Abstract
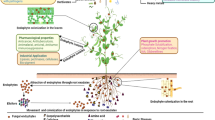
One of the most diverse groups of bioactive bacterial metabolites is the ribosomally synthesised and post-translationally modified peptides (RiPPs) with different bioactivities. The process of genome mining has made it possible to predict the presence of such clusters among the huge genomic data available today. Despite the great potential of actinobacteria in producing natural products and the myriad of completely sequenced genomes available, a comprehensive genome mining of these bacteria for RiPPs is lacking. Here, a collection of 629 complete actinobacterial genomes were analysed to explore their RiPP biosynthesis potential. Using BAGEL3 genome mining tool, the presence of 477 RiPP biosynthesis gene clusters (BGCs) was shown, including all known classes of bacterial RiPPs. RiPP-encoding potential was shown to be widespread among different members of actinobacteria especially within the plant and soil-inhabiting strains. The notable presence of LAP BGCs in plant-associating actinobacteria was also illustrated. Streptomyces, Amycolatopsis, Kitasatospora and Frankia showed greater potential in RiPP biosynthesis while lanthipeptides and lasso peptides were the most distributed RiPPs. Three cyanobactin BGCs were also detected. Generally evidence of promising ability of actinobacteria to synthesise diverse classes of RiPPs as well as information needed to rationally select appropriate taxa for rational screening of specific RiPPs are presented.








Similar content being viewed by others
References
Abriouel H, Valdivia E, Martí M, Maqueda M, Gálvez A (2003) A simple method for semi-preparative-scale production and recovery of enterocin AS-48 derived from Enterococcus faecalis subsp. liquefaciens A-48-32. J Microbiol Methods 55:599–605
Agrawal P, Khater S, Gupta M, Sain N, Mohanty D (2017) RiPPMiner: a bioinformatics resource for deciphering chemical structures of RiPPs based on prediction of cleavage and cross-links. Nucleic acids research 45:80–88
Allenby NE, Watts CA, Homuth G, Prágai Z, Wipat A, Ward AC, Harwood CR (2006) Phosphate starvation induces the sporulation killing factor of Bacillus subtilis. J Bacteriol 188:5299–5303
Arnison PG et al (2013) Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature. Nat Prod Rep 30:108–160
Babalola OO (2010) Beneficial bacteria of agricultural importance Biotechnology letters 32:1559–1570
Babasaki K, Takao T, Shimonishi Y, Kurahashi K, Subtilosin A (1985) A new antibiotic peptide produced by Bacillus subtilis 168: isolation, structural analysis, and biogenesis. J Biochem 98:585–603
Bagley MC, Dale JW, Merritt EA, Xiong X (2005) Thiopeptide antibiotics. Chem Rev 105:685–714
Bartholomae M, Buivydas A, Viel JH, Montalban-Lopez M, Kuipers OP (2017) Major gene-regulatory mechanisms operating in ribosomally synthesized and post-translationally modified peptide (RiPP) biosynthesis. Mol Microbiol 106:186–206
Bérdy J (2012) Thoughts and facts about antibiotics: where we are now and where we are heading. J Antibiot 65:385
Bierbaum G, Sahl H-G (2009) Lantibiotics: mode of action, biosynthesis and bioengineering. Curr Pharm Biotechnol 10:2–18
Bonelli RR, Schneider T, Sahl H-G, Wiedemann I (2006) Insights into in vivo activities of lantibiotics from gallidermin and epidermin mode-of-action studies. Antimicrob Agents Chemother 50:1449–1457
Cascales L, Craik DJ (2010) Naturally occurring circular proteins: distribution, biosynthesis and evolution. Org Biomol Chem 8:5035–5047
Chertkov O et al (2011) Complete genome sequence of Thermomonospora curvata type strain (B9 T). Stand Genom Sci 4:13
