Abstract
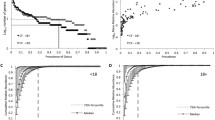
The aim of this study was to investigate the polymicrobial communities in an adult Cystic Fibrosis population stratified by gender and the most common CFTR mutation, F508del. In this pilot study, DNA was extracted from sputum samples of 29 adult patients (16 male: 13 female) with an F508del mutation in a stable clinical state. Universal primers were used to amplify DNA from bacterial and fungal communities and the resulting fragments were analysed by denaturing gradient gel electrophoresis. Bacterial profiles showed a significant effect of gender (P = 0.046) and P. aeruginosa carriage (P = 0.034) on community structure. Bacterial communities were found to be randomly assembled. Fungal community analysis found that F508del homozygous patients had a greater diversity than heterozygous patients (P = 0.007). This study indicates that the bacterial lung communities of adult CF patients are randomly assembled but have distinct gender based differences. Furthermore, the fungal communities colonising the CF lung are more diverse in F508 homozygotes. This is the first paper to identify a reduced bacterial diversity in female patients with CF and to implicate more severe CFTR genotypes with increased risk of infection with multiple fungal species.



Similar content being viewed by others
References
Amin R, Dupuis A, Aaron SD, Ratjen F (2010) The effect of chronic infection with Aspergillus fumigatus on lung function and hospitalization in patients with cystic fibrosis. Chest 137:171–176. doi:10.1378/chest.09-1103
Bittar F, Richet H, Dubus JC, Reynaud-Gaubert M, Stremler N, Sarles J, Raoult D, Rolain JM (2008) Molecular detection of multiple emerging pathogens in sputa from cystic fibrosis patients. PLoS ONE 3:e2908. doi:10.1371/journal.pone.0002908
Block JK, Vandemheen KL, Tullis E et al (2006) Predictors of pulmonary exacerbations in patients with cystic fibrosis infected with multi-resistant bacteria. Thorax 61:969–974. doi:10.1136/thx.2006.061366
Bouchara JP, Hsieh HY, Croquefer S, Barton R, Marchais V, Pihet M, Chang TC (2009) Development of an oligonucleotide array for direct detection of fungi in sputum samples from patients with cystic fibrosis. J Clin Microbiol 47:142–152. doi:10.1128/JCM.01668-08
Burns JL, Emerson J, Stapp JR, Yim DL, Krzewinski J, Louden L, Ramsey BW, Clausen CR (1998) Microbiology of sputum from patients at cystic fibrosis centres in the United States. Clin Infect Dis 27:158–163
Charlson ES, Bittinger K, Haas AR, Fitzgerald AS, Frank I, Yadav A, Bushman FD, Collman RG (2011) Topographical continuity of bacterial populations in the healthy human respiratory tract. Am Respir Crit Care Med 184:957–963. doi:10.1164/rccm.201104-0655OC
Chotirmall SH, Greene CM, Oglesby IK, Thomas W, O’Neill SJ, Harvey BJ, McElvaney NG (2010a) Beta-estradiol inhibits IL-8 in cystic fibrosis by up-regulating secretory leucoprotease inhibitor. Am Respir Crit Care Med 182:62–72. doi:10.1164/rccm.201001-0053OC
Chotirmall SH, O’Donoghue E, Bennett K, Gunaratnam C, O’Neill SJ, McElvaney NG (2010b) Sputum Candida albicans presages FEV1 decline and hospital-treated exacerbations in cystic fibrosis. Chest 138:1186–1195. doi:10.1378/chest.09-2996
Cox MJ, Allgaier M, Taylor B et al (2010) Airway microbiota and pathogen abundance in age-stratified cystic fibrosis patients. PLoS ONE 5:e11044. doi:10.1371/journal.pone.0011044
Dumbrell AJ, Nelson M, Helgason T, Dytham C, Fitter AH (2010) Relative roles of niche and neutral processes in structuring a soil microbial community. ISME J 4:337–345. doi:10.1038/ismej.2009.122
Emerson J, McNamara S, Buccat AM, Worrell K, Burns JL (2010) Changes in cystic fibrosis sputum microbiology in the United States between 1995 and 2008. Pediatr Pulmonol 45:363–370. doi:10.1002/ppul.21198
Erb-Downward JR, Thompson DL, Han MK et al (2011) Analysis of the lung microbiome in the “healthy” smoker and in COPD. PLoS ONE 6:e16384. doi:10.1371/journal.pone.0016384
