A, González-Vallinas M, Lahrmann B, Grabe N, Kauczor H, Breuhahn K, Vignon-Clementel IE, Drasdo D (2018) Tumor cell load and heterogeneity estimation from diffusion-weighted MRI calibrated with histological data: an example from lung cancer. IEEE Trans Med Imaging 37(1):35–46. https://doi.org/10.1109/TMI.2017.2698525
Zhou T, Ruan S, Canu S (2019) A Review: Deep learning for medical image segmentation using multi-modality fusion. Array 100004:3–4. https://doi.org/10.1016/j.array.2019.100004
Guo Z, Li X, Huang H, Guo N (2019) Li Q:Deep learning-based image segmentation on multimodal medical imaging. IEEE Transactions on Radiation and Plasma Medical Sciences 3(2):162–169. https://doi.org/10.1109/TRPMS.2018.2890359
Ghaffari M, Sowmya A, Oliver R (2020) Automated brain tumor segmentation using multimodal brain scans: A survey based on models submitted to the brats 2012–2018 Challenges. IEEE Rev Biomed Eng 13:156–168. https://doi.org/10.1109/RBME.2019.2946868https://doi.org/10.1109/RBME.2019.2946868
Kumar A, Fulham M, Feng D, Kim J (2020) Co-learning feature fusion maps from PET-CT images of lung cancer. IEEE Trans Med Imaging 39(1):204–217. https://doi.org/10.1109/TMI.2019.2923601https://doi.org/10.1109/TMI.2019.2923601
Huo Y, Xu Z, Bao S, Bermudez C, Moon H, Parvathaneni P, Moyo TK, Savona MR, Assad A, Abramson RG, Landman BA (2019) Splenomegaly segmentation on multi-modal MRI using deep convolutional networks. IEEE Trans Med Imaging 38(5):1185–1196. https://doi.org/10.1109/TMI.2018.2881110
Dolz J, Gopinath K, Yuan J, Lombaert H, Desrosiers C, Ben Ayed I (2019) Hyperdense-net: A hyper-densely connected CNN for multi-modal image segmentation. IEEE Trans Med Imaging 38 (5):1116–1126. https://doi.org/10.1109/TMI.2018.2878669
Valindria V, Pawlowski N, Rajchl M, Lavdas I, Aboagye EO, Rockall AG, Rueckert D, Glocker B (2018) Multi-modal learning from unpaired images: Application to multi-organ segmentation in ct and mri. Proceedings IEEE Winter Conference Application Computers Vision (WACV), 547–556
Dou Q, Liu Q, Heng PA, Glocker B (2020) Unpaired multi-modal segmentation via knowledge distillation. IEEE Trans Med Imaging 39(7):2415–2425. https://doi.org/10.1109/tmi.2019.2963882
Wan Q, Deng Y, Lei Q, Bao Y, Wang Y, Zhou J, Zou Q, Li X (2019) Differentiating between malignant and benign solid solitary pulmonary lesions: are intravoxel incoherent motion and diffusion kurtosis imaging superior to conventional diffusion-weighted imaging? Eur Radiol 29 (3):1607–1615. https://doi.org/10.1007/s00330-018-5714-6
Nie D, Wang L, Gao Y, Shen D (2016) Fully convolutional networks for multi-modality isointense infant brain image segmentation. In: Proceedings IEEE 13th international symposium. Biomedical imaging (ISBI), pp 1342–1345. https://doi.org/10.1109/ISBI.2016.7493515
Tang P, Yan X, Nan Y, Xiang S, Krammer S, Lasser T (2022) Fusionm4net: A multi-stage multi-modal learning algorithm for multi-label skin lesion classification. Med Image Anal 76:102307. https://doi.org/10.1016/j.media.2021.102307
Tzeng E, Hoffman J, Saenko K, Darrell T (2017) Adversarial discriminative domain adaptation. In: Proceedings IEEE conference Computer vision and pattern recognition (CVPR), pp 2962–2971. https://doi.org/10.1109/CVPR.2017.316
Chadha A, Andreopoulos Y (2020) Improved techniques for adversarial discriminative domain adaptation. IEEE Trans Image Process 29:2622–2637. https://doi.org/10.1109/TIP.2019.2950768
Javanmardi M, Tasdizen T (2018) Domain adaptation for biomedical image segmentation using adversarial training. In: 2018 IEEE 15Th international symposium on biomedical imaging (ISBI 2018), pp 554–558. https://doi.org/10.1109/ISBI.2018.8363637
Guan H, Liu M (2022) Domain adaptation for medical image analysis: A survey. IEEE Trans Biomed Eng 69(3):1173–1185. https://doi.org/10.1109/TBME.2021.3117407
Dou Q, Ouyang C, Chen C, Chen H, Glocker B, Zhuang X, Heng P-A (2019) Pnp-adanet: Plug-and-play adversarial domain adaptation network at unpaired cross-modality cardiac segmentation. IEEE Access 7:99065–99076. https://doi.org/10.1109/ACCESS.2019.2929258https://doi.org/10.1109/ACCESS.2019.2929258
