Abstract
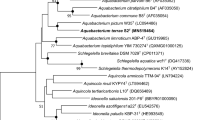
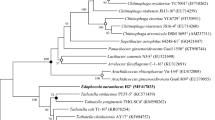
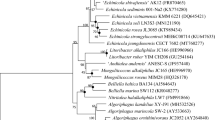
A Gram-stain-positive, orange-pigmented, rod-shaped and flagellated bacterial strain T12T was isolated from wetland soil in Kunyu Mountain Wetland in Yantai, China. The strain was able to grow at 15–40 °C (optimum 37 °C), at 0.0–9.0% NaCl (optimum 2%, w/v) and at pH 5.5–9.0 (optimum 8.5). A phylogenetic analysis based on the 16S rRNA gene sequence indicated that strain T12T is a member of the family Planococcaceae, sharing 97.6% and 97.1% sequence similarity with the type strains of Jeotgalibacillus salarius and Jeotgalibacillus marinus, respectively. Genome-based analyses revealed a genome size of 3,506,682 bp and a DNA G + C content of 43.7%. Besides, the genome sequence led to 55.0–74.6% average amino acid identity values and 67.8–74.7% average nucleotide identity values between strain T12T and the current closest relatives. Digital DNA-DNA hybridization of strain T12T with the type strains of Jeotgalibacillus proteolyticus and J. marinus demonstrated 19.0% and 20.3% relatedness, respectively. The chemotaxonomic analysis showed that the sole quinone was MK-7. The predominant cellular fatty acids were iso-C15:0, anteiso-C15:0, C16:1ω7c alcohol and iso-C14:0. The polar lipids consisted of an unidentified aminolipid, phosphatidylglycerol, diphosphatidylglycerol and two unidentified phospholipids. Based on the polyphasic characterization, strain T12T is considered to represent a novel species, for which the name Jeotgalibacillus aurantiacus sp. nov. is proposed. The type strain is T12T (= KCTC 43296 T = MCCC 1K07171T).



Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this published article, its supplementary information files and GenBank/EMBL/DDBJ. The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain T12T is MW527064 and the number for the whole genome sequence is JACNMS000000000. Supplementary figures and Supplementary tables are available with the online version of this paper.
Abbreviations
- AAI:
-
Average amino acid identity
- AL:
-
Aminolipid
- ANI:
-
Average nucleotide identity
- BGCs:
-
Biosynthetic gene clusters
- COG:
-
Cluster of orthologous groups
- dDDH:
-
Digital DNA-DNA hybridization
- DPG:
-
Diphosphatidylglycerol
- GGDC:
-
Genome-to-genome distance calculator
- HPLC:
-
High performance liquid chromatography
- KCTC:
-
Korean collection for type cultures
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- MA:
-
Marine agar 2216
- MB:
-
Marine broth 2216
- MEGA:
-
Molecular evolutionary genetics analysis
- MIDI:
-
Microbial identification system
- MCCC:
-
Marine culture collection of China
- PC:
-
Phosphatidylcholine
- PG:
-
Phosphatidylglycerol
- RAST:
-
Rapid annotations using subsystems technology
References
Aziz RK (2008) The RAST server: rapid annotations using subsystems technology. BMC Genomics 9:75. https://doi.org/10.1186/1471-2164-9-75
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:W81–W87. https://doi.org/10.1093/nar/gkz310
Chen YG, Peng DJ, Chen QH, Zhang YQ, Tang SK, Zhang DC, Peng QZ, Li WJ (2010) Jeotgalibacillus soli sp. nov., isolated from non-saline forest soil, and emended description of the genus Jeotgalibacillus. Antonie Van Leeuwenhoek 98:415–421. https://doi.org/10.1007/s10482-010-9455-z
Consortium Tu (2019) UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res 47:D506–D515. https://doi.org/10.1093/nar/gky1049
Cunha S, Tiago I, Paiva G, Nobre F, Da Costa MS, Verissimo A (2012) Jeotgalibacillus soli sp. nov., a gram-stain-positive bacterium isolated from soil. Int J Syst Evolut Microbiol 62:608–612. https://doi.org/10.1099/ijs.0.028878-0
Dundas LD, Larsen H (1962) The physiological role of the carotenoid pigments of halobacterium salinarium. Arch Microbiol 44:233–239. https://doi.org/10.1007/BF00510943
