Abstract
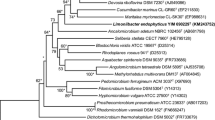
A motile, rod-shaped and yellow coloured proteobacterium, designated strain SYSU D60017T, was isolated from a desert soil sample. The bacterium was found to be an obligately aerobic, mesophilic and neutrophilic chemo-heterotroph. Cells were observed to be Gram-stain negative, catalase positive and oxidase positive. The major cellular fatty acids were identified as C19:0ω8c cyclo and Summed Feature 8 (C18:1ω7c and/or C18:1ω6c). The main respiratory quinone identified was ubiquinone-10. The DNA G + C content was determined to be 63.8% based on draft genome sequence data. The polar lipids detected were identified as diphosphatidylglycerol, phosphatidylcholine, phosphatidylethanolamine, phosphatidylglycerol and five unidentified polar lipids. Phylogenetic analysis based on 16S rRNA gene sequences indicated that strain SYSU D60017T is a member of the order Rhizobiales, but forms a distinct phylogenetic lineage. The differences in the phenotypic characteristics from members of related genera and its distinct phylogenetic position suggested that the isolate SYSU D60017T represents a novel species of a novel genus within the order Rhizobiales, for which the name Flaviflagellibacter deserti gen. nov., sp. nov. is proposed. The type strain of the new taxon is SYSU D60017T (= CGMCC 1.16444T = NBRC 112958T).

Similar content being viewed by others
References
Buck JD (1982) Nonstaining (KOH) method for determination of gram reactions of marine bacteria. Appl Environ Microbiol 44:992–993
Chu JN, Arun AB, Chen WM, Chou JH, Shen FT, Rekha PD, Kämpfer P, Young LS, Lin SY, Young CC (2010) Agaricicola taiwanensis gen. nov., sp. nov., an alphaproteobacterium isolated from the edible mushroom Agaricus blazei. Int J Syst Evol Microbiol 60:2032–2035. https://doi.org/10.1099/ijs.0.016485-0
Collins MD, Jones D (1980) Lipids in the classification and identification of coryneform bacteria containing peptidoglycans based on 2,4-diaminobutyric acid. J Appl Bacteriol 48:459–470. https://doi.org/10.1111/j.1365-2672.1980.tb01036.x
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230
Doronina NV, Trotsenko YA (2015) Incertae sedis IV. Methylopila. In: Whitman WB, Rainey F, Kämpfer P, Trujillo M, Chun J, DeVos P, Hedlund B, Dedysh S (eds) Bergey’s manual of systematics of archaea and bacteria. Wiley, Hoboken. https://doi.org/10.1002/9781118960608.gbm00834
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376. https://doi.org/10.1007/bf01734359
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.2307/2408678
Gonzalez C, Gutierrez C, Ramirez C (1978) Halobacterium vallismortis sp. nov., an amylolytic and carbohydrate-metabolizing, extremely halophilic bacterium. Can J Microbiol 24:710–715
Harrison PG, Strulo B (2000) SPADES—a process algebra for discrete event simulation. J Logic Comput 10:3–42. https://doi.org/10.1093/logcom/10.1.3
Hyatt D, Chen GL, LoCascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform 11:119. https://doi.org/10.1186/1471-2105-11-119
Ivanova E, Doronina N, Trotsenko Y (2007) Hansschlegelia plantiphila gen. nov. sp. nov., a new aerobic restricted facultative methylotrophic bacterium associated with plants. Syst Appl Microbiol 30:444–452. https://doi.org/10.1016/j.syapm.2007.03.001
Kim D, Kang K, Ahn TY (2017) Chthonobacter albigriseus gen. nov., sp. nov., isolated from grass-field soil. Int J Syst Evol Microbiol 67:883–888. https://doi.org/10.1099/ijsem.0.001695
Kovacs N (1956) Identification of Pseudomonas pyocyanea by the oxidase reaction. Nature 178:703–704. https://doi.org/10.1038/178703a0
Leifson E (1960) Atlas of bacterial flagellation. Academic Press, New York
Li WJ, Xu P, Schumann P, Zhang YQ, Pukall R, Xu LH, Stackebrandt E, Jiang CL (2007) Georgenia ruanii sp. nov., a novel actinobacterium isolated from forest soil in Yunnan (China), and emended description of the genus Georgenia. Int J Syst Evol Microbiol 57:1424–1428. https://doi.org/10.1099/ijs.0.64749-0