Tsai Y-H, Sohn K, Schulter S, Chandraker M (2019) Domain adaptation for structured output via discriminative patch representations. In: Proceedings IEEE/CVF international conference Computer vision (ICCV), pp 1456–1465. https://doi.org/10.1109/ICCV.2019.00154
Tsai Y-H, Hung W-C, Schulter S, Sohn K, Yang M-H, Chandraker M (2018) Learning to adapt structured output space for semantic segmentation. In: Proceedings IEEE/CVF conference Computer vision and pattern recognition, pp 7472–7481. https://doi.org/10.1109/CVPR.2018.00780
Johnson JM, Khoshgoftaar TM (2019) Survey on deep learning with class imbalance. Journal of Big Data 6(1):27
Salehi SS, Erdogmus D, Gholipour A (2017) Tversky loss function for image segmentation using 3d fully convolutional deep networks, pp 379–387. https://doi.org/10.1007/978-3-319-67389-9_44
Lin T-Y, Goyal P, Girshick R, He K, Dollár P (2020) Focal loss for dense object detection. IEEE Trans Pattern Anal Mach Intell 42(2):318–327. https://doi.org/10.1109/TPAMI.2018.2858826
Zhou Y, Chen H, Li Y, Liu Q, Xu X, Wang S, Yap P-T, Shen D (2021) Multi-task learning for segmentation and classification of tumors in 3d automated breast ultrasound images. Med Image Anal 101918:70. https://doi.org/10.1016/j.media.2020.101918
Nie D, Gao Y, Wang L, Shen D (2018) Asdnet: Attention based semi-supervised deep networks for medical image segmentation. In: Medical image computing and computer assisted intervention – MICCAI 2018 - 21st international conference, 2018, Proceedings. https://doi.org/10.1007/978-3-030-00937-3_43, pp 370–378
Srivastava N, Salakhutdinov R (2014) Multimodal learning with deep boltzmann machines. J Mach Learn Res 15(1):2949–2980
Pinto A, Pereira S, Meier R, Alves V, Wiest R, Silva CA, Reyes M (2018) Enhancing clinical mri perfusion maps with data-driven maps of complementary nature for lesion outcome prediction. In: Frangi AF, Schnabel JA, Davatzikos C, Alberola-lópez C, Fichtinger G (eds) Medical image computing and computer assisted intervention – MICCAI 2018, pp 107–115. Springer, Cham
Sheng Z, Wang H, Chen G, Zhou B, Sun J (2021) Convolutional residual network to short-term load forecasting. Appl Intell 51:1–15. https://doi.org/10.1007/s10489-020-01932-9
Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, Killeen T, Lin Z, Gimelshein N, Antiga L, Desmaison A, Kopf A, Yang E, DeVito Z, Raison M, Tejani A, Chilamkurthy S, Steiner B, Fang L, Bai J, Chintala S (2019) Pytorch: An imperative style, high-performance deep learning library. In: Advances in neural information processing systems, vol 32. Curran Associates, Inc, Red Hook, pp 8026–8037
Goodfellow I, Pouget-Abadie J, Mirza M, Xu B, Warde-Farley D, Ozair S, Courville A, Bengio Y (2014) Generative adversarial nets. In: Advances in neural information processing systems, pp 2672–2680
Isola P, Zhu J, Zhou T, Efros AA (2017) Image-to-image translation with conditional adversarial networks. In: 2017 IEEE Conference on computer vision and pattern recognition (CVPR), pp 5967–5976. https://doi.org/10.1109/CVPR.2017.632
Milletari F, Navab N, Ahmadi S-A (2016) V-net: Fully convolutional neural networks for volumetric medical image segmentation. 2016 Fourth International Conference on 3D Vision (3DV), 565–571
Rahate A, Walambe R, Ramanna S, Kotecha K (2022) Multimodal co-learning: challenges, applications with datasets, recent advances and future directions. Information Fusion 81:203–239. https://doi.org/10.1016/j.inffus.2021.12.003
Guo W, Wang J, Wang S (2019) Deep multimodal representation learning: A survey. IEEE Access 7:63373–63394. https://doi.org/10.1109/ACCESS.2019.2916887https://doi.org/10.1109/ACCESS.2019.2916887
Acknowledgments
This work was supported by National Science Foundation of China under Grant No. 61771039, No. 62172029.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhou, P., Chen, H., Li, Y. et al. Unpaired multi-modal tumor segmentation with structure adaptation. Appl Intell 53, 3639–3651 (2023). https://doi.org/10.1007/s10489-022-03610-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10489-022-03610-4