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evolut 17:368–376. https://doi.org/10.1007/BF01734359
Galasso C, Corinaldesi C, Sansone C (2017) Carotenoids from marine organisms: biological functions and industrial applications. Antioxidants (basel). https://doi.org/10.3390/antiox6040096
Gonzalez-Beltran A, Li P, Zhao J, Avila-Garcia MS, Roos M, Thompson M, Van Der Horst E, Kaliyaperumal R, Luo R, Lee TL, Lam TW, Edmunds SC, Sansone SA, Rocca-Serra P (2015) From peer-reviewed to peer-reproduced in scholarly publishing: the complementary roles of data models and workflows in bioinformatics. GigaScience. https://doi.org/10.1186/2047-217X-1-18
Hiraishi A, Ueda Y, Ishihara J, Mori T (1996) Comparative lipoquinone analysis of influent sewage and activated sludge by high-performance liquid chromatography and photodiode array detection. J Gen Appl Microbiol 42:457–469. https://doi.org/10.2323/jgam.42.457
Kallscheuer N, Moreira C, Airs R, Llewellyn CA, Wiegand S, Jogler C, Lage OM (2019) Pink- and orange-pigmented planctomycetes produce saproxanthin-type carotenoids including a rare C45 carotenoid. Environ Microbiol Rep 11:741–748. https://doi.org/10.1111/1758-2229.12796
Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M (2016) KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44:D457–D462. https://doi.org/10.1093/nar/gkv1070
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evolut 16:111–120. https://doi.org/10.1007/BF01731581
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Li Y, Zhang ZP, Xu ZX, Fang DB, Wang ET, Shao S, Du ZJ, Liu W, Xie ZH (2018) Jeotgalibacillus proteolyticus sp. nov., a protease-producing bacterium isolated from ocean sediments. Int J Syst Evolut Microbiol 68:3790–3795. https://doi.org/10.1099/ijsem.0.003060
Meier-Kolthoff JP, Auch AF, Klenk HP, Goker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:14. https://doi.org/10.1186/1471-2105-14-60
Moyes RB, Reynolds J, Breakwell DP (2009) Differential staining of bacteria: gram stain. Curr Protoc Microbiol. https://doi.org/10.1002/9780471729259.mca03cs15
Richter M, Rossello-Mora R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106:19126–19131. https://doi.org/10.1073/pnas.0906412106
Ruger HJ (1983) Differentiation of Bacillus globisporus, Bacillus marinus comb. nov., Bacillus aminovorans, and Bacillus insolitus. Int J Syst Bacteriol 33:157–161. https://doi.org/10.1099/00207713-33-2-157
Ruger HJ, Richter G (1979) Bacillus globisporus subsp. marinus subsp. nov. Int J Syst Bacteriol 29:196–203. https://doi.org/10.1099/00207713-29-3-196
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evolut. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sawant SS, Salunke BK, Kim BS (2015) A rapid, sensitive, simple plate assay for detection of microbial alginate lyase activity. Enzyme Microb Technol 77:8–13. https://doi.org/10.1016/j.enzmictec.2015.05.003
Shindo K, Misawa N (2014) New and rare carotenoids isolated from marine bacteria and their antioxidant activities. Mar Drugs 12:1690–1698. https://doi.org/10.3390/md12031690
Simpson JT, Wong K, Jackman SD, Schein JE, Jones SJM, Birol İ (2009) ABySS: a parallel assembler for short read sequence data. Genome Res 19:1117–1123. https://doi.org/10.1101/gr.089532.108
Srinivas A, Divyasree B, Sasikala C, Tushar L, Bharti D, Ramana CV (2016) Description of Jeotgalibacillus alkaliphilus sp. nov., isolated from a solar salt pan, and Jeotgalibacillus terrae sp. nov., a name to replace ‘Jeotgalibacillus soli’ Chen et al. 2010. Int J Syst Evolut Microbiol 66:5167–5172. https://doi.org/10.1099/ijsem.0.001491
Tindall BJ, Sikorski J, Smibert RA, Krieg NR (2007) Phenotypic characterization and the principles of comparative systematics. Methods Gen Mol Microbiol. https://doi.org/10.1128/9781555817497.ch15