Giraud S, Pihet M, Razafimandimby B, Carrère J, Degand N, Mely L, Favennec L, Dannaoui E, Bouchara JP, Calenda A (2010) Geosmithia argillacea: an emerging pathogen in patients with cystic fibrosis. J Clin Microbiol 48:2381–2386. doi:10.1128/JCM.00047-10
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: palaeontological statistics software package for education and data analysis. Palaeontologia Electronica 4:9
Kerem B, Rommens JM, Buchanan JA, Markiewicz D, Cox TK, Chakravarti A, Buchwald M, Tsui LC (1989) Identification of the cystic fibrosis gene: genetic analysis. Science 245:1073–1080
Kerem E, Reisman J, Corey M, Canny GJ, Levison H (1992) Prediction of mortality in patients with cystic fibrosis. N Engl J Med 326:1187–1191. doi:10.1056/NEJM199204303261804
Klepac-Ceraj V, Lemon KP, Martin TR et al (2010) Relationship between cystic fibrosis respiratory tract bacterial communities and age, genotype, antibiotics and Pseudomonas aeruginosa. Environ Microbiol 12:1293–1303. doi:10.1111/j.1462-2920.2010.02173.x
Lane D (1991) Nucleic acid techniques in bacterial systematics, 1st edn. Wiley, New York
Levy H, Kalish LA, Cannon CL, García KC, Gerard C, Goldmann D, Pier GB, Weiss ST, Colin AA (2008) Predictors of mucoid Pseudomonas colonization in cystic fibrosis patients. Pediatr Pulmonol 43:463–471. doi:10.1002/ppul.20794
McKone EF, Goss CH, Aitken ML (2006) CFTR genotype as a predictor of prognosis in cystic fibrosis. Chest 130:1441–1447. doi:10.1378/chest.130.5.1441
Muyzer G, de Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Nelson A, De Soyza A, Bourke SJ, Perry JD, Cummings SP (2010) Assessment of sample handling practices on microbial activity in sputum samples from patients with cystic fibrosis. Lett Appl Microbiol 51:272–277. doi:10.1111/j.1472-765X.2010.02891.x
Nelson A, De Soyza A, Perry JD, Sutcliffe IC, Cummings SP (2012) Polymicrobial challenges to Koch’s postulates: ecological lessons from the bacterial vaginosis and cystic fibrosis microbiomes. Innate Immun 18:774–783. doi:10.1177/1753425912439910
Rogers GB, Carroll MP, Serisier DJ, Hockey PM, Jones G, Bruce KD (2004) Characterization of bacterial community diversity in cystic fibrosis lung infections by use of 16S ribosomal DNA terminal restriction fragment length polymorphism profiling. J Clin Microbiol 42:5176–5183. doi:10.1128/JCM.42.11.5176-5183.2004
Sandhu GS, Kline BC, Stockman L, Roberts GD (1995) Molecular probes for diagnosis of fungal infections. J Clin Microbiol 33:2913–2919
Stressmann FA, Rogers GB, Klem ER et al (2011) Analysis of the bacterial communities present in lungs of patients with cystic fibrosis from American and British centers. J Clin Microbiol 49:281–291. doi:10.1128/JCM.01650-10
Tourlomousis P, Kemsley EK, Ridgway KP, Toscano MJ, Humphrey TJ, Narbad A (2010) PCR-denaturing gradient gel electrophoresis of complex microbial communities: a two-step approach to address the effect of gel-to-gel variation and allow valid comparisons across a large dataset. Microb Ecol 59:776–786. doi:10.1007/s00248-009-9613-x
Tunney MM, Field TR, Moriarty TF et al (2008) Detection of anaerobic bacteria in high numbers in sputum from patients with cystic fibrosis. Am J Respir Crit Care Med 177:995–1001. doi:10.1164/rccm.200708-1151OC
Van Der Gast CJ, Walker AW, Stressmann FA, Rogers GB, Scott P, Daniels TW, Carroll MP, Parkhill J, Bruce KD (2010) Partitioning core and satellite taxa from within cystic fibrosis lung bacterial communities. ISME J. doi:10.1038/ismej.2010.175
Yoon SS, Hennigan RF, Hilliard GM et al (2002) Pseudomonas aeruginosa anaerobic respiration in biofilms: relationships to cystic fibrosis pathogenesis. Dev Cell 3:593–603
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
Stephen P. Cummings and Anthony De Soyza: Joint senior authors.
Rights and permissions
About this article
Cite this article
Nelson, A., Perry, A., Perry, J.D. et al. Determining the influence of environmental and patient specific factors on the polymicrobial communities of the cystic fibrosis airway. Antonie van Leeuwenhoek 103, 755–762 (2013). https://doi.org/10.1007/s10482-012-9857-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-012-9857-1