Abstract
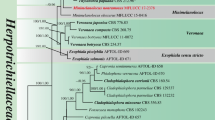
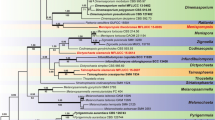
During an investigation of the diversity of aquatic hyphomycetes from southern China, two interesting isolates were collected. These two isolates were cultured and sequenced, and a BLAST search of their LSU sequences against data in GenBank revealed that the closest related taxa were in the genus Microthyrium. Phylogenetic analyses, based on the combined sequence data from the internal transcribed spacer (ITS) and large nuclear subunit ribosomal DNA (LSU), revealed that our isolates belong to the Microthyriaceae. Combined morphological characters allowed us to describe our isolates as two new genera and species in Microthyriaceae, named as: Keqinzhangia aquatica and Pseudocoronospora hainanense. The full descriptions, illustrations, and a phylogenetic tree showing the position of the two new genera were provided in this paper.






Similar content being viewed by others
Availability of data and material
These new generated sequences were uploaded to the GenBank database at the National Center for Biotechnology Information (NCBI), and are available.
Code availability
Not applicable.
References
Abarca GH, Castañeda-Ruiz RF, Mota RMA, Hernandez CIB, Gomez S et al (2011) A new species of Heliocephala from Mexico with an assessment of the systematic position of the anamorph genera Heliocephala and Holubovaniella. Mycologia 103:631–640. https://doi.org/10.3852/10-230
Ariyawansa HA, Hyde KD, Jayasiri SC, Buyck B, Chethana KTW et al (2015) Fungal diversity notes 111–252—taxonomic and phylogenetic contributions to fungal taxa. Fungal Diversity 75:27–274. https://doi.org/10.1007/s13225-015-0346-5
Arnaud G (1918) Lés Asterinées. Ann Éc Natl Agric Montp 2(16):1–288
Boonmee S, Ko Ko TW, Chukeatirote E, Hyde KD, Chen H et al. (2012) Two new Kirschsteiniothelia species with Dendryphiopsis anamorphs cluster in Kirschsteiniotheliaceae fam. nov. Mycologia 104(3):698–714. https://doi.org/10.3852/11-089
Cole GT (1986) Models of cell-differentiation in conidia fungi. Microbiol Rev 50(2):95–132
Crous PW, Wingfield MJ, Richardson DM, Le Roux JJ, Strasberg D et al (2016) Fungal planet description sheets: 400–468. Persoonia 36:316–458. https://doi.org/10.3767/003158516X692185
Crous PW, Wingfield MJ, Burgess TI, Carnegie AJ, Hardy GESJ et al (2017) Fungal Planet description sheets: 625–715. Persoonia 39:270–467. https://doi.org/10.3767/persoonia.2017.39.11
Crous PW, Luangsa-Ard JJ, Wingfield MJ, Carnegie AJ, Hernández-Restrepo M et al (2018) Fungal Planet description sheets: 785–867. Persoonia 41:238–417. https://doi.org/10.3767/persoonia.2018.41.12
Crous PW, Wingfield MJ, Lombard L, Roets F, Swart WJ et al (2019) Fungal Planet description sheets: 951–1041. Persoonia 43:223–425. https://doi.org/10.3767/persoonia.2019.43.06
Ellis MB (1971) Dematiaceous Hyphmycetes x. Mycol Paper 125:1–30
Ferrer A, Miller AN, Shearer CA (2011) Minutisphaera and Natipusilla: two new genera of freshwater Dothideomycetes. Mycologia 103:411–423. https://doi.org/10.3852/10-177
Gonzalez II, Garcia D, Guarro J, Gene J (2020) Heliocephala variabilis and Pseudopenidiella vietnamensis: two new Hyphomycetous species in the Microthyriaceae (Dothideomycetes) from Vietnam. Microorganisms 8(4):478. https://doi.org/10.3390/microorganisms8040478
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41(41):95–98. https://doi.org/10.1021/bk-1999-0734.ch008
Hongsanan S, Hyde KD (2017) Phylogenetic placement of Micropeltidaceae. Mycosphere 8(10):1930–1942. https://doi.org/10.5943/mycosphere/8/10/15
Hongsanan S, Chomnunti P, Crous PW, Chukeatirote E, Hyde KD (2014) Introducing Chaetothyriothecium, a new genus of Microthyriales. Phytotaxa 161:157–164
Hongsanan S, Hyde KD, Phookamsak R, Wanasinghe DN, McKenzie EHC et al (2020) Refined families of Dothideomycetes: orders and families incertae sedis in Dothideomycetes. Fungal Diversity 105(1):17–318. https://doi.org/10.1007/s13225-020-00462-6