Yaakop AS, Chan KG, Ee R, Kahar UM, Kon WC, Goh KM (2015) Isolation of Jeotgalibacillus malaysiensis sp. nov. from a sandy beach, and emended description of the genus Jeotgalibacillus. Int J Syst Evolut Microbiol 65:2215–2221. https://doi.org/10.1099/ijs.0.000242
Yaakop AS, Chan KG, Ee R, Lim YL, Lee SK, Manan FA, Goh KM (2016) Characterization of the mechanism of prolonged adaptation to osmotic stress of Jeotgalibacillus malaysiensis via genome and transcriptome sequencing analyses. Sci Rep 6:33660. https://doi.org/10.1038/srep33660
Yang SJ, Cho JC (2008) Gaetbulibacter marinus sp. nov., isolated from coastal seawater, and emended description of the genus Gaetbulibacter. Int J Syst Evolut Microbiol 58:315–318. https://doi.org/10.1099/ijs.0.65382-0
Yoon JH, Weiss N, Lee KC, Lee IS, Kang KH, Park YH (2001) Jeotgalibacillus alimentarius gen. nov., sp. nov., a novel bacterium isolated from jeotgal with l-lysine in the cell wall, and reclassification of Bacillus marinus ruger 1983 as Marinibacillus marinus gen. nov., comb. nov. Int J Syst Evolut Microbiol 51:2087–2093. https://doi.org/10.1099/00207713-51-6-2087
Yoon JH, Kim IG, Schumann P, Oh TK, Park YH (2004) Marinibacillus campisalis sp. nov., a moderate halophile isolated from a marine solar saltern in Korea, with emended description of the genus Marinibacillus. Int J Syst Evolut Microbiol 54:1317–1321. https://doi.org/10.1099/ijs.0.02779-0
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evolut Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Yoon JH, Kang SJ, Schumann P, Oh TKJIJOS, Microbiology E (2010) Jeotgalibacillus salarius sp. nov., isolated from a marine saltern, and reclassification of Marinibacillus marinus and Marinibacillus campisalis as Jeotgalibacillus marinus comb. nov. and Jeotgalibacillus campisalis comb. nov., respectively. Int J Syst Evolut Microbiol 60:15. https://doi.org/10.1099/ijs.0.008318-0
Zhou YX, Du ZJ, Chen GJ (2016) Seonamhaeicola algicola sp. nov., a complex-polysaccharide-degrading bacterium isolated from Gracilaria blodgettii, and emended description of the genus Seonamhaeicola. Int J Syst Evolut Microbiol 66:2064–2068. https://doi.org/10.1099/ijsem.0.000991
Zhu S, Hegemann JD, Fage CD, Zimmermann M, Xie X, Linne U, Marahiel MA (2016) Insights into the unique phosphorylation of the lasso peptide paeninodin. J Biol Chem 291:13662–13678. https://doi.org/10.1074/jbc.M116.722108
Acknowledgements
We are very grateful to Prof. Aharon Oren for his advice on naming the species.
Funding
This work was supported by the National Natural Science Foundation of China (31700116), the Natural Science Foundation of Shandong Province (ZR2017MC019), the China Postdoctoral Science Foundation (2017M62218), and the Key Science and Technology Program of Weihai (1070413421511).
Author information
Authors and Affiliations
Contributions
HNJ drafted the manuscript. BXW, STY, HNJ and YM performed isolation, deposition and identification. STY, BXW and MJZ performed the genome analysis. STY and YXZ revised the manuscript. YXZ designed all the experiments and supervised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that there is no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain T12T is MW527064 and the number for the whole genome sequence is JACNMS000000000 (assembly accession GCA_020595125.1).
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiang, HN., Yun, ST., Wang, BX. et al. Jeotgalibacillus aurantiacus sp. nov., a novel orange-pigmented species with a carotenoid biosynthetic gene cluster, isolated from wetland soil. Antonie van Leeuwenhoek 115, 773–782 (2022). https://doi.org/10.1007/s10482-022-01731-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-022-01731-x