Abstract
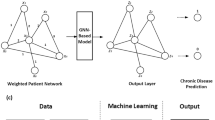
The prediction of chronic diseases and their comorbidities is an essential task in healthcare, aiming to predict patients’ future disease risk based on their previous medical records. The accumulation of administrative data has laid a solid foundation for applying deep learning approaches in healthcare. Existing studies focused on the patients’ characteristics such as gender, age and location to predict the risk of the different diseases. However, there are high dimensional, incomplete and noisy problems in the administrative data. In this research, using administrative health data, we implemented graph theory and content-based recommender system approaches to analyse and predict chronic diseases and their comorbidities. Firstly, we used bipartite graphs to represent the relationships between patients and diseases. Then, we projected this graph to a one-mode graph, namely ‘disease network’. After that, six recommender system models with patient features and network features were trained. The outputs of these models are the severity levels of diseases and the predicted diseases with rank. Finally, we evaluated the performance of these models against the same models without network features. The results demonstrated that the models with network features have lower prediction error and better performances for predicting chronic diseases and their latent comorbidities on large administrative data. Among these models, the graph convolution matrix completion model reveals the least amount of error and the best performance for prediction. Further, using a case study of a specific patient, we demonstrated the application of these models in predictive disease risk analysis. Thus, this study showed the potential application of the recommender system approaches to the health sector utilising administrative claim data, which could significantly contribute to healthcare services and stakeholders.



Similar content being viewed by others
References
World Health Organization (2020) Chronic disease. https://www.who.int/chp/about/integrated_cd/en/, Accessed 8 June 2021
AIHW (2021) Deaths in australia. https://www.aihw.gov.au/reports/life-expectancy-death/deaths-in-australia/contents/leading-causes-of-death, Accessed 26 June 2021
Piane GM, Smith TC (2014) Peer reviewed: Building an evidence base for the co-occurrence of chronic disease and psychiatric distress and impairment. Preventing chronic disease 11
AIHW (2020) Chronic disease. https://www.aihw.gov.au/reports-data/health-conditions-disability-deaths/chronic-disease/overviewhttps://www.aihw.gov.au/reports- https://www.aihw.gov.au/reports-data/health-conditions-disability-deaths/chronic-disease/overviewdata/health-conditions-disability-deaths/chronic-disease/overview, Accessed 26 June 2021
Hossain ME, Khan A, Moni MA, Uddin S (2019) Use of electronic health data for disease prediction. A comprehensive literature review. IEEE/ACM transactions on computational biology and bioinformatics
Shickel B, Tighe PJ, Bihorac A, Rashidi P (2017) Deep ehr: A survey of recent advances in deep learning techniques for electronic health record (ehr) analysis. IEEE J Biomed Health Inform 22(5):1589–1604
Hossain ME, Khan A, Uddin S (2019) Understanding the progression of congestive heart failure of type 2 diabetes patient using disease network and hospital claim data. In: International conference on complex networks and their applications. Springer, pp 774–788
Khan A, Uddin S, Srinivasan U (2019) Chronic disease prediction using administrative data and graph theory: The case of type 2 diabetes. Expert Syst Appl 136:230–241
Uddin S, Khan A, Hossain ME, Moni MA (2019) Comparing different supervised machine learning algorithms for disease prediction. BMC Med Inform Decis Making 19(1):1–16
Lu H, Uddin S, Hajati F, Moni MA, Khushi M (2021) A patient network-based machine learning model for disease prediction: The case of type 2 diabetes mellitus. Appl Intell 1–12
Hossain ME, Uddin S, Khan A (2021) Network analytics and machine learning for predictive risk modelling of cardiovascular disease in patients with type 2 diabetes. Expert Syst Appl 164:113918
Miotto R, Li L, Kidd BA, Dudley JT (2016) Deep patient: an unsupervised representation to predict the future of patients from the electronic health records. Scientif Rep 6(1):1–10
Davis DA, Chawla NV, Blumm N, Christakis N, Barabási AL (2008) Predicting individual disease risk based on medical history. In: Proceedings of the 17th ACM conference on Information and knowledge management, pp 769–778
Davis DA, Chawla NV, Christakis NA, Barabási AL (2010) Time to care: a collaborative engine for practical disease prediction. Data Min Knowl Disc 20(3):388–415
Chao G, Mao C, Wang F, Zhao Y, Luo Y (2018) Supervised nonnegative matrix factorization to predict icu mortality risk. In: 2018 IEEE International Conference on Bioinformatics and Biomedicine (BIBM). IEEE, pp 1189–1194
Xue Y, Klabjan D, Luo Y (2019) Predicting icu readmission using grouped physiological and medication trends. Artif Intell Med 95:27–37