Hongsanan S, Tian Q, Bahkali AH, Yang J-B, McKenzie EHC et al. (2015) Zeloasperisporiales ord. nov., and two new species of Zeloasperisporium. Cryptogamie Mycologie 36(3):301–317. https://doi.org/10.7872/crym/v36.iss3.2015.301
Kindermann J, El-Ayouti Y, Samuels GJ, Kubicek CP (1998) Phylogeny of the genus Trichoderma based on sequence analysis of the internal transcribed spacer region 1 of the rDNA cluster. Fungal Genet Biol 24(3):298–309. https://doi.org/10.1006/fgbi.1998.1049
Kirk P, Cannon P, Minter D, Stalpers J (2008) Dictionary of the fungi, 10th edn. CAB International, Wallingford, UK
Lumbsch HT, Lindemuth R, Schmitt I (2000) Evolution of filamentous Ascomycetes inferred from LSU rDNA sequence data. Plant Biol 2:525–529. https://doi.org/10.1055/s-2000-7472
Matsushima T (2001) Matsushima Mycological Memoirs No. 10. Matsushima fungus collection, Published by author, Kobe, Japan.
Myers N, Mittermeier RA, Mittermeier CG, Fonseca G, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403(6772):853–858. https://doi.org/10.1038/35002501
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25(7):1253–1256. https://doi.org/10.1093/molbev/msn083
Qiao M, Zheng H, Lv R, Yu Z (2020) Neodactylariales, Neodactylariaceae (Dothideomycetes, Ascomycota): new order and family, with a new species from China. Mycokeys 73(6):69–85. https://doi.org/10.3897/mycokeys.73.54054
Qiao M, Huang Y, Deng C, Yu ZF (2017) Tripospermum sinense sp. nov. from China. Mycotaxon 132(3):513–517. https://doi.org/10.5248/132.513
Qiao M, Guo JS, Tian WG, Yu ZF (2018a) Ellisembia hainanensis sp. nov. from Hainan, China. Mycotaxon 133(1):97–10. https://doi.org/10.5248/133.97
Qiao M, Du X, Bian ZH, Peng J, Yu ZF (2018b) Ellisembia pseudokaradkensis sp. nov. from Hainan, China. Mycotaxon 132(4):813–817. https://doi.org/10.5248/132.813
Qiao M, Zheng H, Zhang Z, Yu ZF (2019) Seychellomyces sinensis sp. nov. from China. Mycotaxon 134(2):391–398. https://doi.org/10.5248/134.391
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A et al. (2012) MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. System Biol 61(3):539–542. https://doi.org/10.1093/sysbio/sys029
Saccardo PA (1883) Sylloge Pyrenomycetum, vol II. Sylloge Fungorum 2:1–815
Samerpitak K, Van der Linde E, Choi HJ, Gerrits van den Ende AHG, Machouart M et al (2014) Taxonomy of Ochroconis, genus including opportunistic pathogens on humans and animals. Fungal Diversity 65:89–126. https://doi.org/10.1007/s13225-013-0253-6
Schoch CL, Crous PW, Groenewald JZ, Boehm EW, Burgess TI et al (2009) A class-wide phylogenetic assessment of Dothideomycetes. Stud Mycol 64:1–15. https://doi.org/10.3114/sim.2009.64.01
Seifert K, Morgan-Jones G, Gams W, Kendrick B (2011) The genera of hyphomycetes. CBS Biodiversity Series 9:997
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22(21):2688–2690. https://doi.org/10.1093/bioinformatics/btl446
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution 30(12):2725–2729.
Theissen F (1913) Hemisphaeriales. Annales Mycologici 11(5):425–467
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882. https://doi.org/10.1093/nar/25.24.4876
Turner D, Kovacs W, Kuhls K, Lieckfeldt E, Peter B et al (1997) Biogeography and phenotypic variation in Trichoderma sect Longibrachiatum and associated Hypocrea species. Mycol Res 101:449–459. https://doi.org/10.1017/S0953756296002845
Vilgalys R, Hester M (1990) Rapid Genetic Identification and mapping of enzymatically amplified ribosomal DNA from Several Cryptococcus Species. J Bacteriol 172(8):4238–4246. https://doi.org/10.1128/jb.172.8.4238-4246.1990
White T, Bruns T, Lee S, Taylor F, White T et al (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols: a guide to methods and applications, vol 18. Academic Press, New York, pp 315–322.
