Abstract
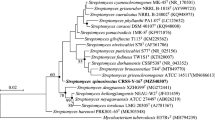
Bacterial strain HV38T was isolated from mangrove soil, which was collected from Thailand. Chemotaxonomic and morphological characteristics were found to be typical of members of the genus Streptomyces. The strain was found to form a distinct phyletic line in the Streptomyces 16S rRNA gene tree and to be closely associated with the type strains of Streptomyces coeruleofuscus CGMCC 4.1667T (98.84 % sequence similarity), Streptomyces chromofuscus CGMCC 4.1451T (98.63 %) and Streptomyces albidoflavus CGMCC 4.1291T (98.56 %). The major menaquinones were identified as MK-9(H8) and MK-9(H10). Its major cellular fatty acids were found to be iso-C14:0, iso-C15:0, anteiso-C15:0, iso-C16:1ω8c, C16:0, anteiso-C16:1ω8c, iso-C16:0 and anteiso-C16:0. The DNA–DNA hybridization values between strain HV38T with S. coeruleofuscus CGMCC 4.1667T, S. chromofuscus CGMCC 4.1451T and S. albidoflavus CGMCC 4.1291T were 32.7 ± 0.9, 21.8 ± 0.3 and 19.9 ± 0.9 %, respectively, which clearly supported the conclusion that they belong to separate genomic species. Cumulatively, the data indicated that strain HV38T represents a novel species of the genus Streptomyces, for which the name Streptomyces ferrugineus sp. nov. is proposed. The type strain is HV38T (=CCTCC AA2014009T = DSM 42152T).

Similar content being viewed by others
References
Cho SH, Han JH, Ko HY, Kim SB (2008) Streptacidiphilus anmyonensis sp. nov., Streptacidiphilus rugosus sp. nov. and Streptacidiphilus melanogenes sp. nov., acidophilic actinobacteria isolated from Pinus soils. Int J Syst Evol Microbiol 58(7):1566–1570
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39(4):783–791
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Hayakawa M, Yoshida Y, Iimura Y (2004) Selective isolation of bioactive soil actinomycetes belonging to the Streptomyces violaceusniger phenotypic cluster. J Appl Microbiol 96(5):973–981
Hong K (2013) Actinomycetes from mangrove and their secondary metabolites. Acta Microbiol Sin 53(11):1142–1148
Hong K, Gao AH, Xie QY, Gao H, Zhuang L, Lin HP, Yu HP, Li J, Yao XS, Goodfellow M, Ruan JS (2009) Actinomycetes for marine drug discovery isolated from mangrove soils and plants in China. Mar Drugs 7:24–44
Hu H, Lin HP, Xie Q, Li L, Xie XQ, Sun M, Hong K (2011) Streptomyces shenzhenensis sp. nov., a novel actinomycete isolated from mangrove sediment. Antonie Van Leeuwenhoek 100(4):631–637
Hu H, Lin HP, Xie Q, Li L, Xie XQ, Hong K (2012) Streptomyces qinglanensis sp. nov., isolated from mangrove sediment. Int J Syst Evol Microbiol 62(3):596–600
Kämpfer P, Labeda DP (2006) International Committee on Systematics of Prokaryotes; Subcommittee on the taxonomy of the Streptomycetaceae: minutes of the meeting, 25 July 2005, San Francisco, CA, USA. Int J Syst Evol Microbiol 56:495
Kämpfer P, Steiof M, Dott W (1991) Microbiological characterization of a fuel-oil contaminated site including numerical identification of heterotrophic water and soil bacteria. Microb Ecol 21(1):227–251
Kim SB, Lonsdale J, Seong CN, Goodfellow M (2003) Streptacidiphilus gen. nov., acidophilic actinomycetes with wall chemotype I and emendation of the family Streptomycetaceae (Waksman and Henrici (1943) AL) emend. Rainey et al. 1997. Antonie van Leeuwenhoek 2:107–116
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62(3):716–721
Lechevalier MP, Lechevalier HA (1980) The chemotaxonomy of actinomycetes. In: Dietz A, Thayer DW (eds) Actinomycete Taxonomy, pp 227–291. Society for Industrial Microbiology, Arlington
