Abstract
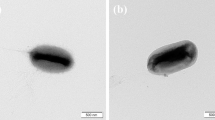
A nitrogen-fixing strain designated SG130T was isolated from paddy soil in Fujian Province, China. Strain SG130T was Gram-staining-negative, rod-shaped, and strictly anaerobic. Strain SG130T showed the highest 16S rRNA gene sequence similarities with the type strains Dendrosporobacter quercicolus DSM 1736T (91.7%), Anaeroarcus burkinensis DSM 6283T (91.0%) and Anaerospora hongkongensis HKU 15T (90.9%). Furthermore, the phylogenetic and phylogenomic analysis also suggested strain SG130T clustered with members of the family Sporomusaceae and was distinguished from other genera within this family. Growth of strain SG130T was observed at 25–45 °C (optimum 30 °C), pH 6.0–9.5 (optimum 7.0) and 0–1% (w/v) NaCl (optimum 0.1%). The quinones were Q-8 and Q-9. The polar lipids were phosphatidylserine (PS), phosphatidylethanolamine (PE), glycolipid (GL), phospholipid (PL) and an unidentified lipid (UL). The major fatty acids (> 10%) were iso-C13:0 3OH (26.6%), iso-C17:1 (15.6%) and iso-C15:1 F (11.4%). The genomic DNA G + C content was 50.7%. The average nucleotide identity (ANI) and digital DNA-DNA hybridization (dDDH) values between strain SG130T and the most closely related type strain D. quercicolus DSM 1736T (ANI 68.0% and dDDH 20.3%) were both below the cut-off level for species delineation. The average amino acid identity (AAI) between strain SG130T and the most closely related type strain D. quercicolus DSM 1736T was 63.2%, which was below the cut-off value for bacterial genus delineation (65%). Strain SG130T possessed core genes (nifHDK) involved in nitrogen fixation, and nitrogenase activity (106.38 μmol C2H4 g−1 protein h−1) was examined using the acetylene reduction assay. Based on the above results, strain SG130T is confirmed to represent a novel genus of the family Sporomusaceae, for which the name Azotosporobacter soli gen. nov., sp. nov. is proposed. The type strain is SG130T (= GDMCC 1.3312T = JCM 35641T).


Similar content being viewed by others
Abbreviations
- ANI:
-
Average Nucleotide Identity
- AAI:
-
Average Amino acid Identity
- dDDH:
-
Digital DNA-DNA Hybridization
- POCP:
-
Percentage Of Conserved Proteins
- PS:
-
Phosphatidylserine
- PE:
-
Phosphatidylethanolamine
- UL:
-
Unidentified Lipid
- GL:
-
Glycolipid
- PL:
-
Phospholipid
References
Besemer J, Lomsadze A, Borodovsky M (2001) GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. implications for finding sequence motifs in regulatory regions. Nucleic Acids Res 29:2607–2618. https://doi.org/10.1093/nar/29.12.2607
Bhattacharjee RB, Singh A, Mukhopadhyay SN (2008) Use of nitrogen-fixing bacteria as biofertiliser for non-legumes: prospects and challenges. Appl Microbiol Biotechnol 80:199–209. https://doi.org/10.1007/s00253-008-1567-2
Blasco F, Iobbi C, Ratouchniak J, Bonnefoy V, Chippaux M (1990) Nitrate reductases of Escherichia coli: sequence of the second nitrate reductase and comparison with that encoded by the narGHJI operon. Mol Gen Genet 222:104–111. https://doi.org/10.1007/BF00283030
Campbell C, Adeolu M, Gupta RS (2015) Genome-based taxonomic framework for the class Negativicutes: division of the class Negativicutes into the orders Selenomonadales emend., Acidaminococcales ord. nov. and Veillonellales ord. nov. Int J Syst Evol Microbiol 65:3203–3215. https://doi.org/10.1099/ijs.0.000347
Chaumeil PA, Mussig AJ, Hugenholtz P, Parks DH (2019) GTDB-Tk: a toolkit to classify genomes with the genome taxonomy database. Bioinformatics 36:1925–1927. https://doi.org/10.1093/bioinformatics/btz848