Wijayawardene NN, Hyde KD, Lumbsch HT, Jian KL, Maharachchikumbura SSN et al (2018) Outline of Ascomycota: 2017. Fungal Diversity 88(2):167–263. https://doi.org/10.1007/s13225-018-0394-8
Wijayawardene N, Hyde KD, Al-Ani L, Tedersoo L, Haelewaters D et al (2020) Outline of fungi and fungi-like taxa. Mycosphere 11(1):1160–1456. https://doi.org/10.5943/mycosphere/11/1/8
Wu HX, Schoch CL, Boonmee S, Bahkali AH, Chomnunti P et al (2011) A reappraisal of Microthyriaceae. Fungal Diversity 51(1):189–248. https://doi.org/10.1007/s13225-011-0143-8
Wu HX, Li YM, Ariyawansa HA, Li WJ, Hyde KD (2014). A new species of Microthyrium from Yunnan, China. Phytotaxa 176(1):213–218. https://doi.org/10.11646/phytotaxa.176.1.21
Yen LTH, Yamaguchi K, Tsurumi Y, Hop DV, Ando K (2018) Hamatispora, a new genus of aquatic fungi in Microthyriales isolated from fallen leaves in Vietnam. Mycoscience 59(6):467–472. https://doi.org/10.1016/j.myc.2018.04.004
Zhang M, Zhang TY (2004) A new species of Coronospora from China. Mycosystema 23:331–332. https://doi.org/10.3969/j.issn.1672-6472.2004.03.004
Zheng H, Wan Y, Li J, Castañeda-Ruiz RF, Yu ZF (2020a) Phialolunulospora vermispora (Chaetosphaeriaceae, Sordariomycetes), a novel asexual genus and species from freshwater in southern China. Mycokeys 76:17–30. https://doi.org/10.3897/mycokeys.76.57410
Zheng H, Yu Z, Xu J, Castañeda-Ruiz RF, Qiao M (2020b) Ramichloridium endophyticum sp. nov., a novel species of endophytic fungus from Potamogeton pectinatus. Int J System Evolut Microbiol 70(5). https://doi.org/10.1099/ijsem.0.004190
Zheng H, Zhang Z, Liu DZ, Yu ZF (2019) Memnoniella sinensis sp. nov., a new species from China and a key to species of the genus. Int J System Evolut Microbiol 69(10). https://doi.org/10.1099/ijsem.0.003605
Zheng H, Li J, Gou JS, Qiao M, Yu ZF (2021a) Anacraspedodidymum submersum sp. nov. (Chaetosphaeriaceae, Chaetosphaeriales), a new species of freshwater hyphomycetes from southwest China. Int J System Evolut Microbiol 71(2). https://doi.org/10.1099/ijsem.0.004650
Zheng H, Qiao M, Lv Y, Du X, Zhang KQ et al (2021b) New species of Trichoderma isolated as endophytes and saprobes from Southwest China. J Fungi 7(6):467. https://doi.org/10.3390/jof7060467
Zheng H, Qiao M, Xu JP, Yu ZF (2021c) Culture-based and culture-independent assessments of endophytic fungal diversity in aquatic plants in Southwest China. Front Fungal Biol 2:27. https://doi.org/10.3389/ffunb.2021.692549
Acknowledgements
We thank Linlin Fang for her work on this manuscript; we are also grateful for the support of Yunnan University Research and Innovation Fund for Postgraduates (2021Y294).
Funding
This work was financed by the National Natural Science Foundation Program of PR China (31770026, 31760012).
Author information
Authors and Affiliations
Contributions
ZY conceived and designed the study. HZ and MQ wrote the manuscript. JG and JP conducted the experiments. RFC contributed actively in the identification and the taxonomy of the fungal strains.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare to have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zheng, H., Qiao, M., Guo, J. et al. Keqinzhangia aquatica gen. et sp. nov. and Pseudocoronospora hainanense gen. et sp. nov., isolated from freshwater in southern China. Antonie van Leeuwenhoek 115, 203–213 (2022). https://doi.org/10.1007/s10482-021-01688-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01688-3