Khan A, Uddin S, Srinivasan U (2018) Comorbidity network for chronic disease: a novel approach to understand type 2 diabetes progression. Int J Med Inform 115:1–9
CBHS Health (2020) CBHS Health Fund. https://www.cbhs.com.au, Accessed 25 Sep 2021
World Health Organization (2020) International classification of diseases (ICD) information sheet. https://www.who.int/classifications/icd/factsheet/en/, Accessed 8 March 2021
The Australian Classification of Health Interventions (2020) ICD-10-AM. http://www.accd.net.au/icd-10-am-achi-acs/, Accessed 8 March 2021
Charlson ME, Pompei P, Ales KL, MacKenzie CR (1987) A new method of classifying prognostic comorbidity in longitudinal studies: development and validation. J Chron Dis 40(5):373–383
Elixhauser A, Steiner C, Harris DR, Coffey RM (1998) Comorbidity measures for use with administrative data. Medical Care 8–27
Asratian AS, Denley TM, Häggkvist R (1998) Bipartite graphs and their applications, vol 131. Cambridge University Press, Cambridge
Freeman LC (1978) Centrality in social networks conceptual clarification. Social networks 1 (3):215–239
Bonacich P (1972) Factoring and weighting approaches to status scores and clique identification. J Math Sociol 2(1):113–120
Shaw ME (1954) Group structure and the behavior of individuals in small groups. J Psychol 38 (1):139–149
Holland PW, Leinhardt S (1971) Transitivity in structural models of small groups. Comparative Group Studies 2(2):107–124
Page L, Brin S, Motwani R, Winograd T (1999) The pagerank citation ranking: Bringing order to the web. Tech. rep., Stanford InfoLab
Ricci F, Rokach L, Shapira B (2011) Introduction to recommender systems handbook. In: Recommender systems handbook. Springer, pp 1–35
Rendle S (2010) Factorization machines. In: 2010 IEEE International Conference on Data Mining, IEEE, pp 995–1000
Cheng HT, Koc L, Harmsen J, Shaked T, Chandra T, Aradhye H, Anderson G, Corrado G, Chai W, Ispir M et al (2016) Wide & deep learning for recommender systems. In: Proceedings of the 1st workshop on deep learning for recommender systems, pp 7–10
Guo H, Tang R, Ye Y, Li Z, He X (2017) Deepfm: a factorization-machine based neural network for ctr prediction. arXiv:170304247
Song W, Shi C, Xiao Z, Duan Z, Xu Y, Zhang M, Tang J (2019) Autoint: Automatic feature interaction learning via self-attentive neural networks. In: Proceedings of the 28th ACM International Conference on Information and Knowledge Management, pp 1161–1170
Zhou G, Zhu X, Song C, Fan Y, Zhu H, Ma X, Yan Y, Jin J, Li H, Gai K (2018) Deep interest network for click-through rate prediction. In: Proceedings of the 24th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, pp 1059–1068
Berg R, Kipf TN, Welling M (2017) Graph convolutional matrix completion. arXiv:170602263
Bastian M, Heymann S, Jacomy M (2009) Gephi: an open source software for exploring and manipulating networks. In: Proceedings of the International AAAI Conference on Web and Social Media, vol 3
MIT (2020) Librecommender library. https://pypi.org/project/LibRecommender/, Accessed 8 June 2021
Paszke A, Gross S, Chintala S, Chanan G, Yang E, DeVito Z, Lin Z, Desmaison A, Antiga L, Lerer A (2017) Automatic differentiation. In: Pytorch
Wang M, Zheng D, Ye Z, Gan Q, Li M, Song X, Zhou J, Ma C, Yu L, Gai Y et al (2019) Deep graph library: A graph-centric, highly-performant package for graph neural networks. arXiv:190901315
Kingma DP, Ba J (2014) Adam: A method for stochastic optimization. arXiv:14126980
Shani G, Gunawardana A (2011) Evaluating recommendation systems. In: Recommender systems handbook, Springer, pp 257–297
Wang R, Chang MC, Radigan M (2020) Modeling latent comorbidity for health risk prediction using graph convolutional network. In: FLAIRS Conference, pp 341–346
Schröder G, Thiele M, Lehner W (2011) Setting goals and choosing metrics for recommender system evaluations. In: UCERSTI2 Workshop at the 5th ACM conference on recommender systems, chicago, USA, vol 23, p 53
Gunawardana A, Shani G (2009) A survey of accuracy evaluation metrics of recommendation tasks. J Mach Learn Res 10(12)
Wang Y, Wang L, Li Y, He D, Chen W, Liu TY (2013) A theoretical analysis of ndcg ranking measures. In: Proceedings of the 26th annual conference on learning theory (COLT 2013), Citeseer, vol 8, p 6
Acknowledgements
We want to acknowledge the contribution of Drs Farshid Hajati, Mohammad Ali Moni and Matloob Khushi for their insightful comments while preparing the revised versions of this article.
Author information
Authors and Affiliations
Contributions
HL: Conceptualisation, Research design, Data analysis and Writing; SU: Conceptualisation, Research design, Writing and Supervision.
Corresponding author
Ethics declarations
Conflict of Interests
The authors declare that they do not have any conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lu, H., Uddin, S. A disease network-based recommender system framework for predictive risk modelling of chronic diseases and their comorbidities. Appl Intell 52, 10330–10340 (2022). https://doi.org/10.1007/s10489-021-02963-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10489-021-02963-6