Li L, Tang YL, Wei B, Xie QY, Hong K (2013) Micromonospora sonneratiae sp. nov., isolated from a root of Sonneratia apetala. Int J Syst Evol Microbiol 63:2383–2388
Lodders N, Kämpfer P (2007) Streptomycetaceae: Phylogeny, Ecology and Pathogenicity. Encyclopedia of Life Science, pp 1–9. Edited by John Wiley & Sons. Ltd: www.els.net
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JK (1984) An integrated procedure for the extraction of isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Pospiech A, Neumann B (1995) A versatile quick-prep of genomic DNA from gram-positive bacteria. Trends Genet 11(6):217–218
Promnuan Y, Kudo T, Ohkuma M, Chantawannakul P (2013) Streptomyces chiangmaiensis sp. nov. and Streptomyces lannensis sp. nov., isolated from the South-East Asian stingless bee (Tetragonilla collina). Int J Syst Evol Microbiol 63(5):1896–1901
Qu Z, Hong K (2009) The identification of the G+C content of DNA by high performance liquid chromatography for mangrove actinomycetes. Biotechnol Bull S1:209–214
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogenetic tree. Mol Biol Evol 4(4):406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. Technical note 101. Microbial ID, Newark
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Sui JL, Xu XX, Qu Z, Wang HL, Lin HP, Xie QY, Ruan JS, Hong K (2011) Streptomyces sanyensis sp. nov., isolated from mangrove sediment. Int J Syst Evol Microbiol 61(7):1632–1637
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882
Waksman SA, Henrici AT (1943) The nomenclature and classification of the actinomycetes. J Bacteriol 46(4):337–341
Wang C, Xu XX, Qu Z, Wang HL, Lin HP, Xie QY, Ruan JS, Hong K (2011) Micromonospora rhizosphaerae sp. nov., isolated from mangrove rhizosphere soil. Int J Syst Evol Microbiol 61:320–324
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE et al (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Williams ST, Goodfellow M, Alderson G, Wellington EM, Sneath PH, Sackin MJ (1983) Numerical classification of Streptomyces and related genera. J Gen Microbiol 129(6):1743–1813
Xie QY, Qu Z, Lin HP, Li L, Hong K (2012) Micromonospora haikouensis sp. nov., isolated from mangrove soil. Antonie Van Leeuwenhoek 101:649–655
Xu DB, Ye WW, Han Y, Deng ZX, Hong K (2014) Natural products from mangrove actinomycetes. Mar Drugs 12(5):2590–2613
Zhang Z, Wang Y, Ruan J (1997) A proposal to revive the genus Kitasatospora (Omura, Takahashi, Iwai, and Tanaka 1982). Int J Syst Bacteriol 47(4):1048–1054
Acknowledgments
Kui Hong, Wasu Pathom-aree, Kannika Duangmal and Rattanaporn Srivibool are grateful for the financial support from the NSFC (31111140297)-NRCT Grant for the Thai–Chinese Cooperative Project, “Actinomycetes from coastal marine and mangrove sediments of Eastern Thailand and their ability to produce bioactive compounds” and for the Chinese MOST-TICA Sino–Thai Joint Research and Development Project, “Selective isolation and prepharmaceutical research on novel and rare actinomycetes from tropical marine and terrestrial habitats” (19-505J).
Author information
Authors and Affiliations
Corresponding authors
Additional information
The GenBank/EMBL/DDBJ accession numbers for the 16S rRNA gene sequence of strain HV38T is KF767859.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ruan, Cy., Zhang, L., Ye, Ww. et al. Streptomyces ferrugineus sp. nov., isolated from mangrove soil in Thailand. Antonie van Leeuwenhoek 107, 39–45 (2015). https://doi.org/10.1007/s10482-014-0301-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-014-0301-6