Choi JK, Shah M, Yee N (2016) Anaerosporomusa subterranea gen. nov., sp. nov., a spore-forming anaerobe belonging to the class Negativicutes isolated from saprolite. Int J Syst Evol Microbiol 66:3848–3854. https://doi.org/10.1099/ijsem.0.001275
Collins MD, Jones D (1980) Lipids in the classification and identification of coryneform bacteria containing peptidoglycans based on 2, 4-diaminobutyric Acid. J Bacteriol 48:459–470. https://doi.org/10.1111/j.1365-2672.1980.tb01036.x
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230. https://doi.org/10.1099/00221287-100-2-221
Dahl C, Franz B, Hensen D, Kesselheim A, Zigann R (2013) Sulfite oxidation in the purple sulfur bacterium Allochromatium vinosum: identification of SoeABC as a major player and relevance of SoxYZ in the process. Microbiology (reading) 159:2626–2638. https://doi.org/10.1099/mic.0.071019-0
Dos Santos PC, Fang Z, Mason SW, Setubal JC, Dixon R (2012) Distribution of nitrogen fixation and nitrogenase-like sequences amongst microbial genomes. BMC Genomics 13:162. https://doi.org/10.1186/1471-2164-13-162
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376. https://doi.org/10.1007/BF01734359
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416. https://doi.org/10.1093/sysbio/20.4.406
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P et al (2007) DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91. https://doi.org/10.1099/ijs.0.64483-0
Govindarajan M, Balandreau J, Kwon SW, Weon HY, Lakshminarasimhan C (2008) Effects of the inoculation of Burkholderia vietnamensis and related endophytic diazotrophic bacteria on grain yield of rice. Microb Ecol 55:21–37. https://doi.org/10.1007/s00248-007-9247-9
Green LS, Laudenbach DE, Grossman AR (1989) A region of a cyanobacterial genome required for sulfate transport. Proc Natl Acad Sci U S A 86:1949–1953. https://doi.org/10.1073/pnas.86.6.1949
Guo K, Yang J, Yu N, Luo L, Wang E (2023) Biological nitrogen fixation in cereal crops: Progress, strategies, and perspectives. Plant Commun 4:100499. https://doi.org/10.1016/j.xplc.2022.100499
Kamlage B (1996) Methods for General and Molecular Bacteriology. Edited by P. Gerhardt, R. G. E. Murray, W. A. Wood and N. R. Krieg. 791 pages, numerous figures and tables. American Society for Microbiology, Washington, D.C., 1994. Price: 55.00 £. Nahrung-food 40:103–103. https://doi.org/10.1002/food.19960400226
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:D277–D280. https://doi.org/10.1093/nar/gkh063
Kanehisa M, Goto S, Hattori M, Aoki-Kinoshita KF, Itoh M et al (2006) From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res 34:D354–D357. https://doi.org/10.1093/nar/gkj102
Kim J, Na SI, Kim D, Chun J (2021) UBCG2: Up-to-date bacterial core genes and pipeline for phylogenomic analysis. J Microbiol 59:609–615. https://doi.org/10.1007/s12275-021-1231-4
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. https://doi.org/10.1007/BF01731581
Kovacs N (1956) Identification of Pseudomonas pyocyanea by the oxidase reaction. Nature 178:703. https://doi.org/10.1038/178703a0
Kroppenstedt RM (1982) Separation of bacterial menaquinones by HPLC using reverse phase (RP18) and a silver loaded Ion exchanger as stationary phases. J Liq Chromatogr 5:2359–2367. https://doi.org/10.1080/01483918208067640
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lagesen K, Hallin P, Rødland EA (2007) RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 35:3100–3108. https://doi.org/10.1093/nar/gkm160