Chun J et al (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466
Claesen J, Bibb M (2010) Genome mining and genetic analysis of cypemycin biosynthesis reveal an unusual class of posttranslationally modified peptides. Proc Natl Acad Sci 107:16297–16302
Claesen J, Bibb MJ (2011) Biosynthesis and regulation of grisemycin, a new member of the linaridin family of ribosomally synthesized peptides produced by Streptomyces griseus IFO 13350. J Bacteriol 193:2510–2516
Cock IE (2012) Species selection for pharmacognostic studies. Pharmacogn Mag 8:182
Cotter PD, Ross RP, Hill C (2013) Bacteriocins—a viable alternative to antibiotics? Nat Rev Microbiol 11:95
Cragg GM, Newman DJ (2013) Natural products: a continuing source of novel drug leads. Biochim Biophys Acta (BBA)-Gen Subj 1830:3670–3695
Czekster CM, Ge Y, Naismith JH (2016) Mechanisms of cyanobactin biosynthesis. Curr Opin Chem Biol 35:80–88
Dangi SK, Dubey S, Bhargava S (2017) Cyanobacterial diversity: a potential source of bioactive compounds situations 3:5
David B, Wolfender J-L, Dias DA (2015) The pharmaceutical industry and natural products: historical status and new trends. Phytochem Rev 14:299–315
Demain AL (2014) Importance of microbial natural products and the need to revitalize their discovery. J Ind Microbiol Biotechnol 41:185–201
Diaz M, Valdivia E, Martínez-Bueno M, Fernández M, Soler-González AS, Ramírez-Rodrigo H, Maqueda M (2003) Characterization of a new operon, as-48EFGH, from the as-48 gene cluster involved in immunity to enterocin AS-48. Appl Environ Microbiol 69:1229–1236
Doroghazi JR, Metcalf WW (2013) Comparative genomics of actinomycetes with a focus on natural product biosynthetic genes. BMC Genom 14:611
Draper LA, Cotter PD, Hill C, Ross RP (2015) Lantibiotic resistance. Microbiol Mol Biol Rev 79:171–191
Duncan KR et al (2015) Molecular networking and pattern-based genome mining improves discovery of biosynthetic gene clusters and their products from Salinispora species. Chem Biol 22:460–471
Duquesne S, Destoumieux-Garzón D, Zirah S, Goulard C, Peduzzi J, Rebuffat S (2007) Two enzymes catalyze the maturation of a lasso peptide in Escherichia coli. Chem Biol 14:793–803
Favret ME, Yousten AA (1989) Thuricin: the bacteriocin produced by Bacillus thuringiensis. J Invertebr Pathol 53:206–216
Flühe L, Marahiel MA (2013) Radical S-adenosylmethionine enzyme catalyzed thioether bond formation in sactipeptide biosynthesis. Curr Opin Chem Biol 17:605–612
Gabrielsen C, Brede DA, Nes IF, Diep DB (2014) Circular bacteriocins: biosynthesis and mode of action. Appl Environ Microbiol 80:6854–6862
Gomes KM, Duarte RS, de Freire Bastos MDC (2017) Lantibiotics produced by Actinobacteria and their potential applications (a review). Microbiology 163:109–121
Gonzalez DJ, Clostridiolysin S et al (2010) A post-translationally modified biotoxin from Clostridium botulinum. J Biol Chem 285:28220–28228
Grove TL et al (2017) Structural insights into thioether bond formation in the biosynthesis of sactipeptides. J Am Chem Soc 139:11734–11744
Gunatilaka A, Wijeratne E (2000) Natural products from bacteria and fungi. Elsevier, Amsterdam
Hamedi J, Poorinmohammad N, Wink J (2017) The role of actinobacteria in biotechnology. In: Biology and biotechnology of actinobacteria. Springer, pp 269–328
Harvey AL, Edrada-Ebel R, Quinn RJ (2015) The re-emergence of natural products for drug discovery in the genomics era. Nat Rev Drug Discov 14:111
Hegemann JD, Zimmermann M, Xie X, Marahiel MA (2015) Lasso peptides: an intriguing class of bacterial natural products. Acc Chem Res 48:1909–1919
Hou Y et al (2012) Structure and biosynthesis of the antibiotic bottromycin D. Org Lett 14:5050–5053