Lv H, Masuda S, Fujitani Y, Sahin N, Tani A (2017) Oharaeibacter diazotrophicus gen. nov., sp. nov., a diazotrophic and facultatively methylotrophic bacterium, isolated from rice rhizosphere. Int J Syst Evol Microbiol 67:576–582. https://doi.org/10.1099/ijsem.0.001660
Minnikin DE, Collins MD, Goodfellow M (1979) Fatty acid and polar lipid composition in the classification of Cellulomonas, Oerskovia and related taxa. J Appl Bacteriol 47:87–95. https://doi.org/10.1111/j.1365-2672.1979.tb01172.x
Nie GX, Ming H, Li S, Zhou E-M, Cheng J, Tang X, Feng HG, Tang SK, Li WJ (2012) Amycolatopsis dongchuanensis sp. nov., an actinobacterium isolated from soil. Int J Syst Evol Microbiol 62:2650–2656. https://doi.org/10.1099/ijs.0.038125-0
Rosselló-Móra R, Trujillo ME, Sutcliffe IC (2017) Introducing a digital protologue: a timely move towards a database-driven systematics of archaea and bacteria. Antonie Van Leeuwenhoek 110:455–456. https://doi.org/10.1007/s10482-017-0841-7
Rzhetsky A, Nei M (1992) A simple method for estimating and testing minimum evolution trees. Mol Biol Evol 9:945–967
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (2001) Identification of bacteria by gas chromatography of cellular fatty acids. http://www.microbialid.com/PDF/TechNote_101.pdf. Accessed 4 May 2016
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340. https://doi.org/10.1099/00207713-16-3-313
Tamaoka J, Katayama-Fujimura Y, Kuraishi H (1983) Analysis of bacterial menaquinone mixtures by high performance liquid chromatography. J Appl Bacteriol 54:31–36. https://doi.org/10.1111/j.1365-2672.1983.tb01297.x
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tindall BJ, Sikorski J, Smibert RA, Krieg NR (2007) Phenotypic characterization and the principles of comparative systematics. In: Reddy CA, Beveridge TJ, Breznak JA, Marzluf GA, Schmidt TM, Snyder LR (eds) Methods for general and molecular microbiology, 3rd edn. American Society of Microbiology, Washington, DC, pp 330–393. https://doi.org/10.1128/9781555817497.ch15
Tindall BJ, Rosselló-Móra R, Busse H-J, Ludwig W, Kämpfer P (2010) Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 60:249–266. https://doi.org/10.1099/ijs.0.016949-0
Webb HK, Ng HJ, Ivanova EP (2014) The family Methylocystaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes: alphaproteobacteria and betaproteobacteria. Springer, Berlin, pp 341–347. https://doi.org/10.1007/978-3-642-30197-1_254
Wen Y, Huang X, Zhou Y, Hong Q, Li S (2011) Hansschlegelia zhihuaiae sp. nov., isolated from a polluted farmland soil. Int J Syst Evol Microbiol 61:1114–1117. https://doi.org/10.1099/ijs.0.021543-0
Yang LQ, Liu L, Salam N, Xiao M, Kim C-J, Hozzein WN, Park DJ, Li WJ, Zhang HW (2016) Chenggangzhangella methanolivorans gen. nov., sp. nov., a member of the family Methylocystaceae, transfer of Methylopila helvetica Doronina et al. 2000 to Albibacter helveticus comb. nov and emended description of the genus Albibacter. Int J Syst Evol Microbiol 66:2825–2830. https://doi.org/10.1099/ijsem.0.001062
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Zou X-L, Li X-A, Wang X-M, Chen Q, Gao M, Qiu T-L, Sun J-G, Gao J-L (2013) Hansschlegelia beijingensis sp. nov., an aerobic, pink-pigmented, facultatively methylotrophic bacterium isolated from watermelon rhizosphere soil. Int J Syst Evol Microbiol 63:3715–3719. https://doi.org/10.1099/ijs.0.052308-0
Acknowledgements
This research was partly funded by the National Natural Science Foundation of China (Project No. 31850410475) and Natural Science Foundation of Guangdong Province, China (Project No. 2016A030312003). WJL is also supported by Guangdong Province Higher Vocational Colleges and Schools Pearl River Scholar Funded Scheme (2014).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animal rights statement
This article does not contain any study with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dong, L., Han, MX., Wang, D. et al. Flaviflagellibacter deserti gen. nov., sp. nov., a novel member of the order Rhizobiales isolated from a desert soil. Antonie van Leeuwenhoek 112, 947–954 (2019). https://doi.org/10.1007/s10482-019-01228-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-019-01228-0