Liu GH, Yang S, Tang R, Xie CJ, Zhou SG (2022) Genome analysis and description of three novel diazotrophs Geomonas species isolated from paddy soils. Front Microbiol 12:801462. https://doi.org/10.3389/fmicb.2021.801462
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964. https://doi.org/10.1093/nar/25.5.955
Maeda S, Omata T (1997) Substrate-binding lipoprotein of the cyanobacterium Synechococcus sp. strain PCC 7942 involved in the transport of nitrate and nitrite. J Biol Chem 272:3036–3041. https://doi.org/10.1074/jbc.272.5.3036
Maeda S, Omata T (2009) Nitrite transport activity of the ABC-type cyanate transporter of the cyanobacterium Synechococcus elongatus. J Bacteriol 191:3265–3272. https://doi.org/10.1128/JB.00013-09
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics 14:60. https://doi.org/10.1186/1471-2105-14-60
Meier-Kolthoff JP, Carbasse JS, Peinado-Olarte RL, Göker M (2022) TYGS and LPSN: a database tandem for fast and reliable genome-based classification and nomenclature of prokaryotes. Nucleic Acids Res 50:D801–D807. https://doi.org/10.1093/nar/gkab902
Minnikin DE, Collins MD, Goodfellow M (1979) Fatty acid and polar lipid composition in the classification of Cellulomonas, Oerskovia and related taxa. J Bacteriol 47:87–95. https://doi.org/10.1111/j.1365-2672.1979.tb01172.x
Mus F, Crook MB, Garcia K et al (2016) Symbiotic nitrogen fixation and the challenges to its extension to nonlegumes. Appl Environ Microbiol 82:36981–43710. https://doi.org/10.1128/AEM.01055-16
Nakajima A, Aono T, Tsukada S, Siarot L, Ogawa T et al (2012) Lon protease of Azorhizobium caulinodans ORS571 is required for suppression of reb gene expression. Appl Environ Microbiol 78:6251–6261. https://doi.org/10.1128/AEM.01039-12
Nojiri Y, Kaneko Y, Azegami Y, Shiratori Y, Ohte N, et al. (2020) Dissimilatory nitrate reduction to ammonium and responsible microbes in Japanese rice paddy soil. Microbes Environ 35:ME20069. https://doi.org/10.1264/jsme2.ME20069
Ogawa K, Akagawa E, Yamane K et al (1995) The nasB operon and nasA gene are required for nitrate/nitrite assimilation in Bacillus subtilis. J Bacteriol 177:1409–1413. https://doi.org/10.1128/jb.177.5.1409-1413.1995
Ouattara AS, Traore AS, Garcia JL (1992) Characterization of Anaerovibrio burkinabensis sp. nov. a lactate fermenting bacterium isolated from rice field soils. Int J Syst Evol Microbiol 42:390–397. https://doi.org/10.1099/00207713-42-3-390
Pandey A, Suter H, He J, Hu H, Chen D (2019) Dissimilatory nitrate reduction to ammonium dominates nitrate reduction in long-term low nitrogen fertilized rice paddies. Soil Biol Biochem 131:149–156. https://doi.org/10.1016/j.soilbio.2019.01.007
Parks DH, Chuvochina M, Waite DW, Rinke C, Skarshewski A et al (2018) A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat Biotechnol 36:996–1004. https://doi.org/10.1038/nbt.4229
Parte AC, Sardà Carbasse J, Meier-Kolthoff JP, Reimer LC, Göker M (2020) List of Prokaryotic names with Standing in Nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol 70:5607–5612. https://doi.org/10.1099/ijsem.0.004332
Postgate JR (1972) Chapter XIII The acetylene reduction test for nitrogen fixation. Methods Microbiol p343–356. https://doi.org/10.1016/S0580-9517(08)70604-4
Qin QL, Xie BB, Zhang XY et al (2014) A proposed genus boundary for the prokaryotes based on genomic insights. J Bacteriol 196:2210–2215. https://doi.org/10.1128/JB.01688-14
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106:19126–19131. https://doi.org/10.1073/pnas.0906412106
Rodriguez-R LM, Konstantinidis KT (2014) Bypassing cultivation to identify bacterial species. Microbe 9:111–118. https://doi.org/10.1128/microbe.9.111.1