Hudson GA, Mitchell DA (2018) RiPP antibiotics: biosynthesis and engineering potential. Curr Opin Microbiol 45:61–69
Just-Baringo X, Albericio F, Álvarez M (2014) Thiopeptide antibiotics: retrospective and recent advances. Mar Drugs 12:317–351
Knappe TA, Linne U, Zirah S, Rebuffat S, Xie X, Marahiel MA (2008) Isolation and structural characterization of capistruin, a lasso peptide predicted from the genome sequence of Burkholderia thailandensis E264. J Am Chem Soc 130:11446–11454
Komiyama K et al (1993) A new antibiotic, cypemycin. J Antibiot 46:1666–1671
Kononova L, Filatova L, Eroshenko D, Korobov V (2017) Suppression of development of vancomycin-resistant Staphylococcus epidermidis by low-molecular-weight cationic peptides of the lantibiotic family. Microbiology 86:571–582
Kröber M et al (2014) Effect of the strain Bacillus amyloliquefaciens FZB42 on the microbial community in the rhizosphere of lettuce under field conditions analyzed by whole metagenome sequencing. Front Microbiol 5:252
Li C, Kelly WL (2010) Recent advances in thiopeptide antibiotic biosynthesis. Nat Prod Rep 27:153–164
Liao R et al (2009) Thiopeptide biosynthesis featuring ribosomally synthesized precursor peptides and conserved posttranslational modifications. Chem Biol 16:141–147
Liu N et al (2016) Unique post-translational oxime formation in the biosynthesis of the azolemycin complex of novel ribosomal peptides from Streptomyces sp. FXJ1. 264. Chem Sci 7:482–488
Lyon WJ, Glatz BA (1993) Isolation and purification of propionicin PLG-1, a bacteriocin produced by a strain of Propionibacterium thoenii. Appl Environ Microbiol 59:83–88
Maksimov MO, Pan SJ, Link AJ (2012) Lasso peptides: structure, function, biosynthesis, and engineering. Nat Prod Rep 29:996–1006
Martins J, Ramos V, Leão P, Vasconcelos V (2012) Cyanobactins and anticancer bioactivity of cyanobacterial extracts. Planta Med 78:PI59
Martin-Visscher LA, Gong X, Duszyk M, Vederas JC (2009) The three-dimensional structure of carnocyclin A reveals that many circular bacteriocins share a common structural motif. J Biol Chem 284:28674–28681
Melby JO, Nard NJ, Mitchell DA (2011) Thiazole/oxazole-modified microcins: complex natural products from ribosomal templates. Curr Opin Chem Biol 15:369–378
Mo T et al (2017) Biosynthetic insights into linaridin natural products from genome mining and precursor peptide mutagenesis. ACS Chem Biol 12:1484–1488
Montalbán-López M, Spolaore B, Pinato O, Martínez-Bueno M, Valdivia E, Maqueda M, Fontana A (2008) Characterization of linear forms of the circular enterocin AS-48 obtained by limited proteolysis. FEBS Lett 582:3237–3242
Onaka H, Tabata H, Igarashi Y, Sato Y, Furumai T (2001) Goadsporin, a chemical substance which promotes secondary metabolism and morphogenesis in streptomycetes. J Antibiot 54:1036–1044
Ortega MA, Van Der Donk WA (2016) New insights into the biosynthetic logic of ribosomally synthesized and post-translationally modified peptide natural products. Cell Chem Biol 23:31–44
Papagianni M (2003) Ribosomally synthesized peptides with antimicrobial properties: biosynthesis, structure, function, and applications. Biotechnol Adv 21:465–499
Rateb ME et al (2015) Legonaridin, a new member of linaridin RiPP from a Ghanaian Streptomyces isolate. Org Biomol Chem 13:9585–9592
Reddy TB et al (2014) The genomes online database (GOLD) v. 5: a metadata management system based on a four level (meta) genome project classification. Nucleic Acids Res 43:D1099–D1106
Rosengren KJ, Craik DJ (2009) How bugs make lassos. Chem Biol 16:1211–1212
Scholz R et al (2011) Plantazolicin, a novel microcin B17/streptolysin S-like natural product from Bacillus amyloliquefaciens FZB42. J Bacteriol 193:215–224
Sievers F, Higgins DG (2018) Clustal Omega for making accurate alignments of many protein sequences. Protein Sci 27:135–145
Sivonen K, Leikoski N, Fewer DP, Jokela J (2010) Cyanobactins—ribosomal cyclic peptides produced by cyanobacteria. Appl Microb Biotechnol 86:1213–1225
Theuretzbacher U (2017) New drugs–will they solve the problem of resistance to antibiotics? Clin Microbiol Infect 23:695–696
Torsvik V, Øvreås L (2002) Microbial diversity and function in soil: from genes to ecosystems. Curr Opin Microbiol 5:240–245
Udwary DW et al (2011) Significant natural product biosynthetic potential of actinorhizal symbionts of the genus Frankia, as revealed by comparative genomic and proteomic analyses. Appl Environ Microbiol 77:3617–3625
van Belkum MJ, Martin-Visscher LA, Vederas JC (2011) Structure and genetics of circular bacteriocins. Trends Microbiol 19:411–418
van Heel AJ, de Jong A, Montalban-Lopez M, Kok J, Kuipers OP (2013) BAGEL3: automated identification of genes encoding bacteriocins and (non-) bactericidal posttranslationally modified peptides. Nucleic Acids Res 41:448–453
van Heel AJ, Montalban-Lopez M, Oliveau Q, Kuipers OP (2017) Genome-guided identification of novel head-to-tail cyclized antimicrobial peptides, exemplified by the discovery of pumilarin. Microb Genom 3:1–9
Waisvisz J, Van Der Hoeven M, Van Peppen J, Zwennis W (1957) Bottromycin. I. A new sulfur-containing antibiotic. J Am Chem Soc 79:4520–4521
Weld JT (1934) The toxic properties of serum extracts of hemolytic streptococci. J Exp Med 59:83
Wohlleben W, Mast Y, Stegmann E, Ziemert N (2016) Antibiotic drug discovery. Microb Biotechnol 9:541–548
Zeiller M, Rothballer M, Iwobi AN, Böhnel H, Gessler F, Hartmann A, Schmid M (2015) Systemic colonization of clover (Trifolium repens) by Clostridium botulinum strain 2301. Front Microb 6:1207
Zerikly M, Challis GL (2009) Strategies for the discovery of new natural products by genome mining. ChemBioChem 10:625–633
Zhang Q, Doroghazi JR, Zhao X, Walker MC, van der Donk WA (2015a) Expanded natural product diversity revealed by analysis of lanthipeptide-like gene clusters in Actinobacteria. Appl Environ Microbiol 81:4339–4350
Zhang Q, Doroghazi JR, Zhao X, Walker MC, van der Donk WA (2015b) Expanded natural product diversity revealed by analysis of lanthipeptide-like gene clusters in Actinobacteria. Appl Environ Microbiol 81:4339–4350
Zheng Q, Fang H, Liu W (2017) Post-translational modifications involved in the biosynthesis of thiopeptide antibiotics. Org Biomol Chem 15:3376–3390
Ziemert N, Alanjary M, Weber T (2016) The evolution of genome mining in microbes–a review. Nat Prod Rep 33:988–1005
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1.
Actinobacterial strains used for RiPP-related analysis. (XLSX 44 kb)
Online Resource 2.
Habitat-based classification of the actinobacterial strains.(XLSX 50 kb)
Online Resource 3.
Relationship between predicted RiPPs’ features with phylogenetic and ecology of the studied actinobacteria (PDF 257 kb)
Online Resource 4.
Predicted RiPPs’ Open Reading Frames (ORFs) and sequences in the actinobacterial genomes.(XLSX 433 kb)
Rights and permissions
About this article
Cite this article
Poorinmohammad, N., Bagheban-Shemirani, R. & Hamedi, J. Genome mining for ribosomally synthesised and post-translationally modified peptides (RiPPs) reveals undiscovered bioactive potentials of actinobacteria. Antonie van Leeuwenhoek 112, 1477–1499 (2019). https://doi.org/10.1007/s10482-019-01276-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-019-01276-6