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. USFCC News 20:16
Simon J, Gross R, Einsle O, Kroneck PM, Kröger A, Klimmek O (2000) A NapC/NirT-type cytochrome c (NrfH) is the mediator between the quinone pool and the cytochrome c nitrite reductase of Wolinella succinogenes. Mol Microbiol 35:686–696. https://doi.org/10.1046/j.1365-2958.2000.01742.x
Strömpl C, Tindall BJ, Jarvis GN, Lünsdorf H, Moore ER et al (1999) A re-evaluation of the taxonomy of the genus Anaerovibrio, with the reclassification of Anaerovibrio glycerini as Anaerosinus glycerini gen. nov., comb. nov., and Anaerovibrio burkinabensis as Anaeroarcus burkinensis [corrig.] gen. nov., comb. nov. Int J Syst Bacteriol 49:1861–1872. https://doi.org/10.1099/00207713-49-4-1861
Strömpl C, Tindall BJ, Lünsdorf H, Wong TY, Moore ER et al (2000) Reclassification of Clostridium quercicolum as Dendrosporobacter quercicolus gen. nov., comb. nov. Int J Syst Evol Microbiol 50:101–106. https://doi.org/10.1099/00207713-50-1-101
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Tyagi J, Ahmad S, Malik M (2022) Nitrogenous fertilizers: impact on environment sustainability, mitigation strategies, and challenges. Int J Environ Sci Technol 19:11649–11672. https://doi.org/10.1007/s13762-022-04027-9
Wirth JS, Whitman WB (2018) Phylogenomic analyses of a clade within the Roseobacter group suggest taxonomic reassignments of species of the genera Aestuariivita, Citreicella, Loktanella, Nautella, Pelagibaca, Ruegeria, Thalassobius, Thiobacimonas and Tropicibacter, and the proposal of six novel genera. Int J Syst Evol Microbiol 68:2393–2411. https://doi.org/10.1099/ijsem.0.002833
Woo PC, Teng JL, Leung KW, Lau SK, Woo GK et al (2005) Anaerospora hongkongensis gen. nov. sp. nov., a novel genus and species with ribosomal DNA operon heterogeneity isolated from an intravenous drug abuser with pseudobacteremia. Microbiol Immunol 49:31–39. https://doi.org/10.1111/j.1348-0421.2005.tb03637.x
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y et al (2017a) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Yoon SH, Ha SM, Lim JM, Kwon SJ, Chun J (2017b) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 110:1281–1286. https://doi.org/10.1007/s10482-017-0844-4
Funding
This work was supported by the National Science Fund for Distinguished Young Scholars of China (41925028), National Natural Science Foundation of China (42277257), Natural Science Foundation of Fujian Province (2023J01202 and 2023J01141342), Natural Science Foundation of Fujian Province for Distinguished Young Scholars (2021J06018) and Research Supporting Project number (RSP2024R205), King Saud University, Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
SGZ and GHL designed research and project outline. CJX performed deposition and polyphasic taxonomy. LY performed isolation. RT, SH and SY performed genome analysis. CJX drafted the manuscript. SGZ, HA, CR and GHL revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflicts of interests
The authors declare that they had no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The GenBank accession numbers for 16S rRNA gene and genome sequence of strain SG130T are OR142399 and JAUAOA000000000, respectively.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xie, CJ., Yao, L., Tang, R. et al. Azotosporobacter soli gen. nov., sp. nov., a novel nitrogen-fixing bacterium isolated from paddy soil. Antonie van Leeuwenhoek 117, 79 (2024). https://doi.org/10.1007/s10482-024-01978-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10482-